Abstract
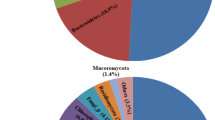
The composition of microbial communities of acid mine drainage (AMD) in two wells drilled in the terrace of the Sherlovaya Gora open-cast polymetallic ores mine (Eastern Siberia) was studied. While drainage water filling two wells, ShG14-1 and ShG14-8, had similar values of pH (2.6), Eh (447–494 mV), and temperature (6.5°C), the water in the first well contained more metals and sulfate. The water in ShG14-1 and ShG14-8 contained, respectively, 1898 and 434 mg/L of iron, 734 and 49 mg/L of manganese, 81 and 7 mg/L of copper, 3597 and 787 mg/L of zinc, and 15990 and 3632 mg/L of sulfate. Molecular analysis of the microbial communities was performed using pyrosequencing of the 16S rRNA gene fragments. The ShG14-8 microbial community included such bacterial taxa typically found in AMD sites as Gallionella (38.8% of total 16S rRNA gene sequences), Ferrovum (4.4%), Acidiphilium (9.1%), Acidisphaera (8.2%), Acidithiobacillus (7.2%), and Leptospirillum (4.6%). In the ShG14-1 sample with higher content of metals, strict acidophiles Acidithiobacillus (16.0%) and Leptospirillum (25.4%) were more abundant, while Gallionella, Ferrovum, Acidiphilium and Acidisphaera were almost absent. Ferrimicrobium (16.8%) and Sulfobacillus (1.4%) were detected in ShG14-1 but not in ShG14-8. Thus, the increase in concentration of metals in the acid mine drainage water under the same value of total acidity substantially altered the composition of the microbial community, preventing the development of “moderate” alpha- and beta-proteobacterial acidophiles, so that the community was dominated by the bacteria characteristic of the extremely acidic drainage waters.
Similar content being viewed by others
References
Anderson, I., Chertkov, O., Chen, A., Saunders, E., Lapidus, A., Nolan, M., Lucas, S., Hammon, N., Deshpande, S., Cheng, J.F., Han, C., Tapia, R., Goodwin, L.A., Pitluck, S., Liolios K., et al., Complete genome sequence of the moderately thermophilic mineral-sulfide-oxidizing firmicute Sulfobacillus acidophilus type strain (NAL(T)), Stand. Genomic Sci., 2012, vol. 6, pp. 1–13.
Bacelar-Nicolau, P. and Johnson, D.B., Leaching of pyrite by acidophilic heterotrophic iron-oxidizing bacteria in pure and mixed cultures, Appl. Environ. Microbiol., 1999, vol. 65, pp. 585–590.
Baker, B.J. and Banfield, J.F., Microbial communities in acid mine drainage, FEMS Microbiol. Ecol., 2003, vol. 44, pp. 139–152.
Baker, B.J., Comolli, L.R., Dick, G.J., Hauser, L.J., Hyatt, D., Dill, B.D., Land, M.L., Verberkmoes, N.C., Hettich, R.L., and Banfield, J.F., Enigmatic, ultrasmall, uncultivated archaea, Proc. Natl. Acad. Sci. U. S. A., 2010, vol. 107, pp. 8806–8811.
Banks, D., Karnachuk, O.V., Kadnikov, V.V., Watts, M., Boyce, A., Ivasenko, D.A., Filenko, R.A., Danilova, E.V., Pimenov, N.V., and Gundersen, P., Hydrochemical data report from sampling of polymetallic mines in Zabaikalskii kray, eastern Siberia, Russian Federation, Norges Geologiske Undersøkelse Report, 2014. 2014.035.
Behnke, A., Engel, M., Christen, R., Nebel, M., Klein, R.R., and Stoeck, T., Depicting more accurate pictures of protistan community complexity using pyrosequencing of hypervariable SSU rRNA gene regions, Environ. Microbiol., 2011, vol. 13, pp. 340–349.
