Abstract
In recent years, advancements in the knowledge of gut microbiota and its relationship with human pathophysiology led to the finding that gut microbiota is deeply implicated in the pathogenesis of liver steatosis and the progression to nonalcoholic steatohepatitis (NASH) and liver fibrosis.
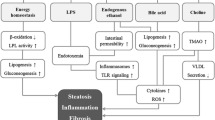
Gut microbiota composition and gene expression are independently linked to NAFLD, as well as to many known risk factors for NAFLD such as obesity and insulin resistance. Furthermore, gut microbiota composition and function are deeply affected by diet and lifestyle. Diets rich in saturated fats, fructose, and cholesterol alter the gut microbiota and intestinal barrier function, favoring the onset and progression of NAFLD. Gut dysbiosis may affect several metabolic pathways, both through direct and indirect mechanisms (e.g., production of short-chain fatty acids, interaction with bile acid metabolism, modulation of intestine-derived hormones), eventually leading to increased hepatic de novo lipogenesis and alterations in glucose and lipid metabolism that perpetuate liver damage. Gut dysbiosis also leads to increased intestinal permeability and translocation of microorganisms and microbial products into the portal circulation (metabolic endotoxemia), triggering pro-inflammatory processes in the liver.
Available strategies to manipulate gut microbiota include the use of probiotic, prebiotic, or synbiotic supplements, or antibiotic treatment. Fecal microbiota transplantation has emerged as a further potential tool to modify gut microbiota composition. Novel therapeutic strategies include targeting the nuclear farnesoid X receptor (FXR), a transcription factor involved in the regulation of bile, glucose, and lipid metabolism, and the use of absorbents that bind to toxic metabolites in the gut.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Betrapally NS, Gillevet PM, Bajaj JS. Changes in the intestinal microbiome and alcoholic and nonalcoholic liver diseases: causes or effects? Gastroenterology. 2016;150:1745–1755.e3.
Pedersen HK, et al. Human gut microbes impact host serum metabolome and insulin sensitivity. Nature. 2016;535:376–81.
Rabot S, et al. Germ-free C57BL/6J mice are resistant to high-fat-diet-induced insulin resistance and have altered cholesterol metabolism. FASEB J. 2010;24:4948–59.
Le Roy T, et al. Intestinal microbiota determines development of non-alcoholic fatty liver disease in mice. Gut. 2013;62:1787–94.
Loomba R, et al. Gut microbiome-based metagenomic signature for non-invasive detection of advanced fibrosis in human nonalcoholic fatty liver disease. Cell Metab. 2017;25:1054–1062.e5.
Boursier J, et al. The severity of nonalcoholic fatty liver disease is associated with gut dysbiosis and shift in the metabolic function of the gut microbiota. Hepatology. 2016;63:764–75.
de Wit NJW, Afman LA, Mensink M, Müller M. Phenotyping the effect of diet on non-alcoholic fatty liver disease. J Hepatol. 2012;57:1370–3.
Miele L, et al. Increased intestinal permeability and tight junction alterations in nonalcoholic fatty liver disease. Hepatology. 2009;49:1877–87.
Luther J, et al. Hepatic injury in nonalcoholic steatohepatitis contributes to altered intestinal permeability. Cell Mol Gastroenterol Hepatol. 2015;1:222–32.
Serino M, et al. Metabolic adaptation to a high-fat diet is associated with a change in the gut microbiota. Gut. 2012;61:543–53.
Wahlström A, Sayin SI, Marschall H-U, Bäckhed F. Intestinal crosstalk between bile acids and microbiota and its impact on host metabolism. Cell Metab. 2016;24:41–50.
Teodoro JS, Rolo AP, Palmeira CM. Hepatic FXR: key regulator of whole-body energy metabolism. Trends Endocrinol Metab. 2011;22:458–66.
Gadaleta RM, et al. Farnesoid X receptor activation inhibits inflammation and preserves the intestinal barrier in inflammatory bowel disease. Gut. 2011;60:463–72.
Midtvedt T. Microbial bile acid transformation. Am J Clin Nutr. 1974;27:1341–7.
