Abstract
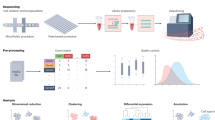
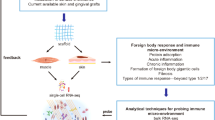
The understanding of the foreign-body responses to implanted biomaterials would benefit from the reconstruction of intracellular and intercellular signalling networks in the microenvironment surrounding the implant. Here, by leveraging single-cell RNA-sequencing data from 42,156 cells collected from the site of implantation of either polycaprolactone or an extracellular-matrix-derived scaffold in a mouse model of volumetric muscle loss, we report a computational analysis of intercellular signalling networks reconstructed from predictions of transcription-factor activation. We found that intercellular signalling networks can be clustered into modules associated with specific cell subsets, and that biomaterial-specific responses can be characterized by interactions between signalling modules for immune, fibroblast and tissue-specific cells. In a Il17ra–/– mouse model, we validated that predicted interleukin-17-linked transcriptional targets led to concomitant changes in gene expression. Moreover, we identified cell subsets that had not been implicated in the responses to implanted biomaterials. Single-cell atlases of the cellular responses to implanted biomaterials will facilitate the design of implantable biomaterials and the understanding of the ensuing cellular responses.





Similar content being viewed by others
Data availability
The raw and processed scRNA-seq data and bulk-sequencing data are available from the Gene Expression Omnibus (GEO) under the accession number GSE175890. All other data are available from the corresponding author on request.
Code availability
The Domino software is available at https://github.com/chris-cherry/domino.
References
Anderson, J. M., Rodriguez, A. & Chang, D. T. Foreign body reaction to biomaterials. Semin. Immunol. 20, 86–100 (2008).
Anderson, J. M. Biological responses to materials. Annu. Rev. Mater. Res. 31, 81–110 (2001).
Zakrzewski, J. L., van den Brink, M. R. M. & Hubbell, J. A. Overcoming immunological barriers in regenerative medicine. Nat. Biotechnol. 32, 786–794 (2014).
Zhang, B., Korolj, A., Lai, B. F. L. & Radisic, M. Advances in organ-on-a-chip engineering. Nat. Rev. Mater. 3, 257–278 (2018).
Chung, L. et al. Interleukin 17 and senescent cells regulate the foreign body response to synthetic material implants in mice and humans. Sci. Transl. Med. 12, eaax3799 (2020).
Sadtler, K. et al. Developing a pro-regenerative biomaterial scaffold microenvironment requires T helper 2 cells. Science 352, 366 (2016).
Papalexi, E. & Satija, R. Single-cell RNA sequencing to explore immune cell heterogeneity. Nat. Rev. Immunol. 18, 35–45 (2018).
Suvà, M. L. & Tirosh, I. Single-cell RNA sequencing in cancer: lessons learned and emerging challenges. Mol. Cell 75, 7–12 (2019).
Zhang, F. et al. Defining inflammatory cell states in rheumatoid arthritis joint synovial tissues by integrating single-cell transcriptomics and mass cytometry. Nat. Immunol. 20, 928–942 (2019).
Das, R. et al. Early B cell changes predict autoimmunity following combination immune checkpoint blockade. J. Clin. Investig. 128, 715–720 (2018).
Steuerman, Y. et al. Dissection of influenza infection in vivo by single-cell RNA sequencing. Cell Syst. 6, 679–691.e674 (2018).
Yao, C. et al. Single-cell RNA-seq reveals TOX as a key regulator of CD8+ T cell persistence in chronic infection. Nat. Immunol. 20, 890–901 (2019).
Efremova, M., Vento-Tormo, M., Teichmann, S. A. & Vento-Tormo, R. CellPhoneDB: inferring cell–cell communication from combined expression of multi-subunit ligand–receptor complexes. Nat. Protoc. 15, 1484–1506 (2020).
Noël, F. et al. Dissection of intercellular communication using the transcriptome-based framework ICELLNET. Nat. Commun. 12, 1089 (2021).
