Abstract
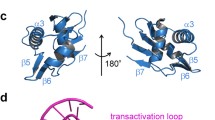
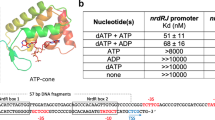
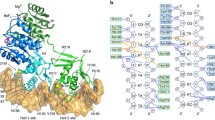
Two-component signal transduction systems are modular phosphorelay regulatory pathways common in prokaryotes. In the co-crystal structure of the Escherichia coli NarL signal output domain bound to DNA, we observe how the NarL family of two-component response regulators can bind DNA. DNA recognition is accompanied by the formation of a new dimerization interface, which could occur only in the full-length protein via a large intramolecular domain rearrangement. The DNA is recognized by the concerted effects of solvation, van der Waals forces and inherent DNA deformability, rather than determined primarily by major groove hydrogen bonding. These subtle forces permit a small DNA-binding domain to perturb the DNA helix, leading to major DNA curvature and a transition from B- to A-form DNA at the binding site, where valine on the recognition helix interacts unexpectedly with the polar major groove floor.





Similar content being viewed by others
Accession codes
References
Parkinson, J.S. & Kofoid, E.C. Communication modules in bacterial signalling proteins. Annu. Rev. Genet. 26, 71–112 (1992).
Stock, A.M., Robinson, V.L. & Goudreau, P.N. Two-component signal transduction. Annu. Rev. Biochem. 69, 183–215 (2000).
Robinson, V.L., Buckler, D.R. & Stock, A.M. A tale of two components: a novel kinase and a regulatory switch. Nature Struct. Biol. 7, 626–633 (2000).
Stewart, V. & Rabin, R.S. in Two-Component Signal Transduction (eds Hoch, J.A. & Silhavy, T.J.) 233–252 (American Society for Microbiology, Washington, D.C.; 1995).
Volz, K. & Matsumura, P. Crystal Structure of Escherichia coli CheY refined at 1.7 Å resolution. J. Biol. Chem 266, 15511–15519 (1991).
Baikalov, I. et al. Structure of the Escherichia coli response regulator NarL. Biochemistry 35, 11053–11061 (1996).
Baikalov, I. et al. NarL dimerization? Suggestive evidence from a new crystal form. Biochemistry 37, 3665–3676 (1998).
Pabo, C.O. & Nekludova, L. Geometric analysis and comparison of protein-DNA interfaces: why is there no simple code? J. Mol. Biol. 301, 597–624 (2000).
Darwin, A.J., Tyson, K.L., Busby, S.J.W. & Stewart, V. Differential regulation by the homologous response regulators NarL and NarP of Escherichia coli K-12 depends on DNA binding site arrangement. Mol. Micro. 25, 583–595 (1997).
Lonetto, M.A. et al. Identification of a contact site for different transcription activators in region 4 of the Escherichia coli RNA polymerase σ70 subunit. J. Mol. Biol. 284, 1353–1365 (1998).
Dickerson, R.E. & Chiu, T.K. Helix bending as a factor in protein/DNA recognition. Biopolymers 44, 361–403 (1998).
Dickerson, R.E. DNA bending: the prevalence of kinkiness and the virtues of normality. Nucleic Acids Res. 26, 1906–1926 (1998).
Lu, X.-J., Shakked, Z. & Olson, W.K. A-form conformational motifs in ligand-bound DNA structures. J. Mol. Biol. 300, 819–840 (2000).
Ng, H.-L., Kopka, M.L. & Dickerson, R.E. The structure of a stable intermediate in the A to B DNA helix transition. Proc. Natl. Acad. Sci. USA 97, 2035–2039 (2000).
Berman, H.M. et al. The Nucleic Acid Database: A comprehensive relational database of three-dimensional structures of nucleic acids. Biophys. J. 63, 751–759 (1992).
Woda, J., Schneider, B., Patel, K., Mistry, K. & Berman, H.M. An analysis of the relationship between hydration and protein-DNA interactions. Biophys. J. 75, 2170–2177 (1998).
Blanco, A.G., Sola, M., Gomis-Rueth, F.X. & Coll, M. Tandem DNA recognition by PhoB, a two-component signal transduction transcriptional activator. Structure 10, 701–713 (2002).
Tran, V.K., Oropeza, R. & Kenney, L.J. A single amino acid substitution in the C terminus of OmpR alters DNA recognition and phosphorylation. J. Mol. Biol. 299, 1257–1270 (2000).
Laughon, A. DNA binding specificity of homeodomains. Biochemistry 30, 11357–11367 (1991).
Fraenkel, E., Rould, M.A., Chambers, K.A. & Pabo, C.O. Engrailed homeodomain-DNA complex at 2.2 Å resolution: a detailed view of the interface and comparison with other engrailed structures. J. Mol. Biol. 284, 351–361 (1998).
Han, G.W., Kopka, M.L., Cascio, D., Grzeskowiak, K. & Dickerson, R.E. Structure of a DNA analog of the primer for HIV-1 RT second strand synthesis. J. Mol. Biol. 269, 811–826 (1997).
Hines, C.S. et al. DNA structure and flexibility in the sequence-specific binding of Papillomavirus E2 proteins. J. Mol. Biol. 276, 809–818 (1998).
