Abstract
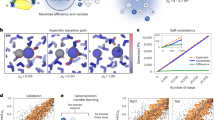
As has been shown recently, the identification of metastable chemical conformations leads to a Perron cluster eigenvalue problem for a reversible Markov operator. Naive discretization of this operator would suffer from combinatorial explosion. As a first remedy, a pre-identification of essential degrees of freedom out of the set of torsion angles had been applied up to now. The present paper suggests a different approach based on neural networks: its idea is to discretize the Markov operator via self-organizing box maps. The thus obtained box decomposition then serves as a prerequisite for the subsequent Perron cluster analysis. Moreover, this approach also permits exploitation of additional structure within embedded simulations. As it turns out, the new method is fully automatic and efficient also in the treatment of biomolecules. This is exemplified by numerical results.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Preview
Unable to display preview. Download preview PDF.
Similar content being viewed by others
References
R. Agrawal, J. Gehrke, D. Gunopulos, and P. Raghavan. Automatic subspace clustering of high dimensional data for data mining applications. In Proc. ACM SIGMOD Int. Conf. on Management of Data, pages 94–105, 1998.
A. Amadei, A.B.M. Linssen, and H.J.C. Berendsen. Essential dynamics of proteins. Proteins, 17, 1993.
P. Deuflhard, M. Dellnitz, O. Junge, and Ch. Schütte. Computation of essential molecular dynamics by subdivision techniques. In [4], pages 98–115.
P. Deuflhard, J. Hermans, B. Leimkuhler, A.E. Mark, S. Reich, and R.D. Skeel, editors, Computational Molecular Dynamics: Challenges, Methods, Ideas. Lecture Notes in Computational Science and Engineering, volume 4. Springer, 1998.
P. Deuflhard, W. Huisinga, A. Fischer, and Ch. Schütte. Identification of almost invariant aggregates in nearly uncoupled Markov chains. Linear Algebra and its Applications 315, pages 39–59, 2000.
B.S. Duran and P.L. Odell. Cluster Analysis. Springer, Berlin, 1974.
U.M. Fayyad, G. Patetsky-Shapiro, P. Smyth, and R. Uthurusamy (Eds.). Advances in Knowledge Discovery and Data Mining. AAAI Press/The MIT Press, California, 1996.
A. Fischer. An uncoupling-coupling technique for Markov Chain Monte Carlo methods. Preprint SC-00-04, Konrad-Zuse Zentrum, Berlin. Available via http://www.zib.de/.
A. Fischer, F. Cordes, and Ch. Schütte. Hybrid Monte Carlo with adaptive temperature in mixed-canonical ensemble: Efficient conformational analysis of RNA. J. Comp. Chem., 19(15):1689–1697, 1998.
A. Fischer, Ch. Schütte, P. Deuflhard, and F. Cordes. Hierarchical uncoupling-coupling of metastable conformations. ZIB Report 01-03, Konrad-Zuse-Zentrum, Berlin. Available via http://www.zib.de/bib/pub/pw.
N.I. Fisher. Statistical Analysis of Circular Data. University Press, Cambridge, 1993.
T. Galliat and P. Deuflhard. Adaptive hierarchical cluster analysis by self-organizing box maps. ZIB-Report 00-13 (April 2000), Konrad-Zuse-Zentrum, Berlin. Available via http://www.zib.de/DataMining, 2000.
T. Galliat, W. Huisinga, and P. Deuflhard. Self-organizing maps combined with eigenmode analysis for automated cluster identification. In H. Bothe and R. Rojas, editors, Proceedings of the 2nd International ICSC Symposium on Neural Computation, pages 227–232. ICSC Academic Press, 2000.
A. Gelman and D.B. Rubin. Inference from iterative simulation using multiple sequences. Statistical Science, 7:457–511, 1992.
A. Gersho and R.M. Gray. Vector Quantization and Signal Compression. Kluwer Academic Publishers, 1992.
T.A. Halgren. Merck molecular force field. i–v. J. Comp. Chem., 17(5&6):490–641, 1996.
M. Van Hulle. Faithful Representations and Topographic Maps. John Wiley Sons, Inc., 2000.
T. Kohonen. Comparison of SOM point densities based on different criteria. Neural Computation, (ll):2081–2095, 1999.
T. Kohonen. Self-Organizing Maps. Springer, Berlin, 3rd edition, 2001.
B.D. Ripley. Pattern Recognition and Neural Networks. Cambridge University Press, 1996.
Ch. Schütte. Conformational Dynamics: Modelling, Theory, Algorithm, and Application to Biomolecules. Habilitation Thesis, Dept. of Mathematics and Computer Science, Free University of Berlin, 1998. Available as ZIB-Report SC-99-18 via http://www.zib.de/bib/pub/pw/.
Ch. Schütte, A. Fischer, W. Huisinga, and P. Deuflhard. A direct approach to conformational dynamics based on hybrid Monte Carlo. J. Comput. Phys., Special Issue on Computational Biophysics, 151:146–168, 1999.
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2002 Springer-Verlag Berlin Heidelberg
About this paper
Cite this paper
Galliat, T., Deuflhard, P., Roitzsch, R., Cordes, F. (2002). Automatic Identification of Metastable Conformations via Self-Organized Neural Networks. In: Schlick, T., Gan, H.H. (eds) Computational Methods for Macromolecules: Challenges and Applications. Lecture Notes in Computational Science and Engineering, vol 24. Springer, Berlin, Heidelberg. https://doi.org/10.1007/978-3-642-56080-4_11
Download citation
DOI: https://doi.org/10.1007/978-3-642-56080-4_11
Publisher Name: Springer, Berlin, Heidelberg
Print ISBN: 978-3-540-43756-7
Online ISBN: 978-3-642-56080-4
eBook Packages: Springer Book Archive