Abstract
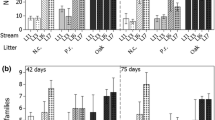
Tank-forming bromeliads, suspended in the rainforest canopy, possess foliage arranged in compact rosettes capable of long-term retention of rainwater. This large and unique aquatic habitat is inhabited by microorganisms involved in the important decomposition of impounded material. Moreover, these communities are likely influenced by environmental factors such as pH, oxygen, and light. Bacterial community composition and diversity was determined for the tanks of several bromeliad species (Aechmea and Werauhia) from northern Costa Rica, which span a range of parameters, including tank morphology and pH. These were compared with a nearby forest soil sample, an artificial tank (amber bottle), and a commercially available species (Aechmea). Bacterial community diversity, as measured by 16S rRNA analysis and tRFLP, showed a significant positive correlation with tank pH. A majority of 16S rRNA bacterial phylotypes found in association with acidic bromeliad tanks of pH < 5.1 were affiliated with the Alphaproteobacteria, Acidobacteria, Planctomycetes, and Bacteroidetes, and were similar to those found in acidic peat bogs, yet distinct from the underlying soil community. In contrast, bromeliads with tank pH > 5.3, including the commercial bromeliad with the highest pH (6.7), were dominated by Betaproteobacteria, Firmicutes, and Bacteroidetes. To empirically determine the effect of pH on bacterial community, the tank pH of a specimen of Aechmea was depressed, in the field, from 6.5 to 4.5, for 62 days. The resulting community changed predictably with decreased abundance of Betaproteobacteria and Firmicutes and a concomitant increase in Alphaproteobacteria and Acidobacteria. Collectively, these results suggest that bromeliad tanks provide important habitats for a diverse microbial community, distinct from the surrounding environment, which are influenced greatly by acid–base conditions. Additionally, total organic carbon (∼46%) and nitrogen (∼2%) of bromeliad-impounded sediment was elevated relative to soil and gene surveys confirmed the presence of both chitinases and nitrogenases, suggesting that bromeliad tanks may provide important habitats for microbes involved in the biological cycling of carbon and nitrogen in tropical forests.







Similar content being viewed by others
References
Beals EW (1984) Bray-Curtis ordination: an effective strategy for analysis of multivariate ecological data. Adv Ecol Res 14:1–55
Benlloch S, Rodriguez-Valera F, Martinez-Murcia AJ (1995) Bacterial diversity in two coastal lagoons deduced from 16S rDNA PCR amplification and partial sequencing. FEMS Microbiol Ecol 18:267–280
Benzing DH (1970) Foliar permeability and the absorption of minerals and organic nitrogen by certain tank bromeliads. Bot Gaz 131:23–31
Benzing DH (2000) Bromeliaceae: profile of an adaptive radiation. Cambridge University Press, Cambridge
Benzing DH, Derr JA, Titus JE (1972) The water chemistry of microcosms associated with the bromeliad Aechmea bracteata. Amer Mid Natr 87:60–70
Bermudes D, Benzing DH (1991) Nitrogen fixation in association with Ecuadorean bromeliads. J Trop Ecol 7:531–536
Bissett A, Bowman J, Burke C (2006) Bacterial diversity in organically-enriched fish farm sediments. FEMS Microbiol Ecol 55:48–56
Brighigna L, Montaini P, Favilli F, Trejo AC (1992) Role of nitrogen-fixing bacterial microflora in the epiphytism of Tillandsia (Bromeliaceae). Am J Bot 79:723–727
Burkert U, Warnecke F, Babenzien D, Zwirnmann E, Pernthaler J (2003) Members of a readily enriched beta-proteobacterial clade are common in surface waters of a humic lake. Appl Environ Microbiol 69:6550–6559
Dedysh SN (2009) Exploring methanotroph diversity in acidic northern wetlands: molecular and cultivation-based studies. Microbiol 78:665–669