Bruneel, O., Duran, R., Casiot, C., Elbaz-Poulichet, F., and Personne, J.C., Diversity of microorganisms in Fe-Asrich acid mine drainage waters of Carnoules, France, Appl. Environ. Microbiol., 2006, vol. 72, pp. 551–556.
Chen, L.X., Hu, M., Huang, L.N., Hua, Z.S., Kuang, J.L., Li, S.J., and Shu, W.S., Comparative metagenomic and metatranscriptomic analyses of microbial communities in acid mine drainage, ISME J., 2015, vol. 9, pp. 1579–1592.
Cole, J.R., Wang, Q., Cardenas, E., Fish, J., Chai, B., Farris, R.J., Kulam-Syed-Mohideen, A.S., McGarrell, D.M., Marsh, T., Garrity, G.M., and Tiedje J.M., The Ribosomal Database Project: improved alignments and new tools for rRNA analysis, Nucl. Acids Res., 2009, vol. 37, pp. 141–145.
Edgar, R.C., Haas, B.J., Clemente, J.C., Quince, C., and Knight, R., UCHIME improves sensitivity and speed of chimera detection, Bioinformatics, 2011, vol. 27, pp. 2194–2200.
Fabisch, M., Beulig, F., Akob, D.M., and Kusel, K., Surprising abundance of Gallionella-related iron oxidizers in creek sediments at pH 4.4 or at high heavy metal concentrations, Front. Microbiol., 2013, vol. 4, p. 390.
Hallberg, K.B., Coupland, K., Kimura, S., and Johnson, D.B., Macroscopic streamer growths in acidic, metal-rich mine waters in North Wales consist of novel and remarkably simple bacterial communities, Appl. Environ. Microbiol., 2006, vol. 72, pp. 2022–2030.
Hallberg, K.B., González-Toril, E., and Johnson, D.B., Acidithiobacillus ferrivorans,sp. nov.; facultatively anaerobic, psychrotolerant iron-,and sulfur-oxidizing acidophiles isolated from metal mine-impacted environments, Extremophiles, 2010, vol. 14, pp. 9–19.
He, Z., Xiao, S., Xie, X., Zhong, H., Hu, Y., Li, Q., Gao, F., Li, G., Liu, J., and Qiu, G., Molecular diversity of microbial community in acid mine drainages of Yunfu sulfide mine, Extremophiles, 2007, vol. 11, pp. 305–314.
Hiraishi, A., Matsuzawa, Y., Kanbe, T., and Wakao, N., Acidisphaera rubrifaciens gen. nov., sp. nov., an aerobic bacteriochlorophyll-containing bacterium isolated from acidic environments, Int. J. Syst. Evol. Microbiol., 2000, vol. 50, pp. 1539–1546.
Holanda, R., Hedrich, S., Falagán, C., Nancucheo, I., Dall’Agnol, H., Grail, B.M., and Johnson, D.B., Characteristics of Acidibacillus spp.: a novel genus of acidophilic iron-oxidising Firmicutes, Adv. Mater. Res., 2015, vol. 1130, pp. 36–39.
Johnson, D.B., Bacelar-Nicolau, P., Okibe, N., Thomas, A., and Hallberg, K.B., Ferrimicrobium acidiphilum gen. nov., sp. nov. and Ferrithrix thermotolerans gen. nov., sp. nov.: heterotrophic, iron-oxidizing, extremely acidophilic actinobacteria, Int. J. Syst. Evol. Microbiol., 2009, vol. 59, pp. 1082–1089.
Johnson, D.B., Hallberg, K.B., and Hedrich, S., Uncovering a microbial enigma: isolation and characterization of the streamer-generating, iron-oxidizing, acidophilic bacterium “Ferrovum myxofaciens,” Appl. Environ. Microbiol., 2014, vol. 80, pp. 672–680.
