Abstract
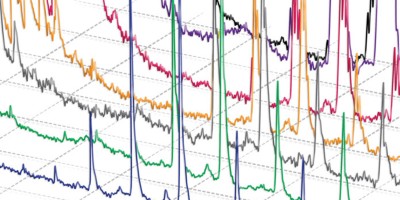
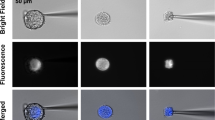
Single-cell mass spectrometry (MS) empowers metabolomic investigations by decreasing analytical dimensions to the size of individual cells and subcellular structures. We describe a protocol for investigating and quantifying metabolites in individual isolated neurons using single-cell capillary electrophoresis (CE) coupled to electrospray ionization (ESI) time-of-flight (TOF) MS. The protocol requires ∼2 h for sample preparation, neuron isolation and metabolite extraction, and 1 h for metabolic measurement. We used the approach to detect more than 300 distinct compounds in the mass range of typical metabolites in various individual neurons (25–500 μm in diameter) isolated from the sea slug (Aplysia californica) central and rat (Rattus norvegicus) peripheral nervous systems. We found that a subset of identified compounds was sufficient to reveal metabolic differences among freshly isolated neurons of different types and changes in the metabolite profiles of cultured neurons. The protocol can be applied to the characterization of the metabolome in a variety of smaller cells and/or subcellular domains.



Similar content being viewed by others
References
Wishart, D.S. et al. HMDB: a knowledgebase for the human metabolome. Nucleic Acids Res. 37, D603–D610 (2009).
Stephens, D.J. & Allan, V.J. Light microscopy techniques for live cell imaging. Science 300, 82–86 (2003).
Fischer, R.S., Wu, Y.C., Kanchanawong, P., Shroff, H. & Waterman, C.M. Microscopy in 3D: a biologist′s toolbox. Trends Cell Biol. 21, 682–691 (2011).
Hell, S.W. Far-field optical nanoscopy. Science 316, 1153–1158 (2007).
Planchon, T.A. et al. Rapid three-dimensional isotropic imaging of living cells using Bessel beam plane illumination. Nat. Methods 8, 417–U468 (2011).
Yan, R.X. et al. Nanowire-based single-cell endoscopy. Nat. Nanotechnol. 7, 191–196 (2012).
Wang, D.J. & Bodovitz, S. Single cell analysis: the new frontier in ′omics′. Trends Biotechnol. 28, 281–290 (2010).
Amantonico, A., Urban, P.L. & Zenobi, R. Analytical techniques for single-cell metabolomics: state of the art and trends. Anal. Bioanal. Chem. 398, 2493–2504 (2010).
Rubakhin, S.S., Romanova, E., Nemes, P. & Sweedler, J.V. Profiling metabolites and peptides in single cells. Nat. Methods 8, S20–S29 (2011).
Lin, Y.Q., Trouillon, R., Safina, G. & Ewing, A.G. Chemical analysis of single cells. Anal. Chem. 83, 4369–4392 (2011).
Nemes, P. & Vertes, A. Ambient mass spectrometry for in vivo local analysis and in situ molecular tissue imaging. Trac-Trends Anal. Chem. 34, 22–34 (2012).
Lee, Y.J., Perdian, D.C., Song, Z.H., Yeung, E.S. & Nikolau, B.J. Use of mass spectrometry for imaging metabolites in plants. Plant J. 70, 81–95 (2012).
Rubakhin, S.S., Lanni, E.J. & Sweedler, J.V. Progress toward single cell metabolomics. Curr. Opin. Biotechnol. 24, 95–104 (2013).
Rubakhin, S.S., Churchill, J.D., Greenough, W.T. & Sweedler, J.V. Profiling signaling peptides in single mammalian cells using mass spectrometry. Anal. Chem. 78, 7267–7272 (2006).
Li, L. & Sweedler, J.V. Peptides in the brain: mass spectrometry-based measurement approaches and challenges. Annu. Rev. Anal. Chem. 1, 451–483 (2008).
Neupert, S., Rubakhin, S.S. & Sweedler, J.V. Targeted single cell microchemical analysis: MS-based peptidomics of individual paraformaldehyde-fixed immunolabeled neurons. Chem. Biol. 19, 1010–1019 (2012).
Rubakhin, S.S. & Sweedler, J.V. Characterizing peptides in individual mammalian cells using mass spectrometry. Nat. Protoc. 2, 1987–1997 (2007).
Northen, T.R. et al. Clathrate nanostructures for mass spectrometry. Nature 449, 1033–1036 (2007).
Greving, M.P., Patti, G.J. & Siuzdak, G. Nanostructure-initiator mass spectrometry metabolite analysis and imaging. Anal. Chem. 83, 2–7 (2011).
