Abstract
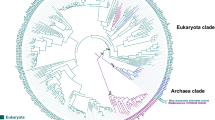
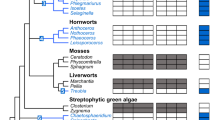
Alternative oxidases (AOXs) are mitochondrial cyanide-resistant membrane-bound metallo-proteins catalyzing the oxidation of ubiquinol and the reduction of oxygen to water bypassing two sites of proton pumping, thus dissipating a major part of redox energy into heat. Here, the structure of Arabidopsis thaliana AOX 1A has been modeled using the crystal structure of Trypanosoma brucei AOX as a template. Analysis of this model and multiple sequence alignment of members of the AOX family from all kingdoms of Life indicate that AOXs display a high degree of conservation of the catalytic core, which is formed by a four-α-helix bundle, hosting the di-iron catalytic site, and is flanked by two additional α-helices anchoring the protein to the membrane. Plant AOXs display a peculiar covalent dimerization mode due to the conservation in the N-terminal region of a Cys residue forming the inter-monomer disulfide bond. The multiple sequence alignment has also been used to infer a phylogenetic tree of AOXs whose analysis shows a polyphyletic origin for the AOXs found in Fungi and a monophyletic origin of the AOXs of Eubacteria, Mycetozoa, Euglenozoa, Metazoa, and Land Plants. This suggests that AOXs evolved from a common ancestral protein in each of these kingdoms. Within the Plant AOX clade, the AOXs of monocotyledon plants form two distinct clades which have unresolved relationships relative to the monophyletic clade of the AOXs of dicotyledonous plants. This reflects the sequence divergence of the N-terminal region, probably due to a low selective pressure for sequence conservation linked to the covalent homo-dimerization mode.






Similar content being viewed by others
Abbreviations
- Δ9:
-
Desaturase
- Stearoyl:
-
Acyl carrier desaturase
- AOX:
-
Alternative oxidase
- ML:
-
Maximum Likelihood
- MMO:
-
Methane–monooxygenase
- RNR:
-
R2 subunit from ribonucleotide reductase
- ROS:
-
Reactive oxygen species
- TAO:
-
Trypanosomal alternative oxidase
- AtAOX:
-
Arabidopsis thaliana AOX 1A
References
Abascal F, Zardoya R, Posada D (2005) ProtTest: selection of best-fit models of protein evolution. Bioinformatics 21:2104–2105
Abele E, Philip E, Gonzalez PM, Puntarulo S (2007) Marine invertebrate mitochondria and oxidative stress. Front Biosci 12:933–946
Ajayi WU, Chaudhuri M, Hill GC (2002) Site-directed mutagenesis reveals the essentiality of the conserved residues in the putative diiron active site of the trypanosome alternative oxidase. J Biol Chem 277:8187–8193
Akaike H (1974) A new look at the statistical model identification. IEEE Trans Automat Contr AC-19:716–723
Albury MS, Elliott C, Moore AL (2009) Towards a structural elucidation of the alternative oxidase in plants. Physiol Plant 137:31–327
Altschul SF, Madden TL, Schaffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucl Acids Res 25:3389–3402
Andersson ME, Nordlund P (1999) A revised model of the active site of alternative oxidase. FEBS Lett 449:17–22
Berthold DA, Andersson ME, Nordlund P (2000) New insight into the structure and function of the alternative oxidase. Biochim Biophys Acta 1460:241–254
Castresana J (2000) Selection of conserved blocks from multiple alignments for their use in phylogenetic analysis. Mol Biol Evol 17:540–552
Chaudhuri M, Hill GC (1996) Cloning, sequencing and functional activity of the Trypanosoma brucei brucei alternative oxidase. Mol Biochem Parasitol 83:125–129
Chen SH, Su SY, Lo CZ, Chen KH, Huang TJ, Kuo BH, Lin CY (2009) PALM: a paralleled and integrated framework for phylogenetic inference with automatic likelihood model selectors. PLoS One 4:e8116
