Abstract
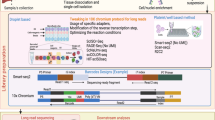
Somatic mutations accumulate in healthy tissues as we age, giving rise to cancer and potentially contributing to ageing. To study somatic mutations in non-neoplastic tissues, we developed a series of protocols to sequence the genomes of small populations of cells isolated from histological sections. Here, we describe a complete workflow that combines laser-capture microdissection (LCM) with low-input genome sequencing, while circumventing the use of whole-genome amplification (WGA). The protocol is subdivided broadly into four steps: tissue processing, LCM, low-input library generation and mutation calling and filtering. The tissue processing and LCM steps are provided as general guidelines that might require tailoring based on the specific requirements of the study at hand. Our protocol for low-input library generation uses enzymatic rather than acoustic fragmentation to generate WGA-free whole-genome libraries. Finally, the mutation calling and filtering strategy has been adapted from previously published protocols to account for artifacts introduced via library creation. To date, we have used this workflow to perform targeted and whole-genome sequencing of small populations of cells (typically 100–1,000 cells) in thousands of microbiopsies from a wide range of human tissues. The low-input DNA protocol is designed to be compatible with liquid handling platforms and make use of equipment and expertise standard to any core sequencing facility. However, obtaining low-input DNA material via LCM requires specialized equipment and expertise. The entire protocol from tissue reception through whole-genome library generation can be accomplished in as little as 1 week, although 2–3 weeks would be a more typical turnaround time.









Similar content being viewed by others
Data availability
Sequencing data referred to in this study have been deposited in the European Genome-phenome Archive with accession code EGAD00001006088.
Change history
27 February 2023
A Correction to this paper has been published: https://doi.org/10.1038/s41596-023-00816-9
References
Gawad, C., Koh, W. & Quake, S. R. Single-cell genome sequencing: current state of the science. Nat. Rev. Genet. 17, 175–188 (2016).
Blokzijl, F. et al. Tissue-specific mutation accumulation in human adult stem cells during life. Nature 538, 260–264 (2016).
Lodato, M. A. et al. Aging and neurodegeneration are associated with increased mutations in single human neurons. Science 359, 555–559 (2018).
Fatehullah, A., Tan, S. H. & Barker, N. Organoids as an in vitro model of human development and disease. Nat. Cell Biol. 18, 246–254 (2016).
Lee-Six, H. et al. The landscape of somatic mutation in normal colorectal epithelial cells. Nature 574, 532–537 (2019).
Brunner, S. F. et al. Somatic mutations and clonal dynamics in healthy and cirrhotic human liver. Nature 574, 538–542 (2019).
Moore, L. et al. The mutational landscape of normal human endometrial epithelium. Nature 580, 640–646 (2020).
Olafsson, S. et al. Somatic evolution in non-neoplastic IBD-affected colon. Cell 182, 672–684 (2020).
Telenius, H. et al. Degenerate oligonucleotide-primed PCR: general amplification of target DNA by a single degenerate primer. Genomics 13, 718–725 (1992).
Dean, F. B., Nelson, J. R., Giesler, T. L. & Lasken, R. S. Rapid amplification of plasmid and phage DNA using Phi29 DNA polymerase and multiply-primed rolling circle amplification. Genome Res 11, 1095–1099 (2001).
Zong, C., Lu, S., Chapman, A. R. & Xie, X. S. Genome-wide detection of single-nucleotide and copy-number variations of a single human cell. Science 338, 1622–1626 (2012).
Deleye, L. et al. Performance of four modern whole genome amplification methods for copy number variant detection in single cells. Sci. Rep. 7, 1–9 (2017).
Mathieson, W. et al. A critical evaluation of the PAXgene tissue fixation system morphology, immunohistochemistry, molecular biology, and proteomics. Am. J. Clin. Pathol. 146, 25–40 (2016).
Lee-Six, H. et al. Population dynamics of normal human blood inferred from somatic mutations. Nature 561, 473–478 (2018).
Chapman, M. S. et al. Lineage tracing of human embryonic development and foetal haematopoiesis through somatic mutations. Preprint at https://www.biorxiv.org/content/10.1101/2020.05.29.088765v1 (2020).
Yoshida, K. et al. Tobacco smoking and somatic mutations in human bronchial epithelium. Nature 578, 266–272 (2020).
Martincorena, I. et al. Somatic mutant clones colonize the human esophagus with age. Science 362, 911–917 (2018).
Martincorena, I. et al. Tumor evolution. High burden and pervasive positive selection of somatic mutations in normal human skin. Science 348, 880–886 (2015).
Picelli, S. et al. Full-length RNA-seq from single cells using Smart-seq2. Nat. Protoc. 9, 171–181 (2014).
