Abstract
Background
The purpose of this study was to develop an in situ immune marker model to predict postoperative oncological outcomes in patients with colorectal cancer (CRC).
Methods
Immunohistochemistry for 13 immune cell markers was performed on tumor tissue microarrays from 300 CRC patients who underwent curative resection from January 2000 to January 2006. Genetic algorithm was applied for the construction of an in situ immune marker model.
Results
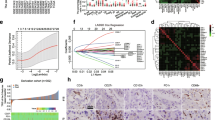
The infiltration of CD3+ cells, CD45RO+ cells, and FOXP3+ cells, but not the infiltration of Tryptase+ cells, in the tumor was significantly associated with better clinical outcome in overall survival (OS) and disease-free survival (DFS) of CRC patients, as assessed by univariate analysis (P < 0.05). Based on the genetic algorithms, a total of 6 markers, including CD3, CD45RO, IL17, CD15, Tryptase, and FOXP3, were selected to construct an immune marker model. Our model was identified to have an independent predictive capability for both OS and DFS in Cox multivariable model (P < 0.001). This was further confirmed by the ROC analysis (area under curve: OS, 0.669; DFS, 0.684).
Conclusions
The in situ immune marker model constructed in this study provides a novel approach to identify CRC patients who were at an increased risk for poor oncological outcomes.



Similar content being viewed by others
References
Jemal A, Bray F, Center MM, Ferlay J, Ward E, Forman D. Global cancer statistics. CA Cancer J Clin. 2011;61:69–90.
O’Connell JB, Maggard MA, Ko CY. Colon cancer survival rates with the new American Joint Committee on Cancer sixth edition staging. J Natl Cancer Inst. 2004; 96:1420–5.
Nitsche U, Maak M, Schuster T, et al. Prediction of prognosis is not improved by the seventh and latest edition of the TNM classification for colorectal cancer in a single-center collective. Ann Surg. 2011;254:793–800; discussion 800–1.
Zlobec I, Karamitopoulou E, Terracciano L, et al. TIA-1 cytotoxic granule-associated RNA binding protein improves the prognostic performance of CD8 in mismatch repair-proficient colorectal cancer. PLoS One. 2010;5:e14282.
Hanahan D, Weinberg RA. Hallmarks of cancer: the next generation. Cell. 2011;144:646–74.
Pages F, Berger A, Camus M, et al. Effector memory T cells, early metastasis, and survival in colorectal cancer. N Engl J Med. 2005;353:2654–66.
Garcia-Martinez E, Gil GL, Benito AC, et al. Tumor-infiltrating immune cell profiles and their change after neoadjuvant chemotherapy predict response and prognosis of breast cancer. Breast Cancer Res. 2014;16:488.
Schalper KA, Brown J, Carvajal-Hausdorf D, et al. Objective measurement and clinical significance of TILs in non-small cell lung cancer. J Natl Cancer Inst. 2015;107:dju435.
Fridman WH, Pages F, Sautes-Fridman C, Galon J. The immune contexture in human tumours: impact on clinical outcome. Nat Rev Cancer. 2012;12:298–306.
Bindea G, Mlecnik B, Fridman WH, Galon J. The prognostic impact of anti-cancer immune response: a novel classification of cancer patients. Semin Immunopathol. 2011;33:335–40.
Galon J, Pages F, Marincola FM, et al. The immune score as a new possible approach for the classification of cancer. J Transl Med. 2012;10:1.
Wu XR, He XS, Chen YF, et al. High expression of CD73 as a poor prognostic biomarker in human colorectal cancer. J Surg Oncol. 2012;106:130–7.
Faith DA, Isaacs WB, Morgan JD, et al. Trefoil factor 3 overexpression in prostatic carcinoma: prognostic importance using tissue microarrays. Prostate. 2004;61:215–27.
Camp RL, Dolled-Filhart M, Rimm DL. X-tile: a new bio-informatics tool for biomarker assessment and outcome-based cut-point optimization. Clin Cancer Res. 2004;10:7252–9.