Wang, Y. et al. iTALK: an R package to characterize and illustrate intercellular communication. Preprint at bioRxiv https://doi.org/10.1101/507871 (2019).
Browaeys, R., Saelens, W. & Saeys, Y. NicheNet: modeling intercellular communication by linking ligands to target genes. Nat. Methods 17, 159–162 (2020).
Regev, A. et al. The Human Cell Atlas. eLife 6, e27041 (2017).
Schaum, N. et al. Single-cell transcriptomics of 20 mouse organs creates a Tabula Muris. Nature 562, 367–372 (2018).
Grubman, A. et al. A single-cell atlas of entorhinal cortex from individuals with Alzheimer’s disease reveals cell-type-specific gene expression regulation. Nat. Neurosci. 22, 2087–2097 (2019).
Sicari, B. M. et al. A murine model of volumetric muscle loss and a regenerative medicine approach for tissue replacement. Tissue Eng. A 18, 1941–1948 (2012).
Badylak, S. F. & Gilbert, T. W. Immune response to biologic scaffold materials. Semin. Immunol. 20, 109–116 (2008).
Sommerfeld, S. D. et al. Interleukin-36γ-producing macrophages drive IL-17-mediated fibrosis. Sci. Immunol. 4, eaax4783 (2019).
Kim, N. et al. Single-cell RNA sequencing demonstrates the molecular and cellular reprogramming of metastatic lung adenocarcinoma. Nat. Commun. 11, 2285 (2020).
Tibbitt, C. A. et al. Single-cell RNA sequencing of the T helper cell response to house dust mites defines a distinct gene expression signature in airway Th2 cells. Immunity 51, 169–184.e165 (2019).
Vallecillo-García, P. et al. Odd skipped-related 1 identifies a population of embryonic fibro-adipogenic progenitors regulating myogenesis during limb development. Nat. Commun. 8, 1218 (2017).
Ashcroft, G. S. et al. Tumor necrosis factor-alpha (TNF-α) is a therapeutic target for impaired cutaneous wound healing. Wound Repair Regen. 20, 38–49 (2012).
Gerarduzzi, C. & Di Battista, J. A. Myofibroblast repair mechanisms post-inflammatory response: a fibrotic perspective. Inflamm. Res. 66, 451–465 (2017).
Stojadinovic, O. et al. Molecular pathogenesis of chronic wounds: the role of β-catenin and c-myc in the inhibition of epithelialization and wound healing. Am. J. Pathol. 167, 59–69 (2005).
Cohen, M. et al. Lung single-cell signalling interaction map reveals basophil role in macrophage imprinting. Cell 175, 1031–1044.e1018 (2018).
Xie, X. et al. Single-cell transcriptome profiling reveals neutrophil heterogeneity in homeostasis and infection. Nat. Immunol. 21, 1119–1133 (2020).
Szczerba, B. M. et al. Neutrophils escort circulating tumour cells to enable cell cycle progression. Nature 566, 553–557 (2019).
Joanisse, S., Nederveen, J. P., Snijders, T., McKay, B. R. & Parise, G. Skeletal muscle regeneration, repair and remodelling in aging: the importance of muscle stem cells and vascularization. Gerontology 63, 91–100 (2017).
Aibar, S. et al. SCENIC: single-cell regulatory network inference and clustering. Nat. Methods 14, 1083–1086 (2017).
Yuk, J.-M. et al. Orphan nuclear receptor ERRα controls macrophage metabolic signalling and A20 expression to negatively regulate TLR-induced inflammation. Immunity 43, 80–91 (2015).
Heredia, J. E. et al. Type 2 innate signals stimulate fibro/adipogenic progenitors to facilitate muscle regeneration. Cell 153, 376–388 (2013).
Zhu, J. T helper 2 (Th2) cell differentiation, type 2 innate lymphoid cell (ILC2) development and regulation of interleukin-4 (IL-4) and IL-13 production. Cytokine 75, 14–24 (2015).
Liu, M. et al. Sox17 is required for endothelial regeneration following inflammation-induced vascular injury. Nat. Commun. 10, 2126 (2019).
