Abstract
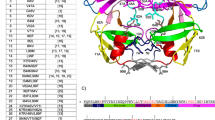
Developing a new HIV-1 protease (HIV-1 PR) inhibitor is still a challenging task to overcome the drug resistance mutations in the HIV-PR. In this study, docking simulations of chromone derivatives against wild type and eleven mutant variants HIV-1 PR were investigated using GOLD and Autodock programs. From both GOLD and Autodock results, chromone 3, the experimentally observed highly potent HIV-1 PR inhibitor, showed stronger binding affinity against every studied mutant strain (2AVS, 2AVO, 2AVV, 1MES, 1MET, 1MEU, 1SDU, 1SDV, 1C6Y, 2F8O, and 1SH9) than the wild-type enzyme (1AJX). Chromone 32, another potent inhibitor as well as chromones 33, 34, 37, and 47 also showed high binding interaction with several mutant-type enzymes. The coherent picture of the interactions at the active sites of mutant PR should facilitate the further design and development of new potent inhibitor against multidrug-resistant virus.






Similar content being viewed by others
References
Ala PJ, Huston EE, Klabe RM, McCabe DD, Duke JL, Rizzo CJ, Korant BD, DeLoskey RJ, Lam PYS, Hodge CN, Chang CH (1997) Molecular basis of HIV-1 protease drug resistance: structural analysis of mutant protease complexed with cyclic urea inhibitors. Biochemistry 36:1573–1580
Barbour JD, Wrin T, Grant RM, Martin JN, Segal MR, Petropoulos CJ, Deeks SG (2002) Evolution of phenotypic drug susceptibility and viral replication capacity during long-term virologic failure of protease inhibitor therapy in human immunodeficiency virus-infected adults. J Virol 76:11104–11112
Boden D, Markowitz M (1998) Resistance to human immunodeficiency virus type 1 protease inhibitors. Antimicrob Agent Chemother 42:2775–2783
Clemente JC, Moose RE, Hemrajani R, Whitford LR, Govindasamy L, Reutzel R, McKenna R, Agbandje-McKenna M, Goodenow MM, Dunn BM (2004) Comparing the accumulation of active- and nonactive-site mutations in the HIV-1 protease. Biochemistry 43:12141–12151
Condra JH, Schleif WA, Blahy OM, Gabryelski LJ, Graham DJ, Quintero J, Rhodes A, Robbins HL, Roth E, Shivaprakash M, Titus D, Yang T, Tepplert H, Squires KE, Deutsch PJ, Emini EA (1995) In vivo emergence of HIV-1 variants resistant to multiple protease inhibitors. Nature 374:569–571
Darke PL, Huff JR (1994) HIV protease as an inhibitor target for the treatment of AIDS. Adv Pharmacol 25:399–455
De Gruttola V, Dix L, D’Aquila R, Holder D, Phillips A, Ait-Khaled M, Baxter J, Clevenbergh P, Hammer S, Harrigan R, Katzenstein D, Lanier R, Miller M, Para M, Yerly S, Zolopa A, Murray J, Patick A, Miller V, Castillo S, Pedneault L, Mellors J (2000) The relation between baseline HIV drug resistance and response to antiretroviral therapy: re-analysis of retrospective and prospective studies using a standardized data analysis plan. Antivir Ther 5:41–48
De Meyer S, Azijn H, Surleraux D, Jochmans D, Tahri A, Pauwels R, Wigerinck P, de Bethune MP (2005) TMC114, a novel human immunodeficiency virus type 1 protease inhibitor active against protease inhibitor-resistant viruses, including a broad range of clinical isolates. Antimicrob Agents Chemother 49:2314–2321
Debouck C, Navia MA, Fitzgerald PMD, McKeever BM, Leu CT, Heimbach JC, Herber WK, Sigal IS, Darke PL, Springer JP (1987) Human immunodeficiency virus protease expressed in Escherichia coli exhibits autoprocessing and specific maturation of the gag precursor. Proc Natl Acad Sci USA 84:8903–8906
Erickson JW, Gulnik SV, Markowitz M (1999) Protease inhibitors: resistance, cross-resistance, fitness and the choice of initial and salvage therapies. AIDS 13(Suppl A):S189–S204
Huff JR (1991) HIV protease: a novel chemotherapeutic target for AIDS. J Med Chem 34:2305–2314
Kaplan AH, Michael SF, Wehbie RS, Knigge MF, Paul DA, Everitt L, Kempf DJ, Norbeck DW, Erickson JW, Swanstrom R (1994) Selection of multiple human sensitivity to an inhibitor of the viral protease. Proc Natl Acad Sci USA 91:5597–5601