Ooi CH, Tan P. Genetic algorithms applied to multi-class prediction for the analysis of gene expression data. Bioinformatics. 2003;19:37–44.
Rothberg BEG, Berger AJ, Molinaro AM, et al. Melanoma prognostic model using tissue microarrays and genetic algorithms. J Clin Oncol. 2009;27:5772–80.
Tejpar S, Bertagnolli M, Bosman F, et al. Prognostic and predictive biomarkers in resected colon cancer: current status and future perspectives for integrating genomics into biomarker discovery. Oncologist. 2010;15:390–404.
Cunningham D, Atkin W, Lenz HJ, et al. Colorectal cancer. Lancet. 2010;375:1030–47.
Mlecnik B, Bindea G, Angell HK, et al. Functional network pipeline reveals genetic determinants associated with in situ lymphocyte proliferation and survival of cancer patients. Sci Transl Med. 2014;6:228ra237.
Bindea G, Mlecnik B, Tosolini M, et al. Spatiotemporal dynamics of intratumoral immune cells reveal the immune landscape in human cancer. Immunity. 2013;39:782–95.
Mlecnik B, Bindea G, Pages F, Galon J. Tumor immunosurveillance in human cancers. Cancer Metastasis Rev. 2011;30:5–12.
Galon J, Fridman WH, Pages F. The adaptive immunologic microenvironment in colorectal cancer: a novel perspective. Cancer Res. 2007;67:1883–6.
Galon J, Costes A, Sanchez-Cabo F, et al. Type, density, and location of immune cells within human colorectal tumors predict clinical outcome. Science. 2006;313:1960–4.
Nosho K, Baba Y, Tanaka N, et al. Tumour-infiltrating T-cell subsets, molecular changes in colorectal cancer, and prognosis: cohort study and literature review. J Pathol. 2010;222:350–66.
Frey DM, Droeser RA, Viehl CT, et al. High frequency of tumor-infiltrating FOXP3(+) regulatory T cells predicts improved survival in mismatch repair-proficient colorectal cancer patients. Int J Cancer. 2010;126:2635–43.
Malfettone A, Silvestris N, Saponaro C, et al. High density of tryptase-positive mast cells in human colorectal cancer: a poor prognostic factor related to protease-activated receptor 2 expression. J Cell Mol Med. 2013;17:1025–37.
Mlecnik B, Tosolini M, Kirilovsky A, et al. Histopathologic-based prognostic factors of colorectal cancers are associated with the state of the local immune reaction. J Clin Oncol. 2011;29:610–8.
Hendriks Y, Franken P, Dierssen JW, et al. Conventional and tissue microarray immunohistochemical expression analysis of mismatch repair in hereditary colorectal tumors. Am J Pathol. 2003;162:469–77.
Xu J, Ding T, He Q, et al. An in situ molecular signature to predict early recurrence in hepatitis B virus-related hepatocellular carcinoma. J Hepatol. 2012;57:313–21.
Acknowledgments
We thank Dr. Xueqin Wang (Sun Yat-sen University) and Dr. Patrick Tan (Genome Institute of Singapore) for valuable insights in data analysis. This work was supported by National Key Clinical Discipline, National High Technology Research and Development Program (863) of China (No. 2012AA02A520), International S&T Cooperation Program of China (ISTCP) (No. 2013DFG32990), National Natural Science Foundation of China (NSFC) (No. 81400603), Science and Technology Program of Guangzhou, China (No. 2013J4100113), and Guangdong Department of Science & Technology Translational Medicine Center Grant (No. 2011A080300002).
Disclosure
All the authors declared that there’s no conflict of interest.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Yufeng Chen, Ruixue Yuan, and Xianrui Wu have contributed equally to this paper.
Electronic Supplementary Material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Chen, Y., Yuan, R., Wu, X. et al. A Novel Immune Marker Model Predicts Oncological Outcomes of Patients with Colorectal Cancer. Ann Surg Oncol 23, 826–832 (2016). https://doi.org/10.1245/s10434-015-4889-1
Received:
Published:
Issue Date:
DOI: https://doi.org/10.1245/s10434-015-4889-1