Chu H, Williams B, Schnabl B. Gut microbiota, fatty liver disease, and hepatocellular carcinoma. Liver Res. 2018;2:43–51.
Mouzaki M, et al. Bile acids and dysbiosis in non-alcoholic fatty liver disease. PLoS One. 2016;11:e0151829.
Neuschwander-Tetri BA, et al. Farnesoid X nuclear receptor ligand obeticholic acid for non-cirrhotic, non-alcoholic steatohepatitis (FLINT): a multicentre, randomised, placebo-controlled trial. Lancet. 2015;385:956–65.
Parséus A, et al. Microbiota-induced obesity requires farnesoid X receptor. Gut. 2017;66:429–37.
Islam KBMS, et al. Bile acid is a host factor that regulates the composition of the cecal microbiota in rats. Gastroenterology. 2011;141:1773–81.
Burkitt DP, Walker AR, Painter NS. Dietary fiber and disease. JAMA. 1974;229:1068–74.
Valdes AM, Walter J, Segal E, Spector TD. Role of the gut microbiota in nutrition and health. BMJ. 2018;361:k2179.
Donohoe DR, et al. The microbiome and butyrate regulate energy metabolism and autophagy in the mammalian colon. Cell Metab. 2011;13:517–26.
Donohoe DR, Wali A, Brylawski BP, Bultman SJ. Microbial regulation of glucose metabolism and cell-cycle progression in mammalian colonocytes. PLoS One. 2012;7:e46589.
De Vadder F, et al. Microbiota-generated metabolites promote metabolic benefits via gut-brain neural circuits. Cell. 2014;156:84–96.
Thorburn AN, et al. Evidence that asthma is a developmental origin disease influenced by maternal diet and bacterial metabolites. Nat Commun. 2015;6:7320.
Davie JR. Inhibition of histone deacetylase activity by butyrate. J Nutr. 2003;133:2485S–93S.
Samuel BS, et al. Effects of the gut microbiota on host adiposity are modulated by the short-chain fatty-acid binding G protein-coupled receptor, Gpr41. Proc Natl Acad Sci U S A. 2008;105:16767–72.
Offermanns S. Free fatty acid (FFA) and hydroxy carboxylic acid (HCA) receptors. Annu Rev Pharmacol Toxicol. 2014;54:407–34.
Canfora EE, Meex RCR, Venema K, Blaak EE. Gut microbial metabolites in obesity, NAFLD and T2DM. Nat Rev Endocrinol. 2019;15:261–73.
Liu R, et al. Gut microbiome and serum metabolome alterations in obesity and after weight-loss intervention. Nat Med. 2017;23:859–68.
Dodd D, et al. A gut bacterial pathway metabolizes aromatic amino acids into nine circulating metabolites. Nature. 2017;551:648–52.
Obeid R, et al. Plasma trimethylamine N-oxide concentration is associated with choline, phospholipids, and methyl metabolism. Am J Clin Nutr. 2016;103(3):703–11. https://www.ncbi.nlm.nih.gov/pubmed/?term=10.3945%2Fajcn.115.121269.
Shan Z, et al. Association between microbiota-dependent metabolite trimethylamine-N-oxide and type 2 diabetes. Am J Clin Nutr. 2017;106(3):888–94. https://www.ncbi.nlm.nih.gov/pubmed/?term=10.3945%2Fajcn.117.157107.
Wilson Tang WH, et al. Intestinal microbial metabolism of phosphatidylcholine and cardiovascular risk. N Engl J Med. 2013;368(17):1575–84. https://www.ncbi.nlm.nih.gov/pubmed/?term=10.1056%2FNEJMoa1109400.
Pingitore A, et al. The diet-derived short chain fatty acid propionate improves beta-cell function in humans and stimulates insulin secretion from human islets in vitro. Diabetes Obes Metab. 2017;19:257–65.
Thomas C, et al. TGR5-mediated bile acid sensing controls glucose homeostasis. Cell Metab. 2009;10:167–77.
Watanabe M, et al. Bile acids induce energy expenditure by promoting intracellular thyroid hormone activation. Nature. 2006;439:484–9.
