Abstract
A mechanism of ion conduction of a voltage-gated potassium channel KcsA was investigated in full-atomic approximation at a trajectory length of 100 ns using the Lomonosov supercomputer. Methods of molecular dynamics were employed. A structure of the KcsA channel in the open state obtained by X-ray structure analysis (PDB ID 3fb7) was used. Free energy profiles of the KcsA pore occupied with either one or three potassium ions were calculated. It was shown that, under physiological conditions, ions pass through the channel pore cooperatively and the mechanism most probably includes three ions permeating in concert. Interactions of the mammalian voltage-gated channel Kv1.2 with neurotoxin were investigated. It was demonstrated that the effect of interionic interactions on binding of a blocker is rather insufficient.
Similar content being viewed by others
References
Molecular Dynamics Simulations of Potassium Channels, Centr. Eur. J. Chem., 2007, vol. 5, pp. 635–671.
Jensen, M., Borhani, D., Lindorff-Larsen, K., Maragakis, P., Jogini, V., Eastwood, M., Dror, R., and Shaw, D., Principles of Conduction and Hydrophobic Gating in K+ Channels, Proc. Natl. Acad. Sci. USA, 2010, vol. 107, pp. 5833–5838.
Berneche, S. and Roux, B., Energetics of Ion Conduction through the K+ Channel, Nature, 2001, vol. 414, pp. 73–77.
Khalili-Araghi, F., Tajkhorshid, E., and Schulten, K., Dynamics of K+ Ion Conduction through Kvl. 2, Biophys. J., 2006, vol. 91, pp. L72–L74.
Sokolova, O.S., Shaitan, K.V., Grizel’, A.V., Popinako, A.V., Karlova, M.G., and Kirpichnikov, M.P., Three-Dimensional Structure of Human VoltageGated Ion Channel Kv10.2 Studied by Electron Microscopy of Macromolecules and Molecular Modeling, Russ. J. Bioorg. Chem., 2012, vol. 38, no. 2, pp. 152–158.
Boiteux, C., Kraszewski, S., Ramseyer, C., Girarder, C., Ion Conductance Vs. Pore Gating and Selectivity in KcsA Channel: Modeling Achievements and Perspectives, J. Mol. Mod., 2007, vol. 13, pp. 699–713.
Cuello, L., Jogini, V., Cortes, M., and Perozo, E., Structural Mechanism of C-Type Inactivation in K+ Channels, Nature, 2010, vol. 466, pp. 203–208.
Smart, O., Neduvelil, J., Wang, X., and Sansom, M., HOLE: A Program for the Analysis of the Pore Dimensions of Ion Channel Structural Models, J. Mol. Graphics, 1996, vol. 14, pp. 354–360.
Duan, Y., Wu, C., Chowdhury, S., Lee, M., Xiong, G., Zhang, W., Yang, R., Cieplak, P., Luo, R., Lee, T., Caldwell, J., Wang, J., and Kollman, P., A PointCharge Force Field for Molecular Mechanics Simulations of Proteins Based on Condensed-Phase Quantum Mechanical Calculations, J. Comput. Chem., 2003, vol. 24, pp. 1999–2012.
Miloshevsky, G. and Jordan, P., Conformational Changes in the Selectivity Filter of the Open-State KcsA Channel: An Energy Minimization Study, Biophys. J., 2008, vol. 95, pp. 3239–3251.
Berger, O., Edholm, O., and Jahnig, F., Molecular Dynamics Simulations of a Fluid Bilayer of Dipalmitoylphosphatidylcholine at Full Hydration, Constant Pressure, and Constant Temperature, Biophys. J., 1997, vol. 72, pp. 2002–2013.
Wolf, M.G., Hoefling, M., Aponte-Santamaia, C., Grubmuller, H., and Groenhof, G., G-membed: Efficient Insertion of a Membrane Protein into an Equilibrated Lipid Bilayer with Minimal Perturbation, J. Comput. Chem., 2010, vol. 31, pp. 2169–2174.
