Abstract
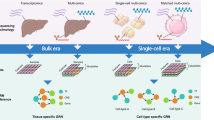
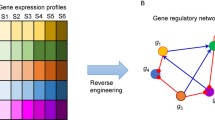
Our understanding of complex gene regulatory networks have been improved by the discovery of ncRNA-ncRNA crosstalk in normal and disease-specific physiological conditions. Previous studies have proposed numerous approaches for constructing ncRNA-ncRNA networks via ncRNA-mRNA regulation, functional information, or phenomics alone, or by combining heterogeneous data. Furthermore, it has been shown that ncRNA-ncRNA crosstalk can be rewired in different tissues or specific diseases. Therefore, it is necessary to integrate transcriptome data to construct context-specific ncRNA-ncRNA networks. In this chapter, we elucidated the commonly used ncRNA-ncRNA network modeling methods, and highlighted the need to integrate heterogeneous multi-mics data. Finally, we suggest future directions for studies of ncRNAs crosstalk. This comprehensive description and discussion elucidated in this chapter will provide constructive insights into ncRNA-ncRNA crosstalk.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Vella MC, Choi EY, Lin SY, Reinert K, Slack FJ (2004) The C. elegans microRNA let-7 binds to imperfect let-7 complementary sites from the lin-41 3′UTR. Genes Dev 18:132–137
Ambros V (2004) The functions of animal microRNAs. Nature 431:350–355
Krek A, Grun D, Poy MN, Wolf R, Rosenberg L, Epstein EJ, MacMenamin P, da Piedade I, Gunsalus KC, Stoffel M et al (2005) Combinatorial microRNA target predictions. Nat Genet 37:495–500
Li Y, Xu J, Chen H, Bai J, Li S, Zhao Z, Shao T, Jiang T, Ren H, Kang C et al (2013) Comprehensive analysis of the functional microRNA-mRNA regulatory network identifies miRNA signatures associated with glioma malignant progression. Nucleic Acids Res 41:e203
Grossi E, Sanchez Y, Huarte M (2016) Expanding the p53 regulatory network: LncRNAs take up the challenge. Biochim Biophys Acta 1859:200–208
Li Y, Chen J, Zhang J, Wang Z, Shao T, Jiang C, Xu J, Li X (2015) Construction and analysis of lncRNA-lncRNA synergistic networks to reveal clinically relevant lncRNAs in cancer. Oncotarget 6:25003–25016
Wu S, Huang S, Ding J, Zhao Y, Liang L, Liu T, Zhan R, He X (2010) Multiple microRNAs modulate p21Cip1/Waf1 expression by directly targeting its 3′ untranslated region. Oncogene 29:2302–2308
Peter ME (2010) Targeting of mRNAs by multiple miRNAs: the next step. Oncogene 29:2161–2164
Frampton AE, Castellano L, Colombo T, Giovannetti E, Krell J, Jacob J, Pellegrino L, Roca-Alonso L, Funel N, Gall TM et al (2014) MicroRNAs cooperatively inhibit a network of tumor suppressor genes to promote pancreatic tumor growth and progression. Gastroenterology 146(1):268–277, e218
Li J, Chen Z, Tian L, Zhou C, He MY, Gao Y, Wang S, Zhou F, Shi S, Feng X et al (2014) LncRNA profile study reveals a three-lncRNA signature associated with the survival of patients with oesophageal squamous cell carcinoma. Gut 63:1700–1710
Antonov AV, Dietmann S, Wong P, Lutter D, Mewes HW (2009) GeneSet2miRNA: finding the signature of cooperative miRNA activities in the gene lists. Nucleic Acids Res 37:W323–W328
Tsang JS, Ebert MS, van Oudenaarden A (2010) Genome-wide dissection of microRNA functions and cotargeting networks using gene set signatures. Mol Cell 38:140–153
Balaga O, Friedman Y, Linial M (2012) Toward a combinatorial nature of microRNA regulation in human cells. Nucleic Acids Res 40:9404–9416
Mal C, Deb A, Aftabuddin M, Kundu S (2015) A network analysis of miRNA mediated gene regulation of rice: crosstalk among biological processes. Mol BioSyst 11:2273–2280
Sun J, Zhou M, Yang H, Deng J, Wang L, Wang Q (2013) Inferring potential microRNA-microRNA associations based on targeting propensity and connectivity in the context of protein interaction network. PLoS One 8:e69719
Xu J, Li CX, Li YS, Lv JY, Ma Y, Shao TT, Xu LD, Wang YY, Du L, Zhang YP et al (2011) MiRNA-miRNA synergistic network: construction via co-regulating functional modules and disease miRNA topological features. Nucleic Acids Res 39:825–836
Wang D, Wang J, Lu M, Song F, Cui Q (2010) Inferring the human microRNA functional similarity and functional network based on microRNA-associated diseases. Bioinformatics 26:1644–1650
Hua L, Xia H, Zhou P, Li D, Li L (2014) Combination of microRNA expression profiling with genome-wide SNP genotyping to construct a coronary artery disease-related miRNA-miRNA synergistic network. Biosci Trends 8:297–307
Xiao S, Ma Y, Zhu H, Sun H, Yin Y, Feng G (2015) miRNA functional synergistic network analysis of mice with ischemic stroke. Neurol Sci Off J Ital Neurol Soc Ital Soc Clin Neurophysiol 36:143–148
Alshalalfa M (2012) MicroRNA response elements-mediated miRNA-miRNA interactions in prostate cancer. Adv Bioinforma 2012:839837
Na YJ, Kim JH (2013) Understanding cooperativity of microRNAs via microRNA association networks. BMC Genomics 14(Suppl 5):S17
Li Y, Liang C, Wong KC, Luo J, Zhang Z (2014) Mirsynergy: detecting synergistic miRNA regulatory modules by overlapping neighbourhood expansion. Bioinformatics 30:2627–2635
Yang Y, Xing Y, Liang C, Hu L, Xu F, Chen Y (2015) Crucial microRNAs and genes of human primary breast cancer explored by microRNA-mRNA integrated analysis. Tumour Biol J Int Soc Oncodev Biol Med 36:5571–5579
Xiao Y, Ping Y, Fan H, Xu C, Guan J, Zhao H, Li Y, Lv Y, Jin Y, Wang L et al (2013) Identifying dysfunctional miRNA-mRNA regulatory modules by inverse activation, cofunction, and high interconnection of target genes: a case study of glioblastoma. Neuro-Oncology 15:818–828
Song R, Catchpoole DR, Kennedy PJ, Li J (2015) Identification of lung cancer miRNA-miRNA co-regulation networks through a progressive data refining approach. J Theor Biol 380:271–279
Meng X, Wang J, Yuan C, Li X, Zhou Y, Hofestadt R, Chen M (2015) CancerNet: a database for decoding multilevel molecular interactions across diverse cancer types. Oncogene 4:e177
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2018 Springer Nature Singapore Pte Ltd.
About this chapter
Cite this chapter
Xu, J., Bai, J., Xiao, J. (2018). Computationally Modeling ncRNA-ncRNA Crosstalk. In: Li, X., Xu, J., Xiao, Y., Ning, S., Zhang, Y. (eds) Non-coding RNAs in Complex Diseases. Advances in Experimental Medicine and Biology, vol 1094. Springer, Singapore. https://doi.org/10.1007/978-981-13-0719-5_8
Download citation
DOI: https://doi.org/10.1007/978-981-13-0719-5_8
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-13-0718-8
Online ISBN: 978-981-13-0719-5
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)