Abstract
During the first lineage segregation, a mammalian totipotent embryo differentiates into the inner cell mass (ICM) and trophectoderm (TE). However, how transcription factors (TFs) regulate this earliest cell-fate decision in vivo remains elusive, with their regulomes primarily inferred from cultured cells. Here, we investigated the TF regulomes during the first lineage specification in early mouse embryos, spanning the pre-initiation, initiation, commitment, and maintenance phases. Unexpectedly, we found that TFAP2C, a trophoblast regulator, bound and activated both early TE and inner cell mass (ICM) genes at the totipotent (two- to eight-cell) stages (‘bipotency activation’). Tfap2c deficiency caused downregulation of early ICM genes, including Nanog, Nr5a2, and Tdgf1, and early TE genes, including Tfeb and Itgb5, in eight-cell embryos. Transcription defects in both ICM and TE lineages were also found in blastocysts, accompanied by increased apoptosis and reduced cell numbers in ICMs. Upon trophoblast commitment, TFAP2C left early ICM genes but acquired binding to late TE genes in blastocysts, where it co-bound with CDX2, and later to extra-embryonic ectoderm (ExE) genes, where it cooperatively co-occupied with the former ICM regulator SOX2. Finally, ‘bipotency activation’ in totipotent embryos also applied to a pluripotency regulator NR5A2, which similarly bound and activated both ICM and TE lineage genes at the eight-cell stage. These data reveal a unique transcription circuity of totipotency underpinned by highly adaptable lineage regulators.







Similar content being viewed by others
Data availability
All datasets generated in this study have been deposited as a super series at the Gene Expression Omnibus (GEO) under accession number GSE216256. Accession codes of the published data in GEO used in this study are as follows: RNA-seq and ATAC-seq of the mouse early embryos, GSE66390; RNA-seq and ATAC-seq of epiblasts, GSE125318; TFAP2C ChIP of mouse TSCs, GSE28455; ATAC, H3K27ac ChIP–seq of mouse TSCs, GSE110950; scRNA of the mouse embryos, GSE45719; scRNA-seq of E4.5 epiblast and PrE, GSE159030; RNA-seq in the Tfap2 knockdown embryos, GSE124755. NR5A2 CUT&RUN and RNA-seq in the Nr5a2 knockdown embryos, GSE229740. Source data are provided with this paper.
References
Rossant, J. & Tam, P. P. Blastocyst lineage formation, early embryonic asymmetries and axis patterning in the mouse. Development 136, 701–713 (2009).
Rossant, J. Genetic control of early cell lineages in the mammalian embryo. Annu. Rev. Genet. 52, 185–201 (2018).
Zernicka-Goetz, M., Morris, S. & Bruce, A. Making a firm decision: multifaceted regulation of cell fate in the early mouse embryo. Nat. Rev. Genet. 10, 467–477 (2009).
Zhu, M. & Zernicka-Goetz, M. Principles of self-organization of the mammalian embryo. Cell 183, 1467–1478 (2020).
Rossant, J. Postimplantation development of blastomeres isolated from 4- and 8-cell mouse eggs. J. Embryol. Exp. Morphol. 36, 283–290 (1976).
Kelly, S. J. Studies of the developmental potential of 4- and 8-cell stage mouse blastomeres. J. Exp. Zool. 200, 365–376 (1977).
De Paepe, C., Krivega, M., Cauffman, G., Geens, M. & Van de Velde, H. Totipotency and lineage segregation in the human embryo. Mol. Hum. Reprod. 20, 599–618 (2014).
Deng, Q., Ramskold, D., Reinius, B. & Sandberg, R. Single-cell RNA-seq reveals dynamic, random monoallelic gene expression in mammalian cells. Science 343, 193–196 (2014).
Williams, T. & Tjian, R. Characterization of a dimerization motif in AP-2 and its function in heterologous DNA-binding proteins. Science 251, 1067–1071 (1991).
Cao, Z. et al. Transcription factor AP-2γ induces early Cdx2 expression and represses HIPPO signaling to specify the trophectoderm lineage. Development 142, 1606–1615 (2015).
Kuckenberg, P., Kubaczka, C. & Schorle, H. The role of transcription factor Tcfap2c/TFAP2C in trophectoderm development. Reprod. Biomed. Online 25, 12–20 (2012).
