Abstract
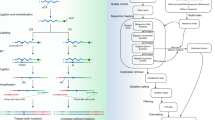
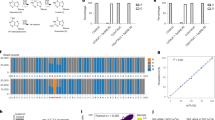
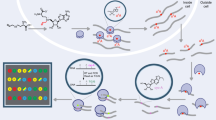
N6-methyladenosine (m6A) is the most abundant posttranscriptional chemical modification in mRNA, involved in regulating various physiological and pathological processes throughout mRNA metabolism. Recently, we developed GLORI, a sequencing method that enables the production of a globally absolute-quantitative m6A map at single-base resolution. Our technique utilizes the glyoxal- and nitrite-based chemical reaction, which selectively deaminates unmethylated adenosines while leaving m6A intact. The RNA library can then be prepared using a modified library construction protocol from enhanced UV crosslinking and immunoprecipitation (eCLIP) or commercial kits. Here we provide a detailed protocol for proper RNA sample handling and provide further guidelines for the use of a tailored bioinformatics pipeline (GLORI-tools) for subsequent data analysis. Compared with current methods, this new method is both exceptionally sensitive and robust, capable of identifying ~80,000 m6A sites with 50 Gb sequencing data in mammalian cells. It also provides a quantitative readout for m6A sites at single-base resolution. We hope the technique will provide a precise and unbiased tool for investigating m6A biology across various fields. Basic expertise in molecular biology and knowledge of bioinformatics are required for the protocol. The entire procedure can be completed within a week, with the library construction process taking ~4 d.
Key points
-
GLORI is a sequencing-based method for the characterization of N6-methyladenosine (m6A) methylome at single-base resolution. The procedure is based on the glyoxal- and nitrite-mediated deamination of unmethylated adenosine with high efficiency. A bioinformatic pipeline (GLORI-tool) is also complemented for optimized analysis of GLORI sequencing data.
-
Compared with other m6A detection methods, GLORI demonstrates exceptional sensitivity and precision, providing a more comprehensive and clearer transcriptome-wide m6A stoichiometric landscape.




Similar content being viewed by others
Data availability
The sequence data generated in this study have been deposited in the NCBI Gene Expression Omnibus, under accession code GSE233875.
Code availability
The code of GLORI-tools is freely available on GitHub (https://github.com/liucongcas/GLORI-tools).
References
Perry, R. P. & Kelley, D. E. Existence of methylated messenger RNA in mouse L cells. Cell 1, 37–42 (1974).
Perry, R. P., Kelley, D. E., Friderici, K. & Rottman, F. The methylated constituents of L cell messenger RNA: evidence for an unusual cluster at the 5′ terminus. Cell 4, 387–394 (1975).
Nachtergaele, S. & He, C. Chemical modifications in the life of an mRNA transcript. Annu. Rev. Genet. 52, 349–372 (2018).
Peer, E., Rechavi, G. & Dominissini, D. Epitranscriptomics: regulation of mRNA metabolism through modifications. Curr. Opin. Chem. Biol. 41, 93–98 (2017).
Wang, X. et al. N6-methyladenosine-dependent regulation of messenger RNA stability. Nature 505, 117–120 (2014).
Wang, X. et al. N6-methyladenosine modulates messenger RNA translation efficiency. Cell 161, 1388–1399 (2015).
Deng, X. et al. RNA N6-methyladenosine modification in cancers: current status and perspectives. Cell Res 28, 507–517 (2018).
Jiang, X. et al. The role of m6A modification in the biological functions and diseases. Signal Transduct. Target Ther. 6, 74 (2021).
Liu, J. et al. A METTL3–METTL14 complex mediates mammalian nuclear RNA N6-adenosine methylation. Nat. Chem. Biol. 10, 93–95 (2014).
Wang, X. et al. Structural basis of N6-adenosine methylation by the METTL3–METTL14 complex. Nature 534, 575–578 (2016).
Patil, D. P. et al. m6A RNA methylation promotes XIST-mediated transcriptional repression. Nature 537, 369–373 (2016).
Fu, Y., Dominissini, D., Rechavi, G. & He, C. Gene expression regulation mediated through reversible m6A RNA methylation. Nat. Rev. Genet. 15, 293–306 (2014).
Jia, G. et al. N6-Methyladenosine in nuclear RNA is a major substrate of the obesity-associated FTO. Nat. Chem. Biol. 7, 885–887 (2011).
Zheng, G. et al. ALKBH5 is a mammalian RNA demethylase that impacts RNA metabolism and mouse fertility. Mol. Cell 49, 18–29 (2013).
Yang, Y., Hsu, P. J., Chen, Y.-S. & Yang, Y.-G. Dynamic transcriptomic m6A decoration: writers, erasers, readers and functions in RNA metabolism. Cell Res 28, 616–624 (2018).
Xiao, W. et al. Nuclear m6A reader YTHDC1 regulates mRNA splicing. Mol. Cell 61, 507–519 (2016).
Dominissini, D. et al. Topology of the human and mouse m6A RNA methylomes revealed by m6A-seq. Nature 485, 201–206 (2012).
Meyer, KateD. et al. Comprehensive analysis of mRNA methylation reveals enrichment in 3′ UTRs and near stop codons. Cell 149, 1635–1646 (2012).
Linder, B. et al. Single-nucleotide-resolution mapping of m6A and m6Am throughout the transcriptome. Nat. Methods 12, 767–772 (2015).
Chen, K. et al. High-resolution N6-methyladenosine (m6A) map using photo-crosslinking-assisted m6A sequencing. Angew. Chem. Int. Ed. Engl. 54, 1587–1590 (2015).
