Abstract
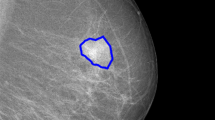
In this paper we present a Markov random field (MRF) driven region-based active contour model (MaRACel) for medical image segmentation. State-of-the-art region-based active contour (RAC) models assume that every spatial location in the image is statistically independent of the others, thereby ignoring valuable contextual information. To address this shortcoming we incorporate a MRF prior into the AC model, further generalizing Chan & Vese’s (CV) and Rousson and Deriche’s (RD) AC models. This incorporation requires a Markov prior that is consistent with the continuous variational framework characteristic of active contours; consequently, we introduce a continuous analogue to the discrete Potts model. To demonstrate the effectiveness of MaRACel, we compare its performance to those of the CV and RD AC models in the following scenarios: (1) the qualitative segmentation of a cancerous lesion in a breast DCE-MR image and (2) the qualitative and quantitative segmentations of prostatic acini (glands) in 200 histopathology images. Across the 200 prostate needle core biopsy histology images, MaRACel yielded an average sensitivity, specificity, and positive predictive value of 71%,95%,74% with respect to the segmented gland boundaries; the CV and RD models have corresponding values of 19%,81%,20% and 53%,88%,56%, respectively.
Chapter PDF
Similar content being viewed by others
Keywords
These keywords were added by machine and not by the authors. This process is experimental and the keywords may be updated as the learning algorithm improves.
References
Caselles, V., Kimmel, R., Sapiro, G.: Geodesic active contours. IJCV 22(1), 61–79 (1997)
Chan, T.F., Vese, L.A.: Active contours without edges. IEEE TIP 10(2), 266–277 (2001)
Sethian, J.A.: A fast marching level set method for monotonically advancing fronts. Proceedings of the National Academy of Sciences of the United States of America 93(4), 1591–1595 (1996)
Rousson, M., Deriche, R.: A variational framework for active and adaptative segmentation of vector valued images, pp. 56–61 (2002)
Zhu, S., Yuille, A.: Region competition: unifying snakes, region growing, and bayes/mdl for multiband image segmentation. IEEE TPAMI 18(9), 884–900 (1996)
Geman, S., Geman, D.: Stochastic relaxation, gibbs distributions, and the bayesian restoration of images. IEEE TPAMI PAMI-6(6), 721–741 (1984)
Agner, S., Soman, S., Libfeld, E., McDonald, M., Thomas, K., Englander, S., Rosen, M., Chin, D., Nosher, J., Madabhushi, A.: Textural kinetics: A novel dynamic contrast-enhanced (dce)-mri feature for breast lesion classification. Journal of Digital Imaging, May 28 (2010) [Epub. ahead of print]
Paragios, N., Deriche, R.: Geodesic active regions and level set methods for supervised texture segmentation. IJCV 46(3), 223–247 (2002)
Besag, J.: Statistical analysis of non-lattice data. Journal of the Royal Statistical Society, Series D (The Statistician) 24(3), 179–195 (1975)
Cremers, D., Mikael, R., Rachid, D.: A review of statistical approaches to level set segmentation: Integrating color, texture, motion and shape. IJCV 72, 195–215 (2007)
Monaco, J.P., Tomaszewski, J.E., Feldman, M.D., Hagemann, I., Moradi, M., Mousavi, P., Boag, A., Davidson, C., Abolmaesumi, P., Madabhushi, A.: High-throughput detection of prostate cancer in histological sections using probabilistic pairwise markov models. Medical Image Analysis 14(4), 617 (2010)
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2010 Springer-Verlag Berlin Heidelberg
About this paper
Cite this paper
Xu, J., Monaco, J.P., Madabhushi, A. (2010). Markov Random Field driven Region-Based Active Contour Model (MaRACel): Application to Medical Image Segmentation. In: Jiang, T., Navab, N., Pluim, J.P.W., Viergever, M.A. (eds) Medical Image Computing and Computer-Assisted Intervention – MICCAI 2010. MICCAI 2010. Lecture Notes in Computer Science, vol 6363. Springer, Berlin, Heidelberg. https://doi.org/10.1007/978-3-642-15711-0_25
Download citation
DOI: https://doi.org/10.1007/978-3-642-15711-0_25
Publisher Name: Springer, Berlin, Heidelberg
Print ISBN: 978-3-642-15710-3
Online ISBN: 978-3-642-15711-0
eBook Packages: Computer ScienceComputer Science (R0)