Winger, Q., Huang, J., Auman, H. J., Lewandoski, M. & Williams, T. Analysis of transcription factor AP-2 expression and function during mouse preimplantation development. Biol. Reprod. 75, 324–333 (2006).
Werling, U. & Schorle, H. Transcription factor gene AP-2γ essential for early murine development. Mol. Cell. Biol. 22, 3149–3156 (2002).
Auman, H. J. et al. Transcription factor AP-2γ is essential in the extra-embryonic lineages for early postimplantation development. Development 129, 2733–2747 (2002).
Kuckenberg, P. et al. The transcription factor TCFAP2C/AP-2γ cooperates with CDX2 to maintain trophectoderm formation. Mol. Cell. Biol. 30, 3310–3320 (2010).
Kidder, B. L. & Palmer, S. Examination of transcriptional networks reveals an important role for TCFAP2C, SMARCA4, and EOMES in trophoblast stem cell maintenance. Genome Res. 20, 458–472 (2010).
Zhu, M. et al. Developmental clock and mechanism of de novo polarization of the mouse embryo. Science 370, abd2703 (2020).
Wu, J. et al. The landscape of accessible chromatin in mammalian preimplantation embryos. Nature 534, 652–657 (2016).
Wu, J. et al. Chromatin analysis in human early development reveals epigenetic transition during ZGA. Nature 557, 256–260 (2018).
Pare, J. F. et al. The fetoprotein transcription factor (FTF) gene is essential to embryogenesis and cholesterol homeostasis and is regulated by a DR4 element. J. Biol. Chem. 279, 21206–21216 (2004).
Heng, J. C. et al. The nuclear receptor Nr5a2 can replace Oct4 in the reprogramming of murine somatic cells to pluripotent cells. Cell Stem Cell 6, 167–174 (2010).
Gu, P. et al. Orphan nuclear receptor LRH-1 is required to maintain Oct4 expression at the epiblast stage of embryonic development. Mol. Cell. Biol. 25, 3492–3505 (2005).
Festuccia, N., Owens, N., Chervova, A., Dubois, A. & Navarro, P. The combined action of Esrrb and Nr5a2 is essential for murine naive pluripotency. Development 148, dev199604 (2021).
Festuccia, N. et al. Nr5a2 is essential for morula development. Preprint at bioRxiv https://doi.org/10.1101/2023.01.16.524255 (2023).
Lai, F. et al. NR5A2 connects zygotic genome activation to the first lineage segregation in totipotent embryos. Cell Res. 33, 952–966 (2023).
Skene, P. J., Henikoff, J. G. & Henikoff, S. Targeted in situ genome-wide profiling with high efficiency for low cell numbers. Nat. Protoc. 13, 1006–1019 (2018).
Karasek, C., Ashry, M., Driscoll, C. S. & Knott, J. G. A tale of two cell-fates: role of the Hippo signaling pathway and transcription factors in early lineage formation in mouse preimplantation embryos. Mol. Hum. Reprod. 26, 653–664 (2020).
Skene, P. J. & Henikoff, S. An efficient targeted nuclease strategy for high-resolution mapping of DNA binding sites. eLife 6, e63274 (2017).
Pfeffer, P. L. & Pearton, D. J. Trophoblast development. Reproduction 143, 231–246 (2012).
Choi, I., Carey, T. S., Wilson, C. A. & Knott, J. G. Transcription factor AP-2γ is a core regulator of tight junction biogenesis and cavity formation during mouse early embryogenesis. Development 139, 4623–4632 (2012).
Xiang, Y. et al. Epigenomic analysis of gastrulation identifies a unique chromatin state for primed pluripotency. Nat. Genet. 52, 95–105 (2020).
Spitz, F. & Furlong, E. E. Transcription factors: from enhancer binding to developmental control. Nat. Rev. Genet. 13, 613–626 (2012).
Du, Z., Zhang, K. & Xie, W. Epigenetic reprogramming in early animal development. Cold Spring Harb. Perspect. Biol. 14, a039677 (2021).
Burton, A. et al. Heterochromatin establishment during early mammalian development is regulated by pericentromeric RNA and characterized by non-repressive H3K9me3. Nat. Cell Biol. 22, 767–778 (2020).
Fabian, D., Koppel, J. & Maddox-Hyttel, P. Apoptotic processes during mammalian preimplantation development. Theriogenology 64, 221–231 (2005).
Pampfer, S. Apoptosis in rodent peri-implantation embryos: differential susceptibility of inner cell mass and trophectoderm cell lineages—a review. Placenta 21, S3–S10 (2000).