Delgado-Olguín, P. et al. CTCF promotes muscle differentiation by modulating the activity of myogenic regulatory factors. J. Biol. Chem. 286, 12483–12494 (2011).
Liu, J. et al. Topical TWEAK accelerates healing of experimental burn wounds in mice. Front. Pharmacol. 9, 00660 (2018).
Wei, K. et al. Notch signalling drives synovial fibroblast identity and arthritis pathology. Nature 582, 259–264 (2020).
Bhattaram, P., Muschler, G., Wixler, V. & Lefebvre, V. Inflammatory cytokines stabilize SOXC transcription factors to mediate the transformation of fibroblast-like synoviocytes in arthritic disease. Arthritis Rheumatol. 70, 371–382 (2018).
Bashirova, A. A. et al. Diversity of the human LILRB3/A6 locus encoding a myeloid inhibitory and activating receptor pair. Immunogenetics 66, 1–8 (2014).
Anderson, A. J., Cummings, B. J. & Cotman, C. W. Increased immunoreactivity for Jun- and Fos-related proteins in Alzheimer’s disease: association with pathology. Exp. Neurol. 125, 286–295 (1994).
Chih-Chung, L. The Role of Bhlhe40 in Autoimmune Neuroinflammation and Mycobacterial Infection. PhD thesis, Washington Univ. (2017).
Zhu, S. & Qian, Y. IL-17/IL-17 receptor system in autoimmune disease: mechanisms and therapeutic potential. Clin. Sci. 122, 487–511 (2012).
Cristiano, C. et al. Neutralization of IL-17 rescues amyloid-β-induced neuroinflammation and memory impairment. Br. J. Pharmacol. 176, 3544–3557 (2019).
Zeng, F. et al. The relationship between single nucleotide polymorphisms of the NTRK2 gene and sporadic Alzheimer’s disease in the Chinese Han population. Neurosci. Lett. 550, 55–59 (2013).
Sakurai, K. & Osumi, N. The neurogenesis-controlling factor, Pax6, inhibits proliferation and promotes maturation in murine astrocytes. J. Neurosci. 28, 4604 (2008).
Takada, N., Kucenas, S. & Appel, B. Sox10 is necessary for oligodendrocyte survival following axon wrapping. Glia 58, 996–1006 (2010).
Aurora, A., Corona, B. T. & Walters, T. J. A porcine urinary bladder matrix does not recapitulate the spatiotemporal macrophage response of muscle regeneration after volumetric muscle loss injury. Cells Tissues Organs 202, 189–201 (2016).
Goldman, S. M. & Corona, B. T. Co-delivery of micronized urinary bladder matrix damps regenerative capacity of minced muscle grafts in the treatment of volumetric muscle loss injuries. PLoS ONE 12, e0186593 (2017).
Yao, Q. et al. Recent development and biomedical applications of decellularized extracellular matrix biomaterials. Mater. Sci. Eng. C. 104, 109942 (2019).
Jain, A. et al. Injectable formulations of poly(lactic acid) and its copolymers in clinical use. Adv. Drug Deliv. Rev. 107, 213–227 (2016).
Paige, J. T. et al. Modulation of inflammation in wounds of diabetic patients treated with porcine urinary bladder matrix. Regen. Med. 14, 269–277 (2019).
Burzyn, D. et al. A special population of regulatory T cells potentiates muscle repair. Cell 155, 1282–1295 (2013).
Gur-Cohen, S. et al. Stem cell–driven lymphatic remodeling coordinates tissue regeneration. Science 366, 1218 (2019).
Macosko, EvanZ. et al. Highly parallel genome-wide expression profiling of individual cells using nanoliter droplets. Cell 161, 1202–1214 (2015).
Satija, R., Farrell, J. A., Gennert, D., Schier, A. F. & Regev, A. Spatial reconstruction of single-cell gene expression data. Nat. Biotechnol. 33, 495–502 (2015).
Korsunsky, I. et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat. Methods 16, 1289–1296 (2019).
Tirosh, I. et al. Dissecting the multicellular ecosystem of metastatic melanoma by single-cell RNA-seq. Science 352, 189 (2016).