Case, D., Gohlke, K., Luo, R., Merz, K., Onufriev, A., Simmerling, C., and Woods, R., The Amber Biomolecular Simulation Programs, J. Comput. Chem., 2005, vol. 26, pp. 1668–1688.
Phillips, J., Braun, R., Wang, W., Gumbart, J., Tajkhorshid, E., Villa, E., Chipot, C., Skeel, R., Kale, L., and Schulten, K., Scalable Molecular Dynamics with NAMD, J. Comput. Chem., 2005, vol. 26, pp. 1781–1802.
Karlova, M.G., Piscshalnikova, A.V., Ramonova, A.A., Moisenovich, M.M., Sokolova, O.S., and Shaitan, K.V., In vitro Fluorescence Assay to Study the Folding of Kv Ion Channels, Biophysics (Moscow), 2011, vol. 56, no. 2, pp. 272–279.
Long, S.B., Campbell, E.B., and Mackinnon, R., Crystal Structure of a Mammalian Voltage-Dependent Shaker Family K+ Channel, Science, 2005, vol. 309, pp. 897–903.
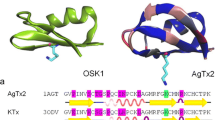
Krezel, A.M., Hidalgo, P., MacKinnon, R., and Wagner, G., Solution Structure of the Potassium Channel Inhibitor Agitoxin 2: Caliper for Probing Channel Geometry, Protein Sci., 1995, vol. 4, pp. 1478–1489.
Maestro, version 9.5, Schrödinger L.L.C., New York, 2007. http://www.schrodinger.corr
Eriksson, M.A. and Roux, B., Modeling the Structure of Agitoxin in Complex with the Shaker K+ Channel: A Computational Approach Based on Experimental Distance Restraints Extracted from Thermodynamic Mutant Cycles, Biophys. J., 2002, vol. 83, no. 5, pp. 2595–609.
Tieleman, D.P., Sansom, M.S.P., and Berendsen, H.J., Alamethicin Helices in a Bilayer and in Solution: Molecular Dynamics Simulations, Biophys. J., 1999, vol. 76, p. 40.
Humphrey, W., Dalke, A., and Schulten, K., VMD: Visual Molecular Dynamics, J. Mol. Graphics, 1996, vol. 14, pp. 33–38.
Dolinsky, T.J., Czodrowski, P., Li, H., Nielsen, J.E., Jensen, J.H., Klebe, G., and Baker, N.A., PDB2PQR: Expanding and Upgrading Automated Preparation of Biomolecular Structures for Molecular Simulations, Nucleic Acids Res., 2007, vol. 35, pp. W522–W525.
Chakrapani, S., Cordero-Morales, J., and Perozo, E., A Quantitative Description of KcsA Gating II: Single-Channel Currents, J. Gen. Physiol., 2007, vol. 130, pp. 479–496.
LeMasurier, M., Hegittbotham, L., and Miller, C., Kcsa. It’s a Potassium Channel, J. Gen. Physiol., 2001, vol. 118, pp. 303–314.
Grottesi, A. and Sansom, M.S., Molecular Dynamics Simulations of a K+ Channel Blocker: Tcl Toxin from Tityus cambridgei, FEBS Lett., 2003, vol. 535, nos. 1–3, pp. 29–33.
Author information
Authors and Affiliations
Corresponding author
Additional information
Original Russian Text © K.V. Shaitan, O.S. Sokolova, A.K. Shaitan, M.A. Kasimova, V.N. Novoseletskii, M.P. Kirpichnikov, 2013, published in Vestnik Moskovskogo Universiteta. Biologiya, 2013, No. 1, pp. 17–23.
About this article
Cite this article
Shaitan, K.V., Sokolova, O.S., Shaitan, A.K. et al. Influence of interionic interactions on functional state and blocker binding of voltage-gated potassium channels. Moscow Univ. Biol.Sci. Bull. 68, 8–14 (2013). https://doi.org/10.3103/S0096392513010057
Received:
Published:
Issue Date:
DOI: https://doi.org/10.3103/S0096392513010057