Bedzhov, I. & Zernicka-Goetz, M. Cell death and morphogenesis during early mouse development: are they interconnected? Bioessays 37, 372–378 (2015).
Adiga, S. K. et al. p21 provides stage specific DNA damage control to preimplantation embryos. Oncogene 26, 6141–6149 (2007).
Cazzalini, O., Scovassi, A. I., Savio, M., Stivala, L. A. & Prosperi, E. Multiple roles of the cell cycle inhibitor p21(CDKN1A) in the DNA damage response. Mutat. Res. 704, 12–20 (2010).
Strumpf, D. et al. Cdx2 is required for correct cell fate specification and differentiation of trophectoderm in the mouse blastocyst. Development 132, 2093–2102 (2005).
Jedrusik, A., Cox, A., Wicher, K. B., Glover, D. M. & Zernicka-Goetz, M. Maternal-zygotic knockout reveals a critical role of Cdx2 in the morula to blastocyst transition. Dev. Biol. 398, 147–152 (2015).
Dietrich, J. E. & Hiiragi, T. Stochastic patterning in the mouse pre-implantation embryo. Development 134, 4219–4231 (2007).
Latos, P. A. et al. Elf5-centered transcription factor hub controls trophoblast stem cell self-renewal and differentiation through stoichiometry-sensitive shifts in target gene networks. Genes Dev. 29, 2435–2448 (2015).
Schorpp-Kistner, M., Wang, Z. Q., Angel, P. & Wagner, E. F. JunB is essential for mammalian placentation. EMBO J. 18, 934–948 (1999).
Schreiber, M. et al. Placental vascularisation requires the AP-1 component fra1. Development 127, 4937–4948 (2000).
Avilion, A. A. et al. Multipotent cell lineages in early mouse development depend on SOX2 function. Genes Dev. 17, 126–140 (2003).
Adachi, K. et al. Context-dependent wiring of Sox2 regulatory networks for self-renewal of embryonic and trophoblast stem cells. Mol. Cell 52, 380–392 (2013).
Guo, G. et al. Resolution of cell fate decisions revealed by single-cell gene expression analysis from zygote to blastocyst. Dev. Cell 18, 675–685 (2010).
Wicklow, E. et al. HIPPO pathway members restrict SOX2 to the inner cell mass where it promotes ICM fates in the mouse blastocyst. PLoS Genet. 10, e1004618 (2014).
Gassler, J. et al. Zygotic genome activation by the totipotency pioneer factor Nr5a2. Science 378, 1305–1315 (2022).
Latos, P. A. & Hemberger, M. From the stem of the placental tree: trophoblast stem cells and their progeny. Development 143, 3650–3660 (2016).
Thompson, J. J. et al. Extensive co-binding and rapid redistribution of NANOG and GATA6 during emergence of divergent lineages. Nat. Commun. 13, 4257 (2022).
Chronis, C. et al. Cooperative binding of transcription factors orchestrates reprogramming. Cell 168, 442–459 e20 (2017).
Cheung, T. H. & Rando, T. A. Molecular regulation of stem cell quiescence. Nat. Rev. Mol. Cell Biol. 14, 329–340 (2013).
Hemberger, M., Dean, W. & Reik, W. Epigenetic dynamics of stem cells and cell lineage commitment: digging Waddington’s canal. Nat. Rev. Mol. Cell Biol. 10, 526–537 (2009).
Blassberg, R. et al. Sox2 levels regulate the chromatin occupancy of WNT mediators in epiblast progenitors responsible for vertebrate body formation. Nat. Cell Biol. 24, 633–644 (2022).
Morgunova, E. & Taipale, J. Structural perspective of cooperative transcription factor binding. Curr. Opin. Struct. Biol. 47, 1–8 (2017).
Stirparo, G. G. et al. OCT4 induces embryonic pluripotency via STAT3 signaling and metabolic mechanisms. Proc. Natl Acad. Sci. USA 118, e2008890118 (2021).
McLean, C. Y. et al. GREAT improves functional interpretation of cis-regulatory regions. Nat. Biotechnol. 28, 495–501 (2010).
Solter, D. & Knowles, B. B. Immunosurgery of mouse blastocyst. Proc. Natl Acad. Sci. USA 72, 5099–5102 (1975).