Koh Y, Nakata H, Maeda K, Ogata H, Bilcer G, Devasamudram T, Kincaid JF, Boross P, Wang YF, Tie Y, Volarath P, Gaddis L, Harrison RW, Weber IT, Ghosh AK, Mitsuya H (2003) Novel bis-tetrahydrofuranylurethane-containing nonpeptidic protease inhibitor (PI) UIC-94017 (TMC114) with potent activity against multi-PI-resistant human immunodeficiency virus in vitro. Antimicrob Agents Chemother 47:3123–3129
Kovalevsky AY, Tie Y, Liu F, Boross PI, Wang YF, Leshchenko S, Ghosh AK, Harrison RW, Weber IT (2006) Effectiveness of nonpeptide clinical inhibitor TMC-114 on HIV-1 protease with highly drug resistant mutations D30N, I50V, and L90M. J Med Chem 49:1379–1387
Ledergerber B, Egger M, Erard V, Weber R, Hirschel B, Furrer H, Battegay M, Vernazza P, Bernasconi E, Opravil M, Kaufmann D, Sudre P, Francioli P, Telenti A (1999) AIDS-related opportunistic illnesses occurring after initiation of potent antiretroviral therapy: the Swiss HIV Cohort Study. JAMA 282:2220–2226
Liu F, Boross PI, Wang YF, Tozser J, Louis JM, Harrison RW, Weber IT (2005) Kinetic, stability, and structural changes in high-resolution crystal structures of HIV-1 protease with drug-resistant mutations L24I, I50V, and G73S. J Mol Biol 354:789–800
Mahalingam B, Wang YF, Boross PI, Tozser J, Louis JM, Harrison RW, Weber IT (2004) Crystal structures of HIV protease V82A and L90M mutants reveal changes in the indinavir-binding site. Eur J Biochem 271:1516–1524
Mushi S, Chen Z, Yan Y, Li Y, Olsen DB, Schock HB, Galvin BB, Dorsey B, Kuo LC (2000) An alternate binding site for P1–P3 group of a class of potent HIV-1 protease inhibitors as a result of concerted structural change in the 80s loop of the protease. Acta Cryst D56:381–388
Muzammil S, Ross P, Freire E (2003) A major role for a set of non-active site mutations in the development of HIV-1 protease drug resistance. Biochemistry 42:631–638
Nunthanavanit P, Anthony NG, Johnston BF, Mackay SP, Ungwitayatorn J (2008) 3D-QSAR studies on chromone derivatives as HIV-1 protease inhibitors: application of molecular field analysis. Arch Pharm 341:357–364
Perez-Valero I, Arribas JR (2011) Protease inhibitor monotherapy. Curr Opin Infect Dis 24(1):7–11
Riva C, De Toma C, Donadel L, Boi C, Pennini R, Motta G, Leonardi A (1997) New DBU (1,8-diazabicyclo[5.4.0]undec-7-ene) assisted one-pot synthesis of 2,8-disubstituted 4H-1-benzopyran-4-ones. Synthesis 2:195–201
Shao W, Everitt L, Manchester M, Loeb DD, Hutchison CA, Swanstrom R (1997) Sequence requirements of the HIV-1 protease flab region determined by saturation mutagenesis and kinetic analysis of flab mutants. Proc Natl Acad Sci USA 94:2243–2248
Temesgen Z, Warnke D, Kasten MJ (2006) Current status of antiretroviral therapy. Expert Opin Pharmacother 7:1541–1554
Toh H, Ono M, Saigo K, Miyama T (1985) Retroviral protease-like sequence in the yeast transposon Ty 1. Nature 315:691
Ungwitayatorn J, Wiwat C, Samee W (2000) Synthesis and evaluation of chromone derivatives as potential HIV-1 protease inhibitors. Thai J Pharm Sci 24:155–156
Ungwitayatorn J, Samee W, Pimthon J (2004) 3D-QSAR studies on chromone derivatives as HIV-1 protease inhibitors. J Mol Struct 689:99–106
Ungwitayatorn J, Wiwat C, Samee W (2011) Synthesis, in vitro evaluation, and docking studies of novel chromone derivatives as HIV-1 protease inhibitor. J Mol Struct 1001:152–161
Wu TD, Schiffer CA, Gonzales MJ, Taylor J, Kantor R, Chou S, Israelski D, Zolopa AR, Fessel WJ, Shafer RW (2003) Mutation patterns and structural correlates in human immunodeficiency virus type 1 protease following different protease inhibitor treatments. J Virol 77:4836–4847
Acknowledgments
This project is supported by the Office of the High Education Commission and Mahidol University under the National Research Universities Initiative.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Nunthanavanit, P., Ungwitayatorn, J. Molecular docking studies of chromone derivatives against wild type and mutant strains of HIV-1 protease. Med Chem Res 23, 4198–4208 (2014). https://doi.org/10.1007/s00044-014-0992-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00044-014-0992-2