Durinck, S., Spellman, P. T., Birney, E. & Huber, W. Mapping identifiers for the integration of genomic datasets with the R/Bioconductor package biomaRt. Nat. Protoc. 4, 1184–1191 (2009).
Acknowledgements
We acknowledge financial support from the Morton Goldberg Chair, NIH Directors Pioneer Award, the Department of Defense, Bloomberg-Kimmel Institute for Cancer Immunotherapy and R01EB028796 awarded from the National Insitutes for Health (to J.H.E.). J.I.A. was supported by 5T32AG058527 awarded by the National Institute on Aging. L.X.G. is supported by grants K01ES025434 awarded by the National Institute of Environmental Health Sciences through funds provided by the trans-National Insitutes for Health Big Data to Knowledge (BD2K) initiative (https://commonfund.nih.gov/bd2k), R01 LM012373 and LM012907 awarded by the National Library of Medicine, and R01 HD084633 awarded by the National Institute of Child Health and Human Development. We thank D. Zack, C. Berlinicke and L. D. Huyer for help and expertise.
Author information
Authors and Affiliations
Contributions
C.C. and J.H.E. conceptualized and drafted the figures and manuscript, contributed to experimental design and interpreted findings. C.C., D.R.M., J.H., J.I.A. and J.H.E. performed experiments and analysed experimental results. C.C. wrote software. C.C., J.H.E., P.C., L.X.G. and E.J.F. contributed to computational methodology. All authors participated in construction of the manuscript and figures.
Corresponding author
Ethics declarations
Competing interests
C.C. is the founder and owner of C M Cherry Consulting, LLC. E.J.F. is a member of the scientific advisory board for Viosera Therapeutics.
Additional information
Peer review information Nature Biomedical Engineering thanks Kai Kessenbrock and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Experimental overview of the assembled datasets.
a, All data sets were taken from mice after volumetric muscle loss treatment. After surgical excision of a large portion of the quadriceps, the wound site was filled with a biomaterial or saline control and stapled shut. Mice were then harvested 1 or 6 weeks after surgery. Young (6 week) or aged (104 week) old animals were used. Representative histological images of PCL and ECM treated muscles 6 week after injury are shown stained by Masson’s Trichrome. Muscle fibers are stained red and connective tissue is stained blue. The posterior (P) is labeled and the location of original defects circled. b, At time of harvest, cells were isolated one of three ways after digestions. For macrophages, cells were sorted as CD45+F4/80HiLy6c+, for fibroblasts cells were sorted as CD45-CD19-CD29+, and for the all-cell dataset CD45+ cells were enriched to ~50% using MACS beads. c, Data sets were integrated for analysis using Harmony. A complete summary of available data sets is given in Supplementary Table 2. d, Enrichment of fibroblasts and macrophages due to inclusion of sorted fibroblast and macrophage data sets. The sorted fibroblasts (left) and macrophages (right) are shown in comparison to the CD45+ enriched sample (middle). E, Cells by condition. Cells are colored by condition plotted on UMAP dimensions. Cells were plotted in order of ECM, PCL, Saline, and Naïve.
Extended Data Fig. 2 Expression characteristics of the CD45+ cluster, and comparison with flow cytometry.
a, Gene expression for myeloid cluster markers. Single cell gene expression for marker genes used to identify myeloid clusters in tandem with CD14, CD11b, and F4/80 expression are shown as violin plots. b, Comparison of flow cytometry cell proportions to single cell proportions. Myeloid (CD45+CD11b+) and T cell (CD45+CD3+) numbers by flow cytometry given as raw counts (top left) and percent of the CD45+ population (top right) from ECM, PCL, or saline treated animals. Mean values are plotted with standard error shown on error bars. CD45- and CD45+ count values are shown, which were used to project single cell proportions to predicted raw numbers. Project single cell counts are shown by cluster (middle) by multipltying raw CD45+ and CD45- counts from flow cytometry with cluster proportions of CD45+ or CD45- populations from single cell (bottom).