Clifton R, Millar AH, Whelan J (2006) Alternative oxidase in Arabidopsis: a comparative analysis of differential expression in gene family provides new insights into function of non-phosphorylating bypasses. Biochim Biophys Acta 1757:730–741
Cooper CE, Brown GC (2008) The inhibition of mitochondrial cytochrome oxidase by the gases carbon monoxide, nitric oxide, hydrogen cyanide and hydrogen sulfide: chemical mechanism and physiological significance. J Bioenerg Biomembr 40:533–539
Finnegan PM, Soole KL, Umbach AL (2004) Alternative mitochondrial electron transport proteins in higher plants. In: Day DA, Millar AH, Whelan J (eds) Plant mitochondria: from gene to function, vol 17., Advances in photosynthesis and respirationKluwer, Dordrecht, pp 163–230
Guex N, Peitsch MC (1997) SWISS-MODEL and the Swiss-PdbViewer: an environment for comparative protein modeling. Electrophoresis 18:2714–2723
Guindon S, Dufayard JF, Lefort V, Anisimova M, Hordijk W, Gascuel O (2010) New algorithms and methods to estimate maximum-likelihood phylogenies: assessing the performance of phyml 3.0. Syst Biol 59:307–321
Hall FR, Hollingworth RM, Shankland DL (1971) Cyanide tolerance in millipedes: the biochemical basis. Comp Biochem Physiol 38:723–737
Huson DH, Bryant D (2006) Application of phylogenetic networks in evolutionary studies. Mol Biol Evol 23:254–267
Keeling P, eander BS, Simpson A (2009) Eukaryotes. Eukaryota, organisms with nucleated cells. Version 28 Oct 2009. http://tolweb.org/Eukaryotes/3/2009.10.28 in The Tree of Life Web Project, http://tolweb.org/
Kellogg EA (2001) Evolutionary history of the grasses. Plant Physiol 125:1198–1205
Kim DE, Chivian D, Baker D (2004) Protein structure prediction and analysis using the Robetta server. Nucl Acids Res 32:526–531
Le SQ, Gascuel O (2008) An improved general amino acid replacement matrix. Mol Biol Evol 25:1307–1320
Li Q, Bai Z, O’Donnell A, Harvey LM, Hoskisson PA, McNeil B (2011) Oxidative stress in fungal fermentation processes: the roles of alternative respiration. Biotechnol Lett 33:457–467
Maxwell DP, Wang Y, McIntosh L (1999) The alternative oxidase lowers mitochondrial reactive oxygen production in plant cells. Proc Natl Acad Sci USA 96:8271–8276
McDonald AE (2008) Alternative oxidase: an inter-kingdom perspective on the function and regulation of this broadly distributed ‘cyanide resistant’ terminal oxidase. Funct Plant Biol 35:535–552
McDonald A, Vanlerberghe G (2004) Branched mitochondrial electron transport in the animalia: presence of alternative oxidase in several animal phyla. IUBMB Life 56:333–341
McDonald AE, Vanlerberghe GC, Staples JF (2009) Alternative oxidase in animals: unique characteristics and taxonomic distribution. J Exp Biol 212:2627–2634
Millenaar FF, Lambers H (2003) The alternative oxidase: in vivo regulation and function. Plant Biol 5:2–15
Moore AL, Umbach AL, Siedow JN (1995) Structure-function relationship of the alternative oxidase of plant mitochondria: a model of the active site. J Bioenerg Biomembr 27:367–377
Moore AL, Shiba T, Young L, Harada S, Kita K, Ito K (2013) Unraveling the heater: new insights into the structure of the alternative oxidase. Annu Rev Plant Biol 64:637–663
Neimanis K, Staples JF, Hüner NP, McDonald AE (2013) Identification, expression, and taxonomic distribution of alternative oxidases in non-angiosperm plants. Gene 526:275–286
Parsons HL, Yip JY, Vanlerberghe GC (1999) Increased respiratory restriction during phosphate-limited growth in transgenic tobacco cells lacking alternative oxidase. Plant Physiol 121:1309–1320
