Abstract
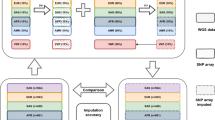
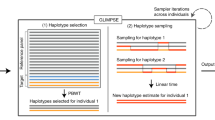
In recent genome-wide association studies, the task of genotype imputation for missing SNPs is a common procedure to increase the power of observed genetic markers. For genotype imputation, they usually employ publicly available resources, such as the International HapMap Project data or the 1000 Genome Project data, as a reference panel. However, lately, the volume of publicly available resources is rapidly increasing with the maturation of high-throughput genotyping technology. Thus, it often requires heavy computation for learning large reference panels, leading to long imputation time. In this work, to handle such problem, we propose a pruning strategy for the construction of imputation reference panels which is to reduce the size of reference panel data by excluding (or pruning) somewhat redundant samples from the reference panel based on the estimation of the kinship coefficients between samples. For evaluation, this approach was implemented under the Beagle framework and was tested on two real datasets, Mao et al.’s prostate cancer data and KNIH’s diabetes data. Our experiment results show that the proposed pruning strategy for reference panel construction can provide fast imputation time without the loss of imputation accuracy.
Similar content being viewed by others
References
Lewis, C.M. Genetic association studies: design, analysis and interpretation. Brief. Bioinform. 3, 146–153 (2002).
Tanaka, T. International HapMap project. Nihon Rinsho 63, 29–34 (2005).
Thorisson, G.A., Smith, A.V., Krishnan, L. & Stein, L.D. The international HapMap project web site. Genome Res. 15, 1592–1593 (2005).
Consortium Genomes Project. A map of human genome variation from population-scale sequencing. Nature 467, 1061–1073 (2010).
Howie, B., Marchini, J. & Stephens, M. Genotype imputation with thousands of genomes. G3 (Bethesda) 1, 457–470 (2011).
Huang, L. et al. The relationship between imputation error and statistical power in genetic association studies in diverse populations. Am. J. Hum. Genet. 85, 692–698 (2009).
Pasaniuc, B. et al. A generic coalescent-based framework for the selection of a reference panel for imputation. Genet. Epidemiol. 34, 773–782 (2010).
Jostins, L., Morley, K.I. & Barrett, J.C. Imputation of low-frequency variants using the HapMap3 benefits from large, diverse reference sets. Eur. J. Hum. Genet. 19, 662–666 (2011).
Sung, Y.J., Wang, L., Rankinen, T., Bouchard, C. & Rao, D.C. Performance of genotype imputations using data from the 1000 genomes project. Hum. Hered. 73, 18–25 (2012).
Huang, L. et al. Genotype-imputation accuracy across worldwide human populations. Am. J. Hum. Genet. 84, 235–250 (2009).
Mao, X. et al. Distinct genomic alterations in prostate cancers in Chinese and Western populations suggest alternative pathways of prostate carcinogenesis. Cancer Res. 70, 5207–5212 (2010).
Danford, T., Rolfe, A. & Gifford, D. GSE: a comprehensive database system for the representation, retrieval, and analysis of microarray data. Pac. Symp. Biocomput. 539–550 (2008).
Edgar, R., Domrachev, M. & Lash, A.E. Gene expression omnibus: NCBI gene expression and hybridization array data repository. Nucleic Acids Res. 30, 207–210 (2002).
Browning, B.L. & Browning, S.R. A unified approach to genotype imputation and haplotype-phase inference for large data sets of trios and unrelated individuals. Am. J. Hum. Genet. 84, 210–223 (2009).
Browning, S.R. Missing data imputation and haplotype phase inference for genome-wide association studies. Hum. Genet. 124, 439–450 (2008).
Browning, S.R. & Browning, B.L. Haplotype phasing: existing methods and new developments. Nat. Rev. Genet. 12, 703–714 (2011).
Sung, Y.J., Wang, L., Rankinen, T., Bouchard, C. & Rao, D.C. Performance of genotype imputations using data from the 1000 genomes project. Hum. Hered. 73, 18–25 (2012).
Manichaikul, A. et al. Robust relationship inference in genome-wide association studies. Bioinformatics 26, 2867–2873 (2010).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Jadamba, E., Shin, M., Chung, M. et al. A pruning strategy of reference panels for fast SNP genotype imputation. BioChip J 7, 6–10 (2013). https://doi.org/10.1007/s13206-013-7102-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13206-013-7102-2