Extended Data Fig. 3 Subset clustering of T/NK cells.
a, Clustering of T/NK cells. After subsetting to only T/NK cells, principle component analysis, clustering, and UMAP was run following the same procedures as the whole dataset. The five resulting clusters are visualized in the T/NK cell specific UMAP space. b, Gene expression for T and NK cell markers.
Extended Data Fig. 4 Expression characteristics of the CD45– cluster.
a, Marker gene expression for non-fibroblast CD45- cell populations. Up to three characteristic gene markers are shown for each cluster by violin plot of normalized gene expression data. Fibroblast markers were used to identify the four fibroblast clusters Fib pre 1, Fib pre 2, Fib immune, and Fib cart. b, Stemness markers for fibroblasts. Pdgfra expression and clusters are shown to demonstrate location of fibroblasts with a dotted line drawn surrounding the fibroblasts. Stem marker Osr1 and CytoTRACE score, an algorithm used to score cells for stemness, is shown below. c, Expression of characteristic markers for the tenocyte-like and immune fibroblast clusters.
Extended Data Fig. 5 Flow cytometry of neutrophils and eosinophils.
a, Cells were gated on scatter (FSC_A, SSC_A) followed by doublet discrimination (FSC_A, FSC_H), selection of CD45+ live cells (Fixable Yellow-, CD45 + ), and selection of myeloid cells (CD3-, CD11b + ). This population was used to identify eosinophils (Ly6g low, Siglec F + ) and neutrophils (Ly6g + , Siglec F-) of correct size (FSC_A, SSC_A). b, Numbers of CD45+ cells, eosinophils, and neutrophils from ECM, PCL, or saline treated animals one week after surgery (top). Eosinophil and neutrophil amounts as proportion of CD45+ cells are given below. Data are mean ± SEM. Statistics shown are after analysis of variance (ANOVA) followed by Dunnett’s multiple comparison testing where P is adjusted p value.
Extended Data Fig. 6 Identification of intercluster signalling with Domino.
a, Identification of cluster-specific signaling subnetworks. Transcription factors enriched by cluster are identified by Wilcoxon rank sum and networks pruned for disconnected nodes to generate signaling subnetworks relevant for biological activation of clusters. b, Calculation of intercluster signaling networks. Once phenotypically relevant receptors are identified by cluster specific signaling subnetworks, cluster-cluster signaling scores are calculated by cluster averaged scaled expression of ligands present in cluster-specific subnetworks. Every potential cluster-cluster combination is scored, and these weights used to generate an intercluster signaling network.
Extended Data Fig. 7 Signalling pathways for ECM and PCL treated cells visualized with their known ligands.
a, A UMAP plot with clusters labeled for reference when viewing feature plots. b, ECM specific signaling pathways specified in Fig. 3. Each pathway contains gene expression of ligands and receptors from each pathway, as well as transcription-factor activation scores for the predicted transcription-factor target. Ligands completely absent from the data set are not shown, although they may still be viable targets for a target receptor. c, PCL specific signaling pathways specified in Fig. 3. No readily accepted ligands for Pirb have been identified, so the Pirb Irf4 pathway is not shown.
Extended Data Fig. 8 Myc signaling in the PCL-treated wound.
Gene expression of ligands and receptors predicted to be involved in activation of Myc in the PCL Domino signaling network for cells from PCL-treated animals (top and middle). Myc expression values are SCENIC transcription-factor activation scores (bottom). A reference of clusters is provided in the bottom left.
Extended Data Fig. 9 In vivo validation of signalling predictions by Domino.