Liu, D. et al. Primary specification of blastocyst trophectoderm by scRNA-seq: new insights into embryo implantation. Sci. Adv. 8, eabj3725 (2022).
Harrison, S. M., Dunwoodie, S. L., Arkell, R. M., Lehrach, H. & Beddington, R. S. Isolation of novel tissue-specific genes from cDNA libraries representing the individual tissue constituents of the gastrulating mouse embryo. Development 121, 2479–2489 (1995).
Zhang, Y. et al. Dynamic epigenomic landscapes during early lineage specification in mouse embryos. Nat. Genet. 50, 96–105 (2018).
Picelli, S. et al. Full-length RNA-seq from single cells using Smart-seq2. Nat. Protoc. 9, 171–181 (2014).
Trapnell, C. et al. Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. Nat. Protoc. 7, 562–578 (2012).
Karolchik, D. et al. The UCSC Table Browser data retrieval tool. Nucleic Acids Res. 32, D493–D496 (2004).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Zhang, Y. et al. Model-based analysis of ChIP–seq (MACS). Genome Biol. 9, R137 (2008).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell 38, 576–589 (2010).
Huang da, W., Sherman, B. T. & Lempicki, R. A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 4, 44–57 (2009).
Tu, S. et al. MAnorm2 for quantitatively comparing groups of ChIP–seq samples. Genome Res. 31, 131–145 (2021).
Scialdone, A. et al. Single-cell landscape of transcriptional heterogeneity and cell fate decisions during mouse early gastrulation. Cell Rep. 20, 1215–1228 (2017).
Acknowledgements
We are grateful to members of the Xie laboratory for the discussion and comments during the preparation of the manuscript, and the Animal Center and Biocomputing Facility at Tsinghua University for their support. We thank Q. Sun (Chinese Academic of Sciences) for providing the Stra8-cre mice. We thank J. Knott (Michigan State University) for insightful discussion. We are grateful to L. Yan, Y. Ren, and N. Wang from Third Hospital of Peking University for help with E4.5 TE collection. This work was funded by the National Key R&D Program of China (2021YFA1100102), the National Natural Science Foundation of China (31988101 and 31725018), the National Key R&D Program of China (2019YFA0508900), and the Tsinghua-Peking Center for Life Sciences. W.X. is a recipient of an HHMI International Research Scholar award and a New Cornerstone Investigator. The funders had no role in study design, data collection and analysis, decision to publish or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
L. Li, F.L., and W.X. conceived and designed the project. L. Li performed the CUT&RUN, ATAC-seq, and RNA-seq experiments. F.L. and L. Liu performed embryo collection and immunostaining with the help of X.L., Z.L., F.K., and Q.X. Q.F., F.L., and L. Li derived the Tfap2c mzKO mouse ES cells. L. Li performed the western blotting experiments. L. Li constructed the Tfap2c-knockout mouse ES cell line with the help from X.H. L. Li performed the bioinformatics analysis with the help from B.L. L. Li and W.X. prepared most figures and wrote the manuscript, with help from all authors. L. Li and F.L. contributed equally. L. Liu, X.L., X.H., and B.L. contributed equally.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Structural & Molecular Biology thanks Pierre Antoine Defossez, Peter Rugg-Gunn, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Dimitris Typas was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Validation of low-input TFAP2C CUT&RUN in TSCs and ESCs.
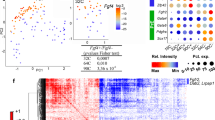
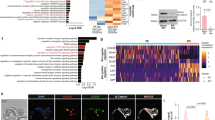
a, The boxplots showing the Spearman correlation between the expression of lineage regulator genes and ICM-TE specific genes across single cells at each stage8. Centre line, median; box, 25th and 75th percentiles; whiskers, 1.5 × interquartile range. Early ICM gene, n = 224; late ICM gene, n = 136; early TE genes, n = 191; late TE genes, n = 149; all genes, n = 22,654. b, Scatter plots comparing the expression of Tfap2c with representative lineage regulator genes across single cells in 8C, 16C embryos, and blastocysts8. The Spearman correlation coefficients are also shown. c, The UCSC browser views showing TFAP2C CUT&RUN using TSCs with cell numbers ranging from 50,000 to 500, and TFAP2C ChIP-seq47 signals in TSCs. d, Sequence logos and ranks of TF motif enrichment for TFAP2C peaks identified from TFAP2C CUT&RUN and ChIP-seq in TSCs. The top four TFs motifs are shown. e, The Western blot showing TFAP2C and GAPDH expression in wild-type (WT) and Tfap2c−/− mESCs. f, The UCSC browser views showing TFAP2C CUT&RUN using WT mESCs with cell numbers ranging from 2,000 to 500, and 2,000 Tfap2c−/− mESCs.