Urban, P.L., Amantonico, A., Fagerer, S.R., Gehrig, P. & Zenobi, R. Mass spectrometric method incorporating enzymatic amplification for attomole-level analysis of target metabolites in biological samples. Chem. Commun. 46, 2212–2214 (2010).
Stolee, J.A., Walker, B.N., Zorba, V., Russo, R.E. & Vertes, A. Laser-nanostructure interactions for ion production. PCCP 14, 8453–8471 (2012).
Monroe, E.B., Jurchen, J.C., Lee, J., Rubakhin, S.S. & Sweedler, J.V. Vitamin E imaging and localization in the neuronal membrane. J. Am. Chem. Soc. 127, 12152–12153 (2005).
Yang, H.J. et al. Detection of characteristic distributions of phospholipid head groups and fatty acids on neurite surface by time-of-flight secondary ion mass spectrometry. Med. Mol. Morphol. 43, 158–164 (2010).
Kurczy, M.E. et al. Mass spectrometry imaging of mating Tetrahymena show that changes in cell morphology regulate lipid domain formation. Proc. Natl Acad. Sci. USA 107, 2751–2756 (2010).
Tsuyama, N., Mizuno, H. & Masujima, T. Mass spectrometry for cellular and tissue analyses in a very small region. Anal. Sci. 27, 163–170 (2011).
Nemes, P. & Vertes, A. Laser ablation electrospray ionization for atmospheric pressure, in vivo, and imaging mass spectrometry. Anal. Chem. 79, 8098–8106 (2007).
Shrestha, B., Patt, J.M. & Vertes, A. In situ cell-by-cell imaging and analysis of small cell populations by mass spectrometry. Anal. Chem. 83, 2947–2955 (2011).
Coello, Y., Jones, A.D., Gunaratne, T.C. & Dantus, M. Atmospheric-pressure femtosecond laser imaging mass spectrometry. Anal. Chem. 82, 2753–2758 (2010).
Cecala, C. & Sweedler, J.V. Sampling techniques for single-cell electrophoresis. Analyst 137, 2922–2929 (2012).
Lapainis, T. & Sweedler, J.V. Contributions of capillary electrophoresis to neuroscience. J. Chromatogr. A 1184, 144–158 (2008).
Lapainis, T., Rubakhin, S.S. & Sweedler, J.V. Capillary electrophoresis with electrospray ionization mass spectrometric detection for single-cell metabolomics. Anal. Chem. 81, 5858–5864 (2009).
Page, J.S., Rubakhin, S.S. & Sweedler, J.V. Single-neuron analysis using CE combined with MALDI MS and radionuclide detection. Anal. Chem. 74, 497–503 (2002).
Nemes, P., Knolhoff, A.M., Rubakhin, S.S. & Sweedler, J.V. Metabolic differentiation of neuronal phenotypes by single-cell capillary electrophoresis-electrospray ionization-mass spectrometry. Anal. Chem. 83, 6810–6817 (2011).
Nemes, P., Knolhoff, A.M., Rubakhin, S.S. & Sweedler, J.V. Single-cell metabolomics: Changes in the metabolome of freshly isolated and cultured neurons. ACS Chem. Neurosci. 3, 782–792 (2012).
Bonner, R.F. et al. Cell sampling-laser capture microdissection: Molecular analysis of tissue. Science 278, 1481–1483 (1997).
Espina, V. et al. Laser-capture microdissection. Nat. Protoc. 1, 586–603 (2006).
Miao, J. & Cui, L.W. Rapid isolation of single malaria parasite–infected red blood cells by cell sorting. Nat. Protoc. 6, 140–146 (2011).
Cohen, D. et al. Chemical cytometry: fluorescence-based single-cell analysis. Annu. Rev. Anal. Chem. 1, 165–190 (2008).
Bodenmiller, B. et al. Multiplexed mass cytometry profiling of cellular states perturbed by small-molecule regulators. Nat. Biotechnol. 30, 858–867 (2012).
Mellors, J.S., Jorabchi, K., Smith, L.M. & Ramsey, J.M. Integrated microfluidic device for automated single cell analysis using electrophoretic separation and electrospray ionization mass spectrometry. Anal. Chem. 82, 967–973 (2010).
Schoenherr, R.M., Ye, M.L., Vannatta, M. & Dovichi, N.J. CE-microreactor-CE-MS/MS for protein analysis. Anal. Chem. 79, 2230–2238 (2007).
Barbula, G.K., Safi, S., Chingin, K., Perry, R.H. & Zare, R.N. Interfacing capillary-based separations to mass spectrometry using desorption electrospray ionization. Anal. Chem. 83, 1955–1959 (2011).
Harris, G.A., Nyadong, L. & Fernandez, F.M. Recent developments in ambient ionization techniques for analytical mass spectrometry. Analyst 133, 1297–1301 (2008).
Knolhoff, A.M., Nemes, P., Rubakhin, S.S. & Sweedler, J.V. in Methodologies for metabolomics: Experimental strategies and techniques (eds. Norbert Lutz, Jonathan V. Sweedler & Ron A. Wevers) 119–139 (Cambridge University Press, 2013).