Hoang, M. L. et al. Genome-wide quantification of rare somatic mutations in normal human tissues using massively parallel sequencing. Proc. Natl Acad. Sci. USA 113, 9846–9851 (2016).
Kennedy, S. R. et al. Detecting ultralow-frequency mutations by Duplex Sequencing. Nat. Protoc. 9, 2586–2606 (2014).
Wang, K., Li, M. & Hakonarson, H. ANNOVAR: functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res 38, e164–e164 (2010).
Li, H. & Durbin, R. Fast and accurate long-read alignment with Burrows–Wheeler transform. Bioinformatics 26, 589–595 (2010).
Jones, D. et al. cgpCaVEManWrapper: simple execution of CaVEMan in order to detect somatic single nucleotide variants in NGS data. Curr. Protoc. Bioinformatics 56, 15.10.1–15.10.18 (2016).
McKenna, A. et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res 20, 1297–1303 (2010).
Broad Institute. Picard Tools. http://broadinstitute.github.io/picard/.
Li, H. et al. The sequence alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Kozarewa, I. et al. Amplification-free Illumina sequencing-library preparation facilitates improved mapping and assembly of GC-biased genomes. Nat. Methods 6, 291–295 (2009).
Loo, P. V. et al. Allele-specific copy number analysis of tumors. Proc. Natl Acad. Sci. USA 107, 16910–16915 (2010).
Ye, K., Schulz, M. H., Long, Q., Apweiler, R. & Ning, Z. Pindel: a pattern growth approach to detect break points of large deletions and medium sized insertions from paired-end short reads. Bioinformatics 25, 2865–2871 (2009).
Acknowledgements
We acknowledge Y. Hooks and K. Roberts (Wellcome Sanger Institute) for assistance in tissue preparation and sectioning, P. Scott (Wellcome Sanger Institute) for advice and training for LCM, D. Rassl (Royal Papworth Hospital) for pathology support and R. Hoogenboezem (Erasmus University Medical Center) for assistance in software development. Finally, we would like to thank L. Apone, K. McKay, V. Panchapakesa and E. Dimalanta (NEB) for assistance in optimizing the library preparation process. This work was supported by the Wellcome Trust. S.F.B. was supported by the Swiss National Science Foundation (P2SKP3-171753 and P400PB-180790). T.M.B. was supported by a Cancer Research UK Grand Challenge Award (C98/A24032). M.A.S. was supported by a Rubicon Fellowship from the Netherlands Organisation for Scientific Research (019.153LW.038). L.M. is a recipient of a CRUK Clinical PhD Fellowship (C20/A20917) and a Pathological Society of Great Britain and Ireland Trainee Small Grant (grant reference no. 1175). I.M. is funded by Cancer Research UK (C57387/A21777). P.J.C. is a Wellcome Trust Senior Clinical Fellow (WT088340MA).
Author information
Authors and Affiliations
Contributions
P.E., L.M., M.A.S., T.M.B., A.R.J.L. and A.C. wrote the manuscript with contributions from all authors. P.E., L.M., R.O., B.F., S.F.B. and H.L.S. devised the protocol for laser-capture microscopy, DNA extraction and sequencing of microbiopsies. M.A.S. developed filters to remove fragmentase-associated artifacts. L.M. and T.M.B. performed LCM experiments, data curation and analysis. T.C. developed the ‘unmatched normal’ filtering strategy. A.R.J.L., M.R.S., I.M. and P.J.C. assisted with data analysis. P.J.C. supervised the study.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Protocols thanks Edwin Cuppen, Subhajyoti De and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Key references using this protocol
Brunner, S. et al. Nature 574, 538–542 (2019): https://doi.org/10.1038/s41586-019-1670-9
Lee-Six, H. et al. Nature 574, 532–537 (2019): https://doi.org/10.1038/s41586-019-1672-7
Moore, L. et al. Nature 580, 640–646 (2020): https://doi.org/10.1038/s41586-020-2214-z
Supplementary information
Supplementary Information
Supplementary Data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Ellis, P., Moore, L., Sanders, M.A. et al. Reliable detection of somatic mutations in solid tissues by laser-capture microdissection and low-input DNA sequencing. Nat Protoc 16, 841–871 (2021). https://doi.org/10.1038/s41596-020-00437-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41596-020-00437-6
- Springer Nature Limited
This article is cited by
-
Mapping cancer biology in space: applications and perspectives on spatial omics for oncology
Molecular Cancer (2024)
-
Tissue mosaicism following stem cell aging: blood as an exemplar
Nature Aging (2024)
-
Genetic variation across and within individuals
Nature Reviews Genetics (2024)
-
Reconstructing phylogenetic trees from genome-wide somatic mutations in clonal samples
Nature Protocols (2024)
-
Spatial architectures of somatic mutations in normal prostate, benign prostatic hyperplasia and coexisting prostate cancer
Experimental & Molecular Medicine (2024)