Abstract
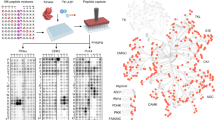
The ubiquitous nature of protein phosphorylation makes it challenging to map kinase-substrate relationships, which is a necessary step toward defining signaling network architecture. To trace the activity of individual kinases, we developed a semisynthetic reaction scheme, which results in the affinity tagging of substrates of the kinase in question. First, a kinase, engineered to use a bio-orthogonal ATPγS analog, catalyzes thiophosphorylation of its direct substrates. Second, alkylation of thiophosphorylated serine, threonine or tyrosine residues creates an epitope for thiophosphate ester–specific antibodies. We demonstrated the generality of semisynthetic epitope construction with 13 diverse kinases: JNK1, p38α MAPK, Erk1, Erk2, Akt1, PKCδ, PKCε, Cdk1/cyclinB, CK1, Cdc5, GSK3β, Src and Abl. Application of this approach, in cells isolated from a mouse that expressed endogenous levels of an analog-specific (AS) kinase (Erk2), allowed purification of a direct Erk2 substrate.
NOTE: In the version of this article initially published online, a sentence was missing a word and did not make sense. The corrected sentence now reads, “Erk2 was immunoprecipitated from each of these cell lines and assayed with A*TPγS analogs; N6-phenethyl ATPγS was a preferred nucleotide substrate for AS Erk2 and was not accepted by wild-type Erk2 (data not shown).” The error has been corrected for all versions of the article.





Similar content being viewed by others
Change history
08 May 2007
In the version of this article initially published online, a sentence was missing a word and did not make sense. The corrected sentence now reads, “Erk2 was immunoprecipitated from each of these cell lines and assayed with A*TPγS analogs; N6-phenethyl ATPγS was a preferred nucleotide substrate for AS Erk2 and was not accepted by wild-type Erk2 (data not shown).” The error has been corrected for all versions of the article.
References
Berwick, D.C. & Tavare, J.M. Identifying protein kinase substrates: hunting for the organ-grinder's monkeys. Trends Biochem. Sci. 29, 227–232 (2004).
Ptacek, J. & Snyder, M. Charging it up: global analysis of protein phosphorylation. Trends Genet. 22, 545–554 (2006).
Ptacek, J. et al. Global analysis of protein phosphorylation in yeast. Nature 438, 679–684 (2005).
Morrison, D.K. & Davis, R.J. Regulation of MAP kinase signaling modules by scaffold proteins in mammals. Annu. Rev. Cell Dev. Biol. 19, 91–118 (2003).
Biondi, R.M. & Nebreda, A.R. Signalling specificity of Ser/Thr protein kinases through docking-site-mediated interactions. Biochem. J. 372, 1–13 (2003).
Shah, K. et al. Engineering unnatural nucleotide specificity for Rous sarcoma virus tyrosine kinase to uniquely label its direct substrates. Proc. Natl. Acad. Sci. USA 94, 3565–3570 (1997).
Witucki, L.A. et al. Mutant tyrosine kinases with unnatural nucleotide specificity retain the structure and phospho-acceptor specificity of the wild-type enzyme. Chem. Biol. 9, 25–33 (2002).
Ubersax, J.A. et al. Targets of the cyclin-dependent kinase Cdk1. Nature 425, 859–864 (2003).
Dephoure, N. et al. Combining chemical genetics and proteomics to identify protein kinase substrates. Proc. Natl. Acad. Sci. USA 102, 17940–17945 (2005).
Lee, W.C. & Lee, K.H. Applications of affinity chromatography in proteomics. Anal. Biochem. 324, 1–10 (2004).
Eblen, S.T. et al. Identification of novel ERK2 substrates through use of an engineered kinase and ATP analogs. J. Biol. Chem. 278, 14926–14935 (2003).
Jaeschke, A. et al. JNK2 is a positive regulator of the cJun transcription factor. Mol. Cell 23, 899–911 (2006).
Wang, H. et al. Inducible protein knockout reveals temporal requirement of CaMKII reactivation for memory consolidation in the brain. Proc. Natl. Acad. Sci. USA 100, 4287–4292 (2003).
Chen, X. et al. A chemical-genetic approach to studying neurotrophin signaling. Neuron 46, 13–21 (2005).
Allen, J.J., Lazerwith, S.E. & Shokat, K.M. Bio-orthogonal affinity purification of direct kinase substrates. J. Am. Chem. Soc. 127, 5288–5289 (2005).
Sondhi, D. et al. Peptide and protein phosphorylation by protein tyrosine kinase Csk: insights into specificity and mechanism. Biochemistry 37, 165–172 (1998).
Olsen, J.V. et al. Global, in vivo, and site-specific phosphorylation dynamics in signaling networks. Cell 127, 635–648 (2006).
Hoffman, C., Genieser, H.G., Veron, M. & Jastorff, B. Novel synthesis of nucleoside 5′-polyphosphates. Bioorg. Med. Chem. Lett. 6, 2571–2574 (1996).
Goody, R.S. & Eckstein, F. Thiophosphate analogs of nucleoside di- and triphosphates. J. Am. Chem. Soc. 93, 6252–6257 (1971).
Kikkawa, U., Matsuzaki, H. & Yamamoto, T. Protein kinase C delta (PKC delta): activation mechanisms and functions. J. Biochem. 132, 831–839 (2002).
Lee, K.S. et al. Yeast polo-like kinases: functionally conserved multitask mitotic regulators. Oncogene 24, 217–229 (2005).
Weston, C.R. & Davis, R.J. The JNK signal transduction pathway. Curr. Opin. Cell. Biol. 19, 142–149 (2007).
Gaarde, W.A., Hunter, T., Brady, H., Murray, B.W. & Goldman, M.E. Development of a nonradioactive, time-resolved fluorescence assay for the measurement of Jun N-terminal kinase activity. J. Biomol. Screen. 2, 213–223 (1997).
Zhang, C. et al. A second-site suppressor strategy for chemical genetic analysis of diverse protein kinases. Nat. Methods 2, 435–441 (2005).
Rush, J. et al. Immunoaffinity profiling of tyrosine phosphorylation in cancer cells. Nat. Biotechnol. 23, 94–101 (2005).
Zhang, H. et al. Phosphoprotein analysis using antibodies broadly reactive against phosphorylated motifs. J. Biol. Chem. 277, 39379–39387 (2002).