Meyer, K. D. DART-seq: an antibody-free method for global m6A detection. Nat. Methods 16, 1275–1280 (2019).
Shu, X. et al. A metabolic labeling method detects m6A transcriptome-wide at single base resolution. Nat. Chem. Biol. 16, 887–895 (2020).
Wang, Y., Xiao, Y., Dong, S., Yu, Q. & Jia, G. Antibody-free enzyme-assisted chemical approach for detection of N6-methyladenosine. Nat. Chem. Biol. 16, 896–903 (2020).
Garcia-Campos, M. A. et al. Deciphering the “m6A code” via antibody-independent quantitative profiling. Cell 178, 731–747.e716 (2019).
Zhang, Z. et al. Single-base mapping of m6A by an antibody-independent method. Sci. Adv. 5, eaax0250 (2019).
Hu, L. et al. m6A RNA modifications are measured at single-base resolution across the mammalian transcriptome. Nat. Biotechnol. 40, 1210–1219 (2022).
Xiao, Y.-L. et al. Transcriptome-wide profiling and quantification of N6-methyladenosine by enzyme-assisted adenosine deamination. Nat. Biotechnol. 41, 993–1003 (2023).
Garalde, D. R. et al. Highly parallel direct RNA sequencing on an array of nanopores. Nat. Methods 15, 201–206 (2018).
Price, A. M. et al. Direct RNA sequencing reveals m6A modifications on adenovirus RNA are necessary for efficient splicing. Nat. Commun. 11, 6016 (2020).
Liu, H. et al. Accurate detection of m6A RNA modifications in native RNA sequences. Nat. Commun. 10, 4079 (2019).
Pratanwanich, P. N. et al. Identification of differential RNA modifications from nanopore direct RNA sequencing with xPore. Nat. Biotechnol. 39, 1394–1402 (2021).
Hendra, C. et al. Detection of m6A from direct RNA sequencing using a multiple instance learning framework. Nat. Methods 19, 1590–1598 (2022).
Vilfan, I. D. et al. Analysis of RNA base modification and structural rearrangement by single-molecule real-time detection of reverse transcription. J. Nanobiotechnol. 11, 8 (2013).
Liu, C. et al. Absolute quantification of single-base m6A methylation in the mammalian transcriptome using GLORI. Nat. Biotechnol. 41, 355–366 (2023).
Broude, N. E. & Budowsky, E. I. The reaction of glyoxal with nucleic acid components III. Kinetics of the reaction with monomers. Biochim. Biophys. Acta 254, 380–388 (1971).
Kivioja, T. et al. Counting absolute number of molecules using unique molecular identifiers. Nat. Methods 9, 72–74 (2011).
Islam, S. et al. Quantitative single-cell RNA-seq with unique molecular identifiers. Nat. Methods 11, 163–166 (2014).
Liu, N. et al. Probing N6-methyladenosine RNA modification status at single-nucleotide resolution in mRNA and long noncoding RNA. RNA 19, 1848–1856 (2013).
Zhang, Z. et al. Systematic calibration of epitranscriptomic maps using a synthetic modification-free RNA library. Nat. Methods 18, 1213–1222 (2021).
Sun, H. et al. m6Am-seq reveals the dynamic m6Am methylation in the human transcriptome. Nat. Commun. 12, 4778 (2021).
Shen, W., Le, S., Li, Y. & Hu, F. SeqKit: a cross-platform and ultrafast toolkit for FASTA/Q file manipulation. PLoS ONE 11, e0163962 (2016).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S. L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009).
Danecek, P. et al. Twelve years of SAMtools and BCFtools. GigaScience 10, giab008 (2021).
Acknowledgements
The authors thank the State Key Laboratory of Natural and Biomimetic Drugs at Peking University for their advice and technological support, the High-Performance Computing Platform of the Center for Life Science for their analysis assistance and the National Center for Protein Sciences at Peking University in Beijing, China for their aid in quantifying the library size distribution. This work was supported by the National Natural Science Foundation of China (nos. 22077006 to J.W., 91940304 to C.Y., 91853107 to J.W. and 21825701 to C.Y.) and the National Key R&D Program of China (nos. 2022YFC3400600 to J.W., 2019YFA0802201 to C.Y., 2017YFA0505202 to J.W., 2019YFA0110902 to C.Y. and 2020YFA0710401 to J.P.).
Author information
Authors and Affiliations
Contributions
The project was conceived and the experiments were designed by J.W. and C.Y. The manuscript was written and edited by C.Y., J.W. and W.S. with the help of C.L., H.S., Y. Hou. and Y. Hu. The chemical method of GLORI was developed by J.W. and Y.Y. Experiments were performed by W.S. and H.S., and sample preparation for next-generation sequencing (NGS) was designed and conducted by H.S. with the help of Y.X. and B.L. The bioinformatics pipeline of GLORI was developed by C.L., who also performed data analysis with the assistance of C.Y. Further optimization of the reaction condition was carried out by W.S. and H.S. The project was supervised by C.Y. and J.W.
Corresponding authors
Ethics declarations
Competing interests
Peking University has been granted a patent for GLORI.
Peer review
Peer review information
Nature Protocols thanks the anonymous reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Key reference using this protocol
Liu, C. et al. Nat Biotechnol. 41, 355–366 (2023): https://doi.org/10.1038/s41587-022-01487-9
Supplementary information
Supplementary Information
Supplementary Figs. 1 and 2.
Supplementary Tables 1–3
Sequences and primers.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Shen, W., Sun, H., Liu, C. et al. GLORI for absolute quantification of transcriptome-wide m6A at single-base resolution. Nat Protoc 19, 1252–1287 (2024). https://doi.org/10.1038/s41596-023-00937-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41596-023-00937-1
- Springer Nature Limited