Kadnikov, V.V., Ivasenko, D.A., Beletsky, A.V., Mardanov, A.V., Danilova, E.V., Pimenov, N.V., Karnachuk, O.V., and Ravin, N.V., A novel uncultured bacterium of the family Gallionellaceae: description and genome reconstruction based on metagenomic analysis of microbial community in acid mine drainage, Microbiology (Moscow), 2016, vol. 85, no. 4, pp. 449–461.
Kimura, S., Bryan, C.G., Hallberg, K.B., and Johnson, D.B., Biodiversity and geochemistry of an extremely acidic, lowtemperature subterranean environment sustained by chemolithotrophy, Environ. Microbiol., 2011, vol. 13, pp. 2092–2104.
Kupka, D., Rzhepishevska, O.I., Dopson, M., Lindström, E.B., Karnachuk, O.V., and Tuovinen, O.H., Bacterial oxidation of ferrous sulfate at low temperatures, Biotechnol. Bioeng., 2007, vol. 97, pp. 1470–1478.
Küsel, K., Dorsch, T., Acker, G., and Stackebrandt, E., Microbial reduction of Fe(III) in acidic sediments: isolation of Acidiphilium cryptum JF-5 capable of coupling the reduction of Fe(III) to the oxidation of glucose, Environ. Microbiol., 1999, vol. 65, pp. 3633–3640.
Liljeqvist, M., Valdes, J., Holmes, D.S., and Dopson, M., Draft genome of the psychrotolerant acidophile Acidithiobacillus ferrivorans SS3, J. Bacteriol., 2011, vol. 193, pp. 4304–4305.
Méndez-García, C., Peláez, A.I, Mesa, V., Sánchez, J. Golyshina, O.V., and Ferrer, M., Microbial diversity and metabolic networks in acid mine drainage habitats, Front. Microbiol., 2015, vol. 29, p. 475.
Quast, C., Pruesse, E., Yilmaz, P., Gerken, J., Schweer, T., Yarza, P., Peplies, J., and Glöckner, F.O., The SILVA ribosomal RNA gene database project: improved data processing and web-based tools, Nucl. Acids Res., 2013, vol. 41 (D1), pp. D590–D596.
Rohwerder, T., Gehrke, T., Kinzler, K., and Sand, W., Bioleaching review part A: progress in bioleaching: fundamentals and mechanisms of bacterial metal sulfide oxidation, Appl. Microbiol. Biotechnol., 2003, vol. 63, pp. 239–248.
Schloss, P.D., Westcott, S.L., Ryabin, T., Hall, J.R., Hartmann, M., Hollister, E.B., Lesniewski, R.A., Oakley, B.B., Parks, D.H., Robinson, C.J., Sahl, J.W., Stres, B., Thallinger, G.G., Van Horn, D.J., and Weber, C.F., Introducing mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities, Appl. Environ. Microbiol., 2009, vol. 75, pp. 7537–7541.
Tyson, G.W., Chapman, J., Hugenholtz, P., Allen, E.E., Ram, R.J., Richardson, P.M., Solovyev, V.V., Rubin, E.M., Rokhsar, D.S., and Banfield, J.F., Community structure and metabolism through reconstruction of microbial genomes from the environment, Nature, 2004, vol. 428, no. 6978, pp. 37–43.
Author information
Authors and Affiliations
Corresponding author
Additional information
Original Russian Text © V.V. Kadnikov, D.A. Ivasenko, A.V. Beletsky, A.V. Mardanov, E.V. Danilova, N.V. Pimenov, O.V. Karnachuk, N.V. Ravin, 2016, published in Mikrobiologiya, 2016, Vol. 85, No. 6, pp. 732–739.
Rights and permissions
About this article
Cite this article
Kadnikov, V.V., Ivasenko, D.A., Beletsky, A.V. et al. Effect of metal concentration on the microbial community in acid mine drainage of a polysulfide ore deposit. Microbiology 85, 745–751 (2016). https://doi.org/10.1134/S0026261716060126
Received:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S0026261716060126