Abstract
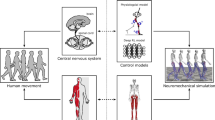
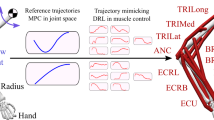
Motor control is a set of time-varying muscle excitations which generate desired motions for a biomechanical system. Muscle excitations cannot be directly measured from live subjects. An alternative approach is to estimate muscle activations using inverse motion-driven simulation. In this article, we propose a deep reinforcement learning method to estimate the muscle excitations in simulated biomechanical systems. Here, we introduce a custom-made reward function which incentivizes faster point-to-point tracking of target motion. Moreover, we deploy two new techniques, namely episode-based hard update and dual buffer experience replay, to avoid feedback training loops. The proposed method is tested in four simulated 2D and 3D environments with 6–24 axial muscles. The results show that the models were able to learn muscle excitations for given motions after nearly 100,000 simulated steps. Moreover, the root mean square error in point-to-point reaching of the target across experiments was less than 1% of the length of the domain of motion. Our reinforcement learning method is far from the conventional dynamic approaches as the muscle control is derived functionally by a set of distributed neurons. This can open paths for neural activity interpretation of this phenomenon.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Berniker M, Kording KP (2015) Deep networks for motor control functions. Front Comput Neurosci 9:35
Broad A (2011) Generating muscle driven arm movements using reinforcement learning. Master’s thesis, Washington University in St. Louis
Chollet F et al (2015) Keras. https://github.com/keras-team/keras
Erdemir A et al (2007) Model-based estimation of muscle forces exerted during movements. Clin Biomech 22(2):131–154
Gu S et al (2016) Continuous deep Q-learning with model-based acceleration. Cogn Process 12(4):319–340
Izawa J et al (2004) Biological arm motion through reinforcement learning. Biol Cybern 91(1):10–22
Khan N, Stavness I (2017) Prediction of muscle activations for reaching movements using deep neural networks. In: 41st Annual meeting of the American Society of Biomechanics, Boulder
Lloyd J et al (2012) ArtiSynth: a fast interactive biomechanical modeling toolkit combining multibody and finite element simulation. In: Payan Y (ed) Soft tissue biomechanical modeling for computer assisted surgery, vol 11, chap. 126. Springer, Berlin, pp 355–394
Mnih V et al (2013) Playing Atari with deep reinforcement learning. arXiv preprint arXiv:1312.5602
Pileicikiene G et al (2007) A three-dimensional model of the human masticatory system, including the mandible, the dentition and the temporomandibular joints. Stomatologija Balt Dent Maxillofac J 9(1):27–32
Plappert M (2016) keras-rl. https://github.com/matthiasplappert/keras-rl
Ravera EP et al (2016) Estimation of muscle forces in gait using a simulation of the electromyographic activity and numerical optimization. Comput Methods Biomech Biomed Engin 19(1):1–12
Sutton RS et al (1998) Reinforcement learning: an introduction. MIT Press, Cambridge
Van Hasselt H et al (2016) Deep reinforcement learning with double q-learning. In: Thirtieth AAAI conference on artificial intelligence, vol 16, pp 2094–2100
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2020 Springer Nature Switzerland AG
About this paper
Cite this paper
Abdi, A.H., Saha, P., Srungarapu, V.P., Fels, S. (2020). Muscle Excitation Estimation in Biomechanical Simulation Using NAF Reinforcement Learning. In: Nash, M., Nielsen, P., Wittek, A., Miller, K., Joldes, G. (eds) Computational Biomechanics for Medicine. Springer, Cham. https://doi.org/10.1007/978-3-030-15923-8_11
Download citation
DOI: https://doi.org/10.1007/978-3-030-15923-8_11
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-15922-1
Online ISBN: 978-3-030-15923-8
eBook Packages: EngineeringEngineering (R0)