Chávez-Talavera O, Tailleux A, Lefebvre P, Staels B. Bile acid control of metabolism and inflammation in obesity, type 2 diabetes, dyslipidemia, and nonalcoholic fatty liver disease. Gastroenterology. 2017;152:1679–1694.e3.
Bäckhed F, et al. The gut microbiota as an environmental factor that regulates fat storage. Proc Natl Acad Sci U S A. 2004;101:15718–23.
Canfora EE, et al. Colonic infusions of short-chain fatty acid mixtures promote energy metabolism in overweight/obese men: a randomized crossover trial. Sci Rep. 2017;7:2360.
Fernandes J, Vogt J, Wolever TMS. Intravenous acetate elicits a greater free fatty acid rebound in normal than hyperinsulinaemic humans. Eur J Clin Nutr. 2012;66:1029–34.
Bouter K, et al. Differential metabolic effects of oral butyrate treatment in lean versus metabolic syndrome subjects. Clin Transl Gastroenterol. 2018;9:155.
Chambers ES, et al. Acute oral sodium propionate supplementation raises resting energy expenditure and lipid oxidation in fasted humans. Diabetes Obes Metab. 2018;20:1034–9.
Gancheva S, Jelenik T, Álvarez-Hernández E, Roden M. Interorgan metabolic crosstalk in human insulin resistance. Physiol Rev. 2018;98:1371–415.
David LA, et al. Diet rapidly and reproducibly alters the human gut microbiome. Nature. 2014;505:559–63.
De Filippo C, et al. Impact of diet in shaping gut microbiota revealed by a comparative study in children from Europe and rural Africa. Proc Natl Acad Sci U S A. 2010;107:14691–6.
Wu GD, et al. Linking long-term dietary patterns with gut microbial enterotypes. Science. 2011;334:105–8.
Cotillard A, et al. Dietary intervention impact on gut microbial gene richness. Nature. 2013;500:585–8.
Kovatcheva-Datchary P, et al. Dietary Fiber-induced improvement in glucose metabolism is associated with increased abundance of Prevotella. Cell Metab. 2015;22:971–82.
Walker AW, et al. Dominant and diet-responsive groups of bacteria within the human colonic microbiota. ISME J. 2011;5:220–30.
Ley RE, et al. Evolution of mammals and their gut microbes. Science. 2008;320:1647–51.
Muegge BD, et al. Diet drives convergence in gut microbiome functions across mammalian phylogeny and within humans. Science. 2011;332:970–4.
Schnorr SL, et al. Gut microbiome of the Hadza hunter-gatherers. Nat Commun. 2014;5:3654.
Yatsunenko T, et al. Human gut microbiome viewed across age and geography. Nature. 2012;486:222–7.
Obregon-Tito AJ, et al. Subsistence strategies in traditional societies distinguish gut microbiomes. Nat Commun. 2015;6:6505.
Clemente JC, et al. The microbiome of uncontacted Amerindians. Sci Adv. 2015;1:e1500183.
Koeth RA, et al. Intestinal microbiota metabolism of L-carnitine, a nutrient in red meat, promotes atherosclerosis. Nat Med. 2013;19:576–85.
Wu GD, et al. Comparative metabolomics in vegans and omnivores reveal constraints on diet-dependent gut microbiota metabolite production. Gut. 2016;65:63–72.
Gratz SW, et al. Dietary carbohydrate rather than protein intake drives colonic microbial fermentation during weight loss. Eur J Nutr. 2019;58:1147–58.
Devkota S, et al. Dietary-fat-induced taurocholic acid promotes pathobiont expansion and colitis in Il10−/− mice. Nature. 2012;487:104–8.
Cani PD, et al. Metabolic endotoxemia initiates obesity and insulin resistance. Diabetes. 2007;56:1761–72.
Ghoshal S, Witta J, Zhong J, de Villiers W, Eckhardt E. Chylomicrons promote intestinal absorption of lipopolysaccharides. J Lipid Res. 2009;50:90–7.
Shi H, et al. TLR4 links innate immunity and fatty acid-induced insulin resistance. J Clin Invest. 2006;116:3015–25.