Dedysh SN, Khmelenina VN, Suzina NE, Trotsenko YA, Semrau JD, Liesack W, Tiedje JM (2002) Methylocapsa acidiphila gen. nov., sp. nov., a novel methane-oxidizing and dinitrogen-fixing acidophilic bacterium from Sphagnum bog. Int J Syst Evol Microbiol 52:251–261
Dedysh SN, Pankratov TA, Belova SE, Kulichevskaya IS, Liesack W (2006) Phylogenetic analysis and in situ identification of bacteria community composition in an acidic Sphagnum peat bog. Appl Environ Microbiol 27:2110–2117
DeSantis TZ, Hugenholtz P, Larsen N, Rojas M, Brodie EL, Keller K (2006) Greengenes, a chimera-checked 16S rRNA gene database and workbench compatible with ARB. Appl Environ Microbiol 72:5069–5072
Eichorst SA, Breznak JA, Schmidt TM (2007) Isolation and characterization of soil bacteria that define Terriglobus gen. nov., in the Phylum Acidobacteria. Appl Environ Microbiol 73:2708–2717
Fish D (1983) Phytotelmata: terrestrial plants as hosts for aquatic insect communities. In: Frank JH, Lounibos LP (eds) Phytotelmata: flora and fauna. Plexus, New Jersey, pp 1–28
Goffredi SK, Orphan VJ (2010) Bacterial community shifts in taxa and diversity in response to localized organic loading in the deep sea. Environ Microbiol 12:344–363
Guimaraes-Souza BA, Mendes GB, Bento L, Morotta H, Santoro AL, Esteves FA, Pinho L, Farjalla VF, Enrich-Prast A (2006) Limnological parameters in the water accumulated in tropical bromeliads. Acta Limnol Bras 18:47–53
Hanson RS, Netrusov AI, Tsuji K (1992) The obligate methanotrophic bacteria Methylococcus, Methylomonas, and Methylosinus. In: Balows A, Truper HG, Dworkin M, Harder W, Schleifer K-H (eds) The prokaryotes. Springer, New York, pp 2350–2364
Hartman WH, Richardson CJ, Vilgalys R, Bruland GL (2008) Environmental and anthropogenic controls over bacterial communities in wetland soils. Proc Natl Acad Sci 105:17842–17847
Imhoff JF (2006) The phototrophic alpha-proteobacteria. In: Dworkin M, Falkow S, Rosenberg E, Schleifer K-H, Stackebrandt E (eds) The prokaryotes, vol 5., pp 41–64
Inselsbacher E, Cambui CA, Richter A, Stange CF, Mercier H, Wanek W (2007) Microbial activities and foliar uptake of nitrogen in the epiphytic bromeliad Vriesea gigantea. New Phytol 175:311–320
Jollès P, Muzzarelli RAA (1999) Chitin and Chitinases. Birkhäuser Verlag, Basel
Kawahara N, Shigematsu K, Miyadai T, Kondo R (2009) Comparison of bacterial communities in fish farm sediments along an organic enrichment gradient. Aquaculture 287:107–113
Kulichevskaya IS, Guzev VS, Gorlenko VM, Liesack W, Dedysh SN (2006) Rhodoblastus sphagnicola sp. nov., a novel acidophilic purple non-sulfur bacterium from Sphagnum peat bog. Int J Syst Evol Microbiol 56:1397–1402
Laessle AM (1961) A micro-limnological study of Jamaican bromeliads. Ecol 42:499–517
Lane DJ (1991) 16S/23S rRNA sequencing. In: Stackebrandt E, Goodfellow M (eds) Nucleic acid techniques in bacterial systematics. Wiley, New York, pp 115–175, In
Lauber CL, Hamady M, Knight R, Fierer N (2009) Pyrosequencing-based assessment of soil pH as a predictor of soil bacterial community structure at the continental scale. Appl Environ Microbiol 75:5111–5120
Lopez LCS, Alves RRD, Rios RI (2009) Micro-environmental factors and the endemism of bromeliad aquatic fauna. Hydrobiol 625:151–156
Loy A, Schulz C, Lucker S, Schopfer-Wendels A, Stoecker K, Baranyi C, Lehner A, Wagner M (2005) 16S rRNA gene-based oligonucleotide microarray for environmental monitoring of the betaproteobacterial order “Rhodocyclales”. Appl Environ Microbiol 71:1373–1386
Ludwig W, Strunk O, Westram R, Richter L, Meier H, Yadhukumar EA (2004) ARB: a software environment for sequence data. Nucleic Acids Res 32:1363–1371
McCune B, Mefford MJ (2006) PC–ORD: multivariate analysis of ecological data, vol. MjM Software, Glenedon Beach.