a, Nanostring gene expression profiling for receptors and transcription factors identified as enriched in either ECM or PCL for specific cell types. After identifying genes specific to either ECM or PCL within the myeloid or T cell populations in Domino, we used Nanostring to probe expression of those genes in sorted myeloid (CD45+CD11b+) or T cells (CD45+CD3+). Of the 9 genes found to overlap between Domino and Nanostring, 7 were predicted correctly and 2 incorrectly. Error bars are standard error. b, The Domino global signaling network for PCL as seen in Fig. 5. The region surrounding the IL17 signaling pathway is enlarged and the components labeled. c, Expression of Il17rc and activation of it’s predicted transcription-factor targets Smarca4 and Mef2a in the PCL signaling network. d, Volcano plot for bulk RNA sequencing of IL17ra-/- and wild type animals one week after VML. Direction of log fold change indicates expression in IL17ra-/- animals compared to wild type. An FDR threshold of 0.05 was used to designate genes as significant. e, Gene set enrichment analysis (GSEA) of Domino transcription-factor modules predicted as downstream of IL17 in the PCL signaling network. Both modules predicted to interact are shown. Genes from the edgeR bulk RNA seq comparison with respect to wild type ordered by FDR were used to calculate enrichment for genes present in the Domino transcription-factor modules. Movement of the running enrichment score (green line) above zero indicates enrichment of genes in the module in the statistically significant portion of expressed genes while movement below zero indicates enrichment of genes in the module in the lower FDR portion of the genes. P values calculated by GSEA adjusted by Benjamini-Hochberg correction are shown.
Extended Data Fig. 10 Pathological signaling found from a public dataset of Alzheimer’s Disease.
a, The Alzheimer’s disease (AD) global signaling network. two modules of receptors and transcription factors are readily apparent and labeled based on enrichment of transcription factors by cluster. b, Heatmaps of transcription-factor activation score for AD-specific transcription factors (left) and correlation of transcription-factor activation score with receptor expression (right). Transcription factors are binned according to their membership to the astrocyte or oligodendrocyte modules from the AD global signaling network. Cells are ordered and colored according to their cluster. Receptors found only in AD are marked with arrows. Connections between receptor and transcription factors are marked with an ‘x’ on the correlation heatmap. c, Example feature plots of gene expression and activation scores for specific receptor–transcription-factor pairs identified by domino in the AD condition. d, The healthy global signaling network. Two modules of receptors and transcription factors are readily apparent and labeled based on enrichment of transcription factors by cluster. e, Heatmaps of transcription-factor activation score for healthy-specific transcription factors (left) and correlation of transcription-factor activation score with receptor expression (right). Transcription factors are binned according to their membership to the astrocyte or oligodendrocyte modules from the global signaling network. Cells are ordered and colored according to their cluster. Receptors found only in the healthy signaling network are marked with arrows. Connections between receptor and transcription factors are marked with an ‘x’ on the correlation heatmap. f, Example feature plots of gene expression and activation scores for specific receptor–transcription-factor pairs identified by domino in healthy cells.
Supplementary information
Supplementary Table 1
Comparison of Nanostring fibrosis-panel gene expression for PCL- and ECM-treated VML injuries, compared with a saline control, one week after injury.
Supplementary Table 2
Summary of samples in the single-cell atlas.
Supplementary Table 3
Differential expression for clusters.
Supplementary Table 4
Intracluster differential expression and gene-set enrichment analysis.
Supplementary Table 5
Numbers of cells allocated to clusters by experimental sample.
Supplementary Table 6
Antibodies used in flow cytometry.
Rights and permissions
About this article
Cite this article
Cherry, C., Maestas, D.R., Han, J. et al. Computational reconstruction of the signalling networks surrounding implanted biomaterials from single-cell transcriptomics. Nat Biomed Eng 5, 1228–1238 (2021). https://doi.org/10.1038/s41551-021-00770-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41551-021-00770-5
- Springer Nature Limited
This article is cited by
-
Modelling and targeting mechanical forces in organ fibrosis
Nature Reviews Bioengineering (2024)
-
Spatial transcriptomics analysis of neoadjuvant cabozantinib and nivolumab in advanced hepatocellular carcinoma identifies independent mechanisms of resistance and recurrence
Genome Medicine (2023)
-
Single-cell transcriptome reveals Staphylococcus aureus modulating fibroblast differentiation in the bone-implant interface
Molecular Medicine (2023)
-
Tracing immune cells around biomaterials with spatial anchors during large-scale wound regeneration
Nature Communications (2023)
-
Transforming bioengineering with unbiased teams and tools
Nature Reviews Bioengineering (2023)