Patti, G.J., Tautenhahn, R. & Siuzdak, G. Meta-analysis of untargeted metabolomic data from multiple profiling experiments. Nat. Protoc. 7, 508–516 (2012).
Pluskal, T., Castillo, S., Villar-Briones, A. & Oresic, M. MZmine 2: modular framework for processing, visualizing, and analyzing mass spectrometry-based molecular profile data. BMC Bioinformatics 11, 395 (2010).
Baran, R. et al. MathDAMP: a package for differential analysis of metabolite profiles. BMC Bioinformatics 7, 530 (2006).
Kilkenny, C., Browne, W.J., Cuthill, I.C., Emerson, M. & Altman, D.G. Improving bioscience research reporting: the ARRIVE guidelines for reporting animal research. PLoS Biol. 8, e1000412 (2010).
Smith, C.A. et al. METLIN - a metabolite mass spectral database. Ther. Drug Monit. 27, 747–751 (2005).
Southey, B.R., Amare, A., Zimmerman, T.A., Rodriguez-Zas, S.L. & Sweedler, J.V. NeuroPred: a tool to predict cleavage sites in neuropeptide precursors and provide the masses of the resulting peptides. Nucleic Acids Res. 34, W267–W272 (2006).
Geigl, J.B. & Speicher, M.R. Single-cell isolation from cell suspensions and whole genome amplification from single cells to provide templates for CGH analysis. Nat. Protoc. 2, 3173–3184 (2007).
Neupert, S. & Predel, R. Mass spectrometric analysis of single identified neurons of an insect. Biochem. Biophys. Res. Commun. 327, 640–645 (2005).
Eyer, K., Kuhn, P., Hanke, C. & Dittrich, P.S. A microchamber array for single cell isolation and analysis of intracellular biomolecules. Lab Chip 12, 765–772 (2012).
De Sousa, B.N. & Horrocks, L.A. Development of rat spinal cord. I. Weight and length with a method for rapid removal. Dev. Neurosci. 2, 115–121 (1979).
Nemes, P., Marginean, I. & Vertes, A. Spraying mode effect on droplet formation and ion chemistry in electrosprays. Anal. Chem. 79, 3105–3116 (2007).
Acknowledgements
T. Lapainis contributed to the development of the CE-ESI-MS platform, X. Wang facilitated the single-neuron preparation work and A.M. Knolhoff helped with the interpretation of CE-ESI-MS data in the original CE-MS studies. This work was supported by the National Science Foundation under award no. CHE 11-11705, the National Institute of Neurological Disorders and Stroke under award no. NS031609, the National Institute of Dental and Craniofacial Research and the Office of the Director, US National Institutes of Health under award no. DE018866 and the National Institute on Drug Abuse under award no. P30 DA018310. The mention of commercial products, their sources or their use in connection with material reported herein is not to be construed as either actual or implied endorsement of such products by the Department of Health and Human Services. The authors thank S. Baker for assistance in the preparation of this publication and the machine shop of the School of Chemical Sciences, University of Illinois at Urbana-Champaign, for providing various components for the construction of the CE-ESI system. They also thank M. Kim for help in photography.
Author information
Authors and Affiliations
Contributions
The contributions of the authors to the original studies that were used to generate the protocol are indicated in the original cited articles. For the protocols described here, P.N. generated many of the protocol steps related to CE-MS operation and wrote the data analysis script and virtual instrument; J.T.A. modified the protocols and validated them using the DRG neurons; S.S.R. performed the cell isolations and optimized the protocols for cell handling; and J.V.S. modified and clarified the protocol. All authors participated in the writing of the protocol.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Data
Supplementary sequence archive (ZIP 120 kb)
Supplementary Methods
Stable ion generation, reproducible separation, and semiautomated data analysis (PDF 901 kb)
Rights and permissions
About this article
Cite this article
Nemes, P., Rubakhin, S., Aerts, J. et al. Qualitative and quantitative metabolomic investigation of single neurons by capillary electrophoresis electrospray ionization mass spectrometry. Nat Protoc 8, 783–799 (2013). https://doi.org/10.1038/nprot.2013.035
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2013.035
- Springer Nature Limited
This article is cited by
-
Lipidomic profiling of single mammalian cells by infrared matrix-assisted laser desorption electrospray ionization (IR-MALDESI)
Analytical and Bioanalytical Chemistry (2020)
-
A microanalytical capillary electrophoresis mass spectrometry assay for quantifying angiotensin peptides in the brain
Analytical and Bioanalytical Chemistry (2019)
-
Tapered-Tip Capillary Electrophoresis Nano-Electrospray Ionization Mass Spectrometry for Ultrasensitive Proteomics: the Mouse Cortex
Journal of the American Society for Mass Spectrometry (2017)