Cani PD, et al. Changes in gut microbiota control metabolic endotoxemia-induced inflammation in high-fat diet-induced obesity and diabetes in mice. Diabetes. 2008;57:1470–81.
Caesar R, Tremaroli V, Kovatcheva-Datchary P, Cani PD, Bäckhed F. Crosstalk between gut microbiota and dietary lipids aggravates WAT inflammation through TLR signaling. Cell Metab. 2015;22:658–68.
van der Beek CM, et al. The prebiotic inulin improves substrate metabolism and promotes short-chain fatty acid production in overweight to obese men. Metab Clin Exp. 2018;87:25–35.
Rahat-Rozenbloom S, Fernandes J, Cheng J, Wolever TMS. Acute increases in serum colonic short-chain fatty acids elicited by inulin do not increase GLP-1 or PYY responses but may reduce ghrelin in lean and overweight humans. Eur J Clin Nutr. 2017;71:953–8.
Chassaing B, et al. Dietary emulsifiers impact the mouse gut microbiota promoting colitis and metabolic syndrome. Nature. 2015;519:92–6.
Suez J, et al. Artificial sweeteners induce glucose intolerance by altering the gut microbiota. Nature. 2014;514:181–6.
Zeevi D, et al. Personalized nutrition by prediction of glycemic responses. Cell. 2015;163:1079–95.
Sowah SA, et al. Effects of weight-loss interventions on short-chain fatty acid concentrations in blood and feces of adults: a systematic review. Adv Nutr. 2019;10:673–84.
Heianza Y, et al. Gut microbiota metabolites, amino acid metabolites and improvements in insulin sensitivity and glucose metabolism: the POUNDS lost trial. Gut. 2019;68:263–70.
Wang Z, et al. Impact of chronic dietary red meat, white meat, or non-meat protein on trimethylamine N-oxide metabolism and renal excretion in healthy men and women. Eur Heart J. 2019;40:583–94.
Martin AM, Sun EW, Rogers GB, Keating DJ. The influence of the gut microbiome on host metabolism through the regulation of gut hormone release. Front Physiol. 2019;10:428.
Holst JJ. The physiology of glucagon-like peptide 1. Physiol Rev. 2007;87:1409–39.
Christiansen CB, et al. The impact of short-chain fatty acids on GLP-1 and PYY secretion from the isolated perfused rat colon. Am J Physiol Gastrointest Liver Physiol. 2018;315:G53–65.
Tolhurst G, et al. Short-chain fatty acids stimulate glucagon-like peptide-1 secretion via the G-protein-coupled receptor FFAR2. Diabetes. 2012;61:364–71.
Yamaguchi Y, et al. Association of Intestinal Microbiota with metabolic markers and dietary habits in patients with type 2 diabetes. Digestion. 2016;94:66–72.
Batterham RL, et al. Gut hormone PYY(3-36) physiologically inhibits food intake. Nature. 2002;418:650–4.
Batterham RL, et al. Inhibition of food intake in obese subjects by peptide YY3-36. N Engl J Med. 2003;349:941–8.
Lin HV, et al. Butyrate and propionate protect against diet-induced obesity and regulate gut hormones via free fatty acid receptor 3-independent mechanisms. PLoS One. 2012;7:e35240.
Martin AM, et al. The diverse metabolic roles of peripheral serotonin. Endocrinology. 2017;158:1049–63.
Young RL, et al. Augmented capacity for peripheral serotonin release in human obesity. Int J Obes. 2018;42:1880–9.
Crane JD, et al. Inhibiting peripheral serotonin synthesis reduces obesity and metabolic dysfunction by promoting brown adipose tissue thermogenesis. Nat Med. 2015;21:166–72.
Yano JM, et al. Indigenous bacteria from the gut microbiota regulate host serotonin biosynthesis. Cell. 2015;161:264–76.
Le Chatelier E, et al. Richness of human gut microbiome correlates with metabolic markers. Nature. 2013;500:541–6.
Turnbaugh PJ, et al. A core gut microbiome in obese and lean twins. Nature. 2009;457:480–4.
Ridaura VK, et al. Gut microbiota from twins discordant for obesity modulate metabolism in mice. Science. 2013;341:1241214.