Pettersen EF, Goddard TD, Huang CC, Couch GS, Greenblatt DM, Meng EC, Ferrin TE (2004) UCSF Chimera—a visualization system for exploratory research and analysis. J Comput Chem 25:1605–1612
Polidoros AN, Mylona PV, Arnholdt-Schmitt B (2009) AOX gene structure, transcript variation and expression in plants. Physiol Plant 137:342–353
Purvis AC, Shewfelt RL (1993) Does the alternative pathway ameliorate chilling injury in sensitive plant tissue? Physiol Plant 88:712–718
Rasmusson AG, Geisler DA, Moller IM (2008) The multiplicity of dehydrogenases in the electron transport chain of plant mitochondria. Mitochondrion 8:47–60
Rhoads DM, Umbach AL, Sweet CR, Lennon AM, Rauch GS, Siedow JN (1998) Regulation of the cyanide-resistant alternative oxidase of plant mitochondria. Identification of the cysteine residue involved in α-keto acid stimulation and intersubunit disulfide bond formation. J Biol Chem 273:30750–30756
Roberts CW, Roberts F, Henriquez FL, Akiyoshi D, Samuel BU, Richards TA, Mihous W, Kyle D, McIntosh L, Hill GC, Chaudhuri M, Tzipori S, McLeod R (2004) Evidence for mitochondrial-derived alternative oxidase in the apicomplexan parasite Cryptosporidium parvum: a potential anti-microbial agent target. Int J Parasitol 34:297–308
Roy A, Kucukural A, Zhang Y (2010) I-TASSER: a unified platform for automated protein structure and function prediction. Nat Protoc 5:725–738
Shiba T, Kido Y, Sakamoto K, Inaoka DK, Tsuge C (2013) Structure of the trypanosome cyanide insensitive alternative oxidase. Proc Natl Acad Sci USA 110:4580–4585
Siedow JN, Umbach AL, Moore AL (1995) The active site of the cyanide-resistant oxidase from plant mitochondria contains a binuclear iron center. FEBS Lett 362:10–14
Sievers F, Wilm A, Dineen D, Gibson TJ, Karplus K, Li W, Lopez R, McWilliam H, Remmert M, Söding J, Thompson JD, Higgins DG (2011) Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol Syst Biol 7:539
Stechmann A, Hamblin K, Pérez-Brocal V, Gaston D, Richmond GS, van der Giezen M, Clark CG, Roger AJ (2008) Organelles in Blastocystis that blur the distinction between mitochondria and hydrogenosomes. Curr Biol 18:580–585
Stenmark P, Nordlund P (2003) A prokaryotic alternative oxidase present in the bacterium Novosphingobium aromaticivorans. FEBS Lett 552:189–192
Sun J, Lu X, Rinas U, Ping Zeng A (2007) Metabolic peculiarities of Aspergillus niger disclosed by comparative metabolic genomics. Genome Biol 8(9):R182
Sussarellu R, Dudognon T, Fabioux C, Soudant P, Moraga D, Kraffe E (2013) Rapid mitochondrial adjustments in response to short-term hypoxia and re-oxygenation in the Pacific oyster, Crassostrea gigas. J Exp Biol 216:1561–1569
Suzuki T, Hashimoto T, Yabu Y, Majiwa PA, Ohshima S, Suzuki M, Lu S, Hato M, Kido Y, Sakamoto K, Nakamura K, Kita K, Ohta N (2005) Alternative oxidase (AOX) genes of African trypanosomes: phylogeny and evolution of AOX and plastid terminal oxidase families. J Eukaryot Microbiol 52(4):374–381
Talavera G, Castresana J (2007) Improvement of phylogenies after removing divergent and ambiguously aligned blocks from protein sequence alignments. Syst Biol 56:564–577
Tanudji M, Sjöling S, Glaser E, Whelan J (1999) Signals required for the import and processing of the alternative oxidase into mitochondria. J Biol Chem 15:1286–1293
Thirkettle-Watts D, McCabe TC, Clifton R, Moore C, Finnegan PM, Day DA, Whelan J (2003) Analysis of the alternative oxidase promoters from soybean. Plant Physiol 133:1158–1169
Thompson JD, Higgins DG, Gibson TJ (1994) CLUSTAL W: improving the sensitivity of progressive multiple sequence alignments through sequence weighting, position specific gap penalties and weight matrix choice. Nucl Acids Res 22:4673–4680