Extended Data Fig. 2 Validation of TFAP2C CUT&RUN in mouse embryos.
a, Immunostaining of TFAP2C (red) and DAPI (blue) in mouse 2-cell, 4-cell, 8-cell embryos, E3.5 and E4.5 blastocysts (two biological replicates). Dotted lines mark inner cells. Scale bar: 20 μm. b, Immunostaining showing TFAP2C, SOX2, and CDX2 protein levels and DAPI in E3.5 (n = 7) and E4.5 (n = 7) blastocysts. Scale bar: 20 μm. Quantification of TFAP2C signal intensity (relative to DAPI signal) in ICM/Epi cells (SOX2 positive or CDX2 negative) and TE cells (SOX2 negative or CDX2 positive) and P values (t-test, two-sided) are shown. Each dot represents a single blastomere. c, The UCSC browser views showing H3K27ac and H3K27me3 enrichment near Tfap2c, and heat maps showing the expression levels of Tfap2c. d, Scatter plots comparing the TFAP2C CUT&RUN signals (5-kb window for the entire genome) between two biological replicates. The Pearson correlation coefficients are also shown. e, Bar charts showing the numbers of strong peaks identified from TFAP2C CUT&RUN in 2C, 4C, and 8C embryos, E3.5 ICM, E4.5 TE, E6.5 ExE, and TSCs (two biological replicates at each stage). f, Bar charts showing the percentages of TFAP2C strong peak distribution in the promoter (TSS ± 2.5 kb), intragenic, and intergenic regions.
Extended Data Fig. 3 Characterization of TFAP2C binding dynamics in mouse early development.
a, Scatter plots showing pairwise correlation of TFAP2C CUT&RUN signals at its promoter or distal binding peaks across different stages and TSCs, for two replicates. Pearson correlation coefficients are also shown. b, Left, heat maps showing the TFAP2C binding, motif density, H3K27ac, ATAC-seq signals, and CpG density at stage-specific and shared TFAP2C binding peaks. Right, the functional enrichment of nearby genes for TFAP2C binding peaks by the GREAT analysis are shown. c, Bar charts showing the TFAP2C peak distribution in promoter (TSS ± 2.5 kb), intragenic and intergenic regions in each cluster.
Extended Data Fig. 4 TFAP2C binds both accessible and inaccessible regions at early stages.
a, Bar charts showing percentages of TFAP2C binding peaks overlapped with ATAC-seq peaks at the promoters and distal regions. Random peaks of the same length of each peak are shuffled and generated. b, The UCSC browser views and heat maps showing TFAP2C binding and ATAC-seq enrichment, and RNA expression of representative genes. TFAP2C binding in accessible regions and inaccessible regions is shaded. c, Scatter plots comparing the TFAP2C CUT&RUN signals (5-kb window for the entire genome) and ATAC-seq signals at each stage. The Pearson correlation coefficients are also shown. d, Immunofluorescence of TFAP2C (red) and DAPI (blue) in mouse control and Tfap2c KO 8-cell embryos (two biological replicates). Scale bar: 20 μm. e, Bright field views of control and Tfap2c KO embryos at E7.5 and E10.5 (two biological replicates). Scale bar: 200 μm. f, Control and Tfap2c mzKO blastocysts, and their outgrowths grown in culture for 6 days (three biological replicates). Scale bar: 50 μm. g, Morphology of 2i mESCs derived from control and Tfap2c KO blastocysts (three biological replicates). Scale bar: 100 μm. h, Western blot showing TFAP2C, NANOG, SOX2, OCT4, and TUBULIN proteins in control and Tfap2c KO mESCs (2 clones).
Extended Data Fig. 5 Transcriptome analyses of Tfap2c mzKO embryos.
a, Volcano plot showing the gene expression fold changes (Tfap2c mzKO vs. control embryos) and the P values from edgeR. 2, 2, 3, 5, 2, 5, and 5 biological replicates are used for 2C, 4C, 8C, E3.5 ICM, E4.5 TE, E6.5 Epi, and ExE, respectively. Up-regulated (log2(Fold change) > 2, P value < 0.01) and down-regulated (log2(Fold change) < 2, P value < 0.01) genes are colored in red and blue, respectively. The GO terms and example genes are also shown.