Haro C, et al. The gut microbial community in metabolic syndrome patients is modified by diet. J Nutr Biochem. 2016;27:27–31.
Santos-Marcos JA, Perez-Jimenez F, Camargo A. The role of diet and intestinal microbiota in the development of metabolic syndrome. J Nutr Biochem. 2019;70:1–27.
Derrien M, Vaughan EE, Plugge CM, de Vos WM. Akkermansia muciniphila gen. nov., sp. nov., a human intestinal mucin-degrading bacterium. Int J Syst Evol Microbiol. 2004;54:1469–76.
Dao MC, et al. Akkermansia muciniphila and improved metabolic health during a dietary intervention in obesity: relationship with gut microbiome richness and ecology. Gut. 2016;65:426–36.
Muñiz Pedrogo DA, et al. Gut microbial carbohydrate metabolism hinders weight loss in overweight adults undergoing lifestyle intervention with a volumetric diet. Mayo Clin Proc. 2018;93:1104–10.
Turnbaugh PJ, et al. An obesity-associated gut microbiome with increased capacity for energy harvest. Nature. 2006;444:1027–31.
Cho I, et al. Antibiotics in early life alter the murine colonic microbiome and adiposity. Nature. 2012;488:621–6.
Schwiertz A, et al. Microbiota and SCFA in lean and overweight healthy subjects. Obesity (Silver Spring). 2010;18:190–5.
Mouzaki M, Allard JP. The role of nutrients in the development, progression, and treatment of nonalcoholic fatty liver disease. J Clin Gastroenterol. 2012;46:457–67.
Zelber-Sagi S, et al. High red and processed meat consumption is associated with non-alcoholic fatty liver disease and insulin resistance. J Hepatol. 2018;68:1239–46.
Duncan SH, et al. Human colonic microbiota associated with diet, obesity and weight loss. Int J Obes. 2008;32:1720–4.
Fava F, et al. The type and quantity of dietary fat and carbohydrate alter faecal microbiome and short-chain fatty acid excretion in a metabolic syndrome ‘at-risk’ population. Int J Obes. 2013;37:216–23.
Pendyala S, Walker JM, Holt PR. A high-fat diet is associated with endotoxemia that originates from the gut. Gastroenterology. 2012;142:1100–1101.e2.
Zhou X, et al. A model of metabolic syndrome and related diseases with intestinal endotoxemia in rats fed a high fat and high sucrose diet. PLoS One. 2014;9:e115148.
Giorgio V, et al. Intestinal permeability is increased in children with non-alcoholic fatty liver disease, and correlates with liver disease severity. Dig Liver Dis. 2014;46:556–60.
Jin R, et al. Fructose induced endotoxemia in pediatric nonalcoholic fatty liver disease. Int J Hepatol. 2014;2014:560620.
Bischoff SC, et al. Intestinal permeability—a new target for disease prevention and therapy. BMC Gastroenterol. 2014;14:189.
Tripathi A, et al. The gut-liver axis and the intersection with the microbiome. Nat Rev Gastroenterol Hepatol. 2018;15:397–411.
Duarte SMB, Stefano JT, Oliveira CP. Microbiota and nonalcoholic fatty liver disease/nonalcoholic steatohepatitis (NAFLD/NASH). Ann Hepatol. 2019;18:416–21.
Zhu L, et al. Characterization of gut microbiomes in nonalcoholic steatohepatitis (NASH) patients: a connection between endogenous alcohol and NASH. Hepatology. 2013;57:601–9.
Kirpich IA, et al. The type of dietary fat modulates intestinal tight junction integrity, gut permeability, and hepatic toll-like receptor expression in a mouse model of alcoholic liver disease. Alcohol Clin Exp Res. 2012;36:835–46.
Liu J, Han L, Zhu L, Yu Y. Free fatty acids, not triglycerides, are associated with non-alcoholic liver injury progression in high fat diet induced obese rats. Lipids Health Dis. 2016;15:27.
Muccioli GG, et al. The endocannabinoid system links gut microbiota to adipogenesis. Mol Syst Biol. 2010;6:392.