Turrens JF (2003) Mitochondrial formation of reactive oxygen species. J Physiol 552:335–344
Umbach AL, Siedow JN (1993) Covalent and noncovalent dimers of the cyanide-resistant alternative oxidase protein in higher plant mitochondria and their relationship to enzyme activity. Plant Physiol 103:845–854
Umbach AL, Wiskich JT, Siedow JN (1994) Regulation of alternative oxidase kinetics by pyruvate and intermolecular disulfide bond redox status in soybean seedling mitochondria. FEBS Lett 348:181–184
Van Aken O, Giraud E, Clifton R, Whelan J (2009) Alternative oxidase: a target and regulator of stress responses. Physiol Plant 137:354–361
Van Hellemond JJ, Simons B, Millenaar FF, Tielens AG (1998) A gene encoding the plant-like alternative oxidase is present in Phytomonas but absent in Leishmania spp. J Eukaryot Microbiol 45:426–430
Vanlerberghe GC (2013) Alternative oxidase: a mitochondrial respiratory pathway to maintain metabolic and signaling homeostasis during abiotic and biotic stress in plants. Int J Mol Sci 14:6805–6847
Vanlerberghe GC, McIntosh L (1997) Alternative oxidase: from gene to function. Annu Rev Plant Physiol Plant Mol Biol 48:703–734
Veiga A, Arrabaça JD, Loureiro-Dias MC (2003) Cyanide–resistant respiration, a very frequent metabolic pathway in yeasts. FEMS Yeast Res 3:239–245
Vishwakarma A, Bashyam L, Senthilkumaran B, Scheibe R, Padmasree K (2014) Physiological role of AOX1a in photosynthesis and maintenance of cellular redox homeostasis under high light in Arabidopsis thaliana. Plant Physiol Biochem 81:44–53
Williams BAP, Elliot C, Burri L, Kido Y, Kita K et al (2010) A broad distribution of the alternative oxidase in microsporidian parasites. PLoS Pathog 6(2):e1000761
Xu D, Zhang Y (2012) Ab initio protein structure assembly using continuous structure fragments and optimized knowledge-based force field. Proteins 80:1715–1735
Yukioka H, Inagaki S, Tanaka R, Katoh K, Miki N, Mizutani A, Masuko M (1998) Transcriptional activation of the alternative oxidase gene of the fungus Magnaporthe grisea by a respiratory-inhibiting fungicide and hydrogen peroxide. Biochim Biophys Acta 1442:161–169
Acknowledgments
D.S. is supported by FCT post-doctoral fellowship SFRH/BPD/105274/2014 from the European Social Fund and Portuguese Ministério da Educação e Ciência, and by the project “Genomics and Evolutionary Biology” cofinanced by North Portugal Regional Operational Programme 2007/2013 (ON.2-O Novo Norte), under the National Strategic Reference Framework (NSRF), through the European Regional Development Fund (ERDF).
Author Contributions
RP collected the AOXs sequences, carried out the molecular modeling studies and drafted the manuscript. DS carried out the phylogenetic analysis and helped to draft the manuscript. VB carried out ab initio molecular modeling studies and electrostatic potential calculations. RA participated in the interpretation of the results and helped to draft the manuscript. PA and FP conceived the study, participated in its design and coordination and helped to draft the manuscript. All authors read and approved the final manuscript.
Author information
Authors and Affiliations
Corresponding author
Additional information
Rosa Pennisi and Daniele Salvi contributed equally to this work.
Database: AtAOX model data are available in the PMDB database under the accession number PM0080189.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Pennisi, R., Salvi, D., Brandi, V. et al. Molecular Evolution of Alternative Oxidase Proteins: A Phylogenetic and Structure Modeling Approach. J Mol Evol 82, 207–218 (2016). https://doi.org/10.1007/s00239-016-9738-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00239-016-9738-8