Kim, S.-S., Tam, J.K., Wang, A.-F. & Hegde, R.S. The structural basis of DNA target discrimination by Papillomavirus E2 proteins. J. Biol. Chem 275, 31245–31254 (2000).
Hizver, J., Rozenberg, H., Frolow, F., Rabinovich, D. & Shakked, Z. DNA bending by an adenine-thymine tract and its role in gene regulation. Proc. Natl. Acad. Sci. USA 98, 8490–8495 (2001).
Gajiwala, K.S. et al. Structure of the winged-helix protein hRFX1 reveals a new mode of DNA binding. Nature 403, 916–919 (2000).
Djordjevic, S., Goudreau, P.N., Xu, Q., Stock, A.M. & West, A.H. Structural basis for methylesterase CheB regulation by a phosphorylation-activated domain. Proc. Natl. Acad. Sci. USA 95, 1381–1386 (1998).
Buckler, D.R., Zhou, Y. & Stock, A.M. Evidence of intradomain and interdomain flexibility in an OmpR/PhoB homolog from Thermotoga maritima. Structure 10, 153–164 (2002).
Zhang, R.-G. et al. Structure of a bacterial quorum-sensing transcription factor complexed with pheromone and DNA. Nature 417, 971–974 (2002).
Schroeder, I., Wolin, C.D., Cavicchioli, R. & Gunsalus, R.P. Phosphorylation and dephosphorylation of the NarX, NarL, and NarQ proteins of the nitrate dependent two-component regulatory system of Escherichia coli. J. Bacteriol. 176, 4985–4992 (1994).
Van Duyne, G.D., Standaert, R.F., Karplus, P.A., Schreiber, S.L. & Clardy, J. Atomic structures of the human immunophilin FKBP-12 complexes with FK506 and rapamycin. J. Mol. Biol. 229, 105–124 (1993).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
Maris, A.E. Molecular studies in protein/DNA recognition: crystallographic trials of the retrovirus integrase protein and the crystal structure of the Escherichia coli response regulator NarLC/DNA complex. PhD. thesis, Univ. California (2002).
Terwilliger, T.C. & Berendzen, J. Automated MAD and MIR structure solution. Acta Crystallogr. D 55, 849–861 (1999).
Jones, T.A., Zhou, J.-Y., Cowan, S.W. & Kjeldgaard, M. Improved methods for the building of protein models in electron density maps and the location of errors in these maps. Acta Crystallogr. A 47, 110–119 (1991).
Collaborative Computational Project, Number 4. The CCP4 suite: programs for protein crystallography. Acta. Crystallogr. D 50, 760–763 (1994).
McRee, D.E. XtalView/Xfit-A versatile program for manipulating atomic coordinates and electron density. J. Struct. Biol. 125, 156–165 (1999).
Tong, L. & Rossman, M.G. The locked rotation function. Acta Crystallogr. A 46, 783–792 (1990).
Brünger, A.T. et al. Crystallography and NMR System (CNS): a new software package for macromolecular structure determination. Acta Crystallogr. D 54, 905–921 (1998).
Bearson, S.M.D., Albrecht, J.A. & Gunsalus, R.P. Oxygen and nitrate-dependent regulation of dmsABC operon expression in Escherichia coli: sites for Fnr and NarL protein interactions. BMC Micro. 2, 13 (2002).
El Hassan, M.A. & Calladine, C.R. Conformational characteristics of DNA: empirical classifications and a hypothesis for the conformational behaviour of dinucleotide steps. Phil. Trans. R. Soc. Lond. A 355, 43–100 (1997).
Lavery, R. & Skelnar, H. The definition of generalized helicoidal parameters and of axis curvature for irregular nucleic acids. J. Biomol. Struct. Dynam. 6, 63–91 (1988).
Laskowski, R.A., MacArthur, M.W., Moss, D.S. & Thornton, J.M. PROCHECK: a program to check the stereochemical quality of protein structures. J. Appl. Crystallogr. 26, 283–291 (1993).
Acknowledgements
We thank A. Colasanti for his insightful discussions regarding B-to-A DNA transitions; D. Cascio for X-ray technical assistance and discussions; and G. Katsir, T. Chiu and H.-L. Ng for technical assistance and discussion. This work was supported by NIH grants.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Maris, A., Sawaya, M., Kaczor-Grzeskowiak, M. et al. Dimerization allows DNA target site recognition by the NarL response regulator. Nat Struct Mol Biol 9, 771–778 (2002). https://doi.org/10.1038/nsb845
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsb845
- Springer Nature America, Inc.
This article is cited by
-
Acetylation of NarL K188 and K192 is involved in regulating Escherichia coli anaerobic nitrate respiration
Applied Microbiology and Biotechnology (2022)
-
Transcription factors in microalgae: genome-wide prediction and comparative analysis
BMC Genomics (2016)
-
Two transcription factors, CabA and CabR, are independently involved in multilevel regulation of the biosynthetic gene cluster encoding the novel aminocoumarin, cacibiocin
Applied Microbiology and Biotechnology (2016)
-
The Escherichia coli NarL receiver domain regulates transcription through promoter specific functions
BMC Microbiology (2015)
-
Exploring comprehensive within-motif dependence of transcription factor binding in Escherichia coli
Scientific Reports (2015)