Cani PD. Crosstalk between the gut microbiota and the endocannabinoid system: impact on the gut barrier function and the adipose tissue. Clin Microbiol Infect. 2012;18(Suppl 4):50–3.
Chen Y, et al. Associations of gut-flora-dependent metabolite trimethylamine-N-oxide, betaine and choline with non-alcoholic fatty liver disease in adults. Sci Rep. 2016;6:19076.
Del Chierico F, et al. Gut microbiota profiling of pediatric nonalcoholic fatty liver disease and obese patients unveiled by an integrated meta-omics-based approach. Hepatology. 2017;65:451–64.
Rinella ME, Tacke F, Sanyal AJ, Anstee QM, participants of the AASLD/EASL Workshop. Report on the AASLD/EASL joint workshop on clinical trial endpoints in NAFLD. J Hepatol. 2019;71:823–33. https://doi.org/10.1002/hep.30782.
European Association for the Study of the Liver (EASL), European Association for the Study of Diabetes (EASD) & European Association for the Study of Obesity (EASO). EASL–EASD–EASO Clinical Practice Guidelines for the management of non-alcoholic fatty liver disease. J Hepatol. 2016;64:1388–402.
Álvarez-Mercado AI, et al. Microbial population changes and their relationship with human health and disease. Microorganisms. 2019;7:68.
Baars A, Oosting A, Knol J, Garssen J, van Bergenhenegouwen J. The gut microbiota as a therapeutic target in IBD and metabolic disease: a role for the bile acid receptors FXR and TGR5. Microorganisms. 2015;3:641–66.
Parnell JA, Raman M, Rioux KP, Reimer RA. The potential role of prebiotic fibre for treatment and management of non-alcoholic fatty liver disease and associated obesity and insulin resistance. Liver Int. 2012;32:701–11.
Brandi G, et al. Microbiota, NASH, HCC and the potential role of probiotics. Carcinogenesis. 2017;38:231–40.
Rahman K, et al. Loss of junctional adhesion molecule A promotes severe steatohepatitis in mice on a diet high in saturated fat, fructose, and cholesterol. Gastroenterology. 2016;151:733–746.e12.
Famouri F, Shariat Z, Hashemipour M, Keikha M, Kelishadi R. Effects of probiotics on nonalcoholic fatty liver disease in obese children and adolescents. J Pediatr Gastroenterol Nutr. 2017;64:413–7.
Wong VW-S, et al. Treatment of nonalcoholic steatohepatitis with probiotics. A proof-of-concept study. Ann Hepatol. 2013;12:256–62.
Engstler AJ, et al. Insulin resistance alters hepatic ethanol metabolism: studies in mice and children with non-alcoholic fatty liver disease. Gut. 2016;65:1564–71.
Bakker GJ, Nieuwdorp M. Fecal microbiota transplantation: therapeutic potential for a multitude of diseases beyond Clostridium difficile. Microbiol Spectr. 2017;5.
Hu J, Luo H, Jiang Y, Chen P. Dietary capsaicin and antibiotics act synergistically to reduce non-alcoholic fatty liver disease induced by high fat diet in mice. Oncotarget. 2017;8:38161–75.
Gangarapu V, et al. Efficacy of rifaximin on circulating endotoxins and cytokines in patients with nonalcoholic fatty liver disease. Eur J Gastroenterol Hepatol. 2015;27:840–5.
Terveer EM, et al. Faecal microbiota transplantation in clinical practice. Gut. 2018;67:196.
Cammarota G, et al. European consensus conference on faecal microbiota transplantation in clinical practice. Gut. 2017;66:569–80.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2020 Springer Nature Switzerland AG
About this chapter
Cite this chapter
Miele, L. et al. (2020). Etiopathogenesis of NAFLD: Diet, Gut, and NASH. In: Bugianesi, E. (eds) Non-Alcoholic Fatty Liver Disease. Springer, Cham. https://doi.org/10.1007/978-3-319-95828-6_5
Download citation
DOI: https://doi.org/10.1007/978-3-319-95828-6_5
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-95827-9
Online ISBN: 978-3-319-95828-6
eBook Packages: MedicineMedicine (R0)