Mehta MP, Butterfield DA, Baross JA (2003) Phylogenetic diversity of nitrogenase (nifH) genes in deep-sea and hydrothermal vent environments of the Juan de Fuca Ridge. Appl Environ Microbiol 69:960–970
Merwin MC, Rentmeester SA, Nadkarni NM (2003) The influence of host tree species on the distribution of epiphytic bromeliads in experimental monospecific plantations, La Selva, Costa Rica. Biotropica 35:37–47
Pittl E, Innerebner G, Wanek W, Insam H (2010) Microbial communities of arboreal and ground soils in the Esquinas rainforest, Costa Rica. Plant Soil 329:65–74
Richardson BA (1999) The bromeliad microcosm and the assessment of faunal diversity in a neotropical forest. Biotropica 31:321–336
Rui J, Peng J, Lu Y (2009) Succesion of bacterial populations during plant residue decomposition in rice field soil. Appl Environ Microbiol 75:4879–4886
Sait M, Davis KER, Janssen PH (2006) Effect of pH on isolation and distribution of members of subdivision 1 of the phylum Acidobacteria occurring in soil. Appl Environ Microbiol 72:1852–1857
Spring S (2006) The genera Leptothrix and Sphaerotilus. In: Dworkin M, Falkow S, Rosenberg E, Schleifer K-H, Stackebrandt E (eds) The Prokaryotes, vol. 5., pp 758–777
Sugden AM, Robins RJ (1979) Aspects of the ecology of vascular epiphytes in colombian cloud forests, I. The distribution of the epiphytic flora. Biotropica 11:173–188
Swofford DL (1998) PAUP*: phylogenetic analysis using parsimony (*and other methods). Sinauer, Sunderland.
van Veen WL, Mulder EG, Deinema MH (1978) The Sphaerotilus-Leptothrix group of bacteria. Microbiol Rev 42:329–356
Wang Q, Garrity GM, Tiedje JM, Cole JR (2007) Naïve Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl Environ Microbiol 73:5261–5267
Ward NL, Challacombe JF, Janssen PH, Henrissat B, Coutinho PM, Wu M, Xie G, Haft DH, Sait M, Badger J, Barabote RD, Bradley B, Brettin TS, Brinkac LM (2009) Three genomes from the phylum Acidobacteria provide insight into the lifestyles of these microorganisms in soils. Appl Environ Microbiol 75:2046–2056
Winkler U, Zotz G (2008) Highly efficient uptake of phosphorus in epiphytic bromeliads. Ann Bot (Lond) 103:477–484
Xiao X, Yin X, Lin J, Sun L, You Z, Wang P, Wang F (2005) Chitinase genes in lake sediments of Ardley Island, Antarctica. Appl Environ Microbiol 71:7904–7909
Yamada A, Inoue T, Noda S, Hongoh Y, Ohkuma M (2007) Evolutionary trend of phylogenetic diversity of nitrogen fixation genes in the gut community of wood-feeding termites. Mol Ecol 16:3768–3777
Yarwood SA, Myrold DD, Högberg MN (2009) Termination of belowground C allocation by trees alters soil fungal and bacterial communities in a boreal forest. FEMS Microbiol Ecol 70:151–162
Zhang W, Ki J-S, Qian P-Y (2008) Microbial diversity in polluted harbor sediments I: bacterial community assessment based on four clone libraries of 16S rDNA. Estuar Coast Shelf Sci 76:668–681
Acknowledgments
The authors thank Dr. Gretchen North for invaluable scientific advice with regard to plant biology and as co-investigator on the bromeliad project overall; Dr. Beth Braker, for introducing us to the wonders of the rainforest; Bernal Matarrita, among others, for his laboratory support at La Selva; Dr. Deedra McClearn and members of the Organization for Tropical Studies for their support of this project; Dr. Victoria Orphan for use of the DNA sequencing facilities at the California Institute of Technology; Occidental College undergraduates involved in the Spring 2010 semester of Microbial Diversity (Bio325); Pamela Imperiale-Hagerman for laboratory assistance and data collection, and, finally, B. Harrison for help and interpretation of the dataset with PC-ORD. Funding for this project included, in part, a NSF grant to B. Braker (Occidental College, OISE-0526551), a HHMI grant to Occidental College, as well as the Undergraduate Research Center (Academic Student and Summer Research Projects), and a Faculty Enrichment grant from Occidental College.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Goffredi, S.K., Kantor, A.H. & Woodside, W.T. Aquatic Microbial Habitats Within a Neotropical Rainforest: Bromeliads and pH-Associated Trends in Bacterial Diversity and Composition. Microb Ecol 61, 529–542 (2011). https://doi.org/10.1007/s00248-010-9781-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00248-010-9781-8