Extended Data Fig. 6 TFAP2C regulates both early ICM and early TE genes at the 8C stage.
a, The PCA analysis based on RNA-seq of wild-type embryos18, control and Tfap2c KO embryos. b, Venn diagram shows the overlap between down- or up-regulated genes in Tfap2c KO 8C embryos (three biological replicates) and TFAP2C target genes with inaccessible (left) or accessible (right) TFAP2C peaks at their promoters. P values, one-sided Fisher exact test. c, The cumulative distributions of down-regulated, up-regulated, and all genes with defined distances (x-axis) between their TSSs and nearest distal 8-cell TFAP2C accessible or inaccessible peaks, with P values (Wilcoxon test, two-sided) indicated. d, Left, the boxplots showing the average enrichment of TFAP2C signals at the promoters of various lineage genes at the 8C stage. Centre line, median; box, 25th and 75th percentiles; whiskers, 1.5 × interquartile range. Early ICM gene, n = 224; early TE genes, n = 191; PrE specific genes73, n = 979; Ectoderm (Ect) specific genes, n = 84; Mesoderm specific genes, n = 222; Endoderm (End) specific genes, n = 585; all genes, n = 22654. Right, the cumulative distributions of various lineage genes with defined distances (x-axis) between their TSSs and nearest distal TFAP2C binding peaks. e, The cumulative distributions of early-late ICM or TE genes with defined distances (x-axis) between their TSSs and nearest distal TFAP2C binding peaks. f, Box plots showing expression of representative genes in control and Tfap2c knockdown 8C embryos (four biological replicates)17, with P values (t-test, two-sided) indicated. g, The cumulative distributions of down-regulated or up-regulated ICM-specific, TE-specific, and other genes in Tfap2c KO 8C embryos with defined distances (x-axis) between their TSSs and nearest distal 8C TFAP2C binding peaks, with P values (Wilcoxon test, two-sided) indicated.
Extended Data Fig. 7 TFAP2C depletion activates cell apoptosis in ICMs.
a, Immunostaining showing NANOG protein levels and DAPI in control and Tfap2c mzKO E3.5 blastocysts (two biological replicates). Scale bar: 20 μm. b, Pie charts showing the percentages of ICM- and TE-specific genes among down-regulated genes in Tfap2c mzKO ICM and TE. c, The cumulative distributions of down-regulated or up-regulated, and all genes in Tfap2c KO ICM embryos with defined distances (x-axis) between their TSSs and nearest distal ICM TFAP2C binding peaks, with P values (Wilcoxon test, two-sided) indicated. d, Left, TUNEL assay and immunostaining showing CDX2 protein levels and DAPI in control and Tfap2c mzKO early blastocysts. Scale bar: 20 μm. Right, bar charts show the percentages of control (n = 12) and Tfap2c mzKO (n = 14) early blastocysts with different TUNEL foci incidence (CDX2 + , TE; CDX2-, ICM, indicated by dashed circles). e, Box plots showing expression of representative genes in control and Tfap2c mzKO embryos. f, TF motifs identified from 8C-specific (n = 2060), ICM-specific (n = 299), TE-specific (with CDX2 binding, n = 1227; without CDX2 binding, n = 843), and 8C-ICM-TE shared (with CDX2 binding, n = 1860; without CDX2 binding, n = 913) distal TFAP2C peaks. Sizes of circles indicate the relative motif enrichment. Expression levels of TFs in 8C embryos, ICM, and TE are color-coded for 8C-specific, ICM-specific, and TE-specific peaks, respectively. Expression of TFs is not shown for 8C-ICM-TE shared peaks. g, Line plots showing average motif densities for selected TFs (10 bp resolution) within 2 Kb around peak centers identified from TE-specific and 8C-ICM-TE shared distal TFAP2C peaks, with or without CDX2 binding in TE.
Extended Data Fig. 8 Co-localization of CDX2 and TFAP2C in TE.
a, Sequence logos and ranks of TF motifs enriched in CDX2 peaks (n = 19,620) identified from CDX2 CUT&RUN at E4.5 TE. The top five TFs motifs are shown. b, The UCSC browser views showing CDX2 binding and TFAP2C binding signals in E4.5 TE (two biological replicates). c, The UCSC browser views showing enrichment of TFAP2C binding, CDX2 binding in the control and Tfap2c mzKO TE. Shades indicate TE-specific TFAP2C binding site (red), 8C-ICM-TE shared TFAP2C binding site (green) and CDX2-only binding site (blue). The arrows indicate reduction of CDX2 binding at TE-specific TFAP2C peaks in TE. d, Left, heat maps showing TFAP2C binding, CDX2 binding at the 8C-specific, ICM-specific, TE-specific, and 8C-ICM-TE shared TFAP2C peaks, and TE CDX2-only peaks. Right, heat maps and average plots show enrichment of CDX2 CUT&RUN (replicate 2) signals in control and Tfap2c mzKO E4.5 TE. e, Immunostaining showing TFAP2C, CDX2 protein levels and DAPI in control and Tfap2c mzKO E4.5 blastocysts (three biological replicates). Scale bar: 20 μm.
Extended Data Fig. 9 Co-localization of SOX2 and TFAP2C in ExE.
a, The UCSC browser views showing SOX2 binding and TFAP2C binding signals in ExE (two biological replicates). b, Sequence logos and ranks of TF motifs enriched in SOX2 peaks (n = 69,743) identified from SOX2 CUT&RUN at E6.5 ExE. The top four TF motifs are shown. c, Scatterplot comparing global binding (5 Kb bin) of SOX2 and TFAP2C in ExE for two replicates. Pearson correlation coefficients are also shown. d, Heat maps showing enrichment of SOX2, TFAP2C binding signals, and motif densities (numbers of motif per bp) at SOX2-only, TFAP2C-only, and SOX2-TFAP2C co-bound peaks in ExE (left). The functional enrichment of nearby genes around binding peaks by the GREAT analysis is also shown (right). e, Box plots showing expression of Sox2 in control and Tfap2c mzKO E6.5 ExE (five biological replicates). f, Immunostaining showing SOX2, OCT4 protein levels and DAPI in control and Tfap2c mzKO E6.5 embryos (two biological replicates). Scale bar: 50 μm.
Extended Data Fig. 10 Comparisons of TFAP2C and NR5A2 binding.
a, The boxplots showing the average enrichment of NR5A2 signals at the promoters of early-late ICM or TE genes at each stage. P values, two-sided Wilcoxon rank-sum test. Centre line, median; box, 25th and 75th percentiles; whiskers, 1.5 × interquartile range (same for b). Early ICM gene, n = 224; late ICM gene, n = 136; early TE genes, n = 191; late TE genes, n = 149; all genes, n = 22654. b, Boxplots showing the average enrichment of NR5A2 binding signals25 at the promoter (in WT embryos) of down-regulated or up-regulated genes identified in Nr5a2 knockdown (KD) 8C embryos25 for ICM-specific, TE-specific, and other genes, with P values (t-test, two-sided) indicated. All genes were similarly analyzed and are shown as controls. Down ICM genes, n = 180; up ICM genes, n = 164; down TE genes, n = 168; up TE genes, n = 69; down other genes, n = 628; up other genes, n = 573. c, Left, Heat maps showing TFAP2C binding, NR5A2 binding25, ATAC-seq signals18, and motif density at TFAP2C-only, NR5A2-only, TFAP2C-NR5A2 shared binding peaks in 2C embryos, separated by promoter and distal regions. d, Heat maps showing TFAP2C binding, NR5A2 binding25, ATAC-seq signal, and motif density at TFAP2C-only, NR5A2-only, TFAP2C-NR5A2 shared binding peaks in 8C embryos, separated by promoter and distal regions. The percentages of open (+) or not-open (-) binding sites for each cluster are also shown. e, Venn diagram showing the overlap between down- (top) or up-regulated (bottom) genes in Tfap2c KO and Nr5a2 KD25 8C embryos. The functional enrichment of shared downregulated genes is also shown. P values, one-sided Fisher exact test.
Supplementary information
Supplementary Table 1
Tfap2c mzKO RNA expression table
Source data
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 1
Unprocessed western blots.
Source Data Extended Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 4
Unprocessed western blots.
Source Data Extended Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 7
Statistical source data.
Source Data Extended Data Fig. 10
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Li, L., Lai, F., Liu, L. et al. Lineage regulators TFAP2C and NR5A2 function as bipotency activators in totipotent embryos. Nat Struct Mol Biol (2024). https://doi.org/10.1038/s41594-023-01199-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41594-023-01199-x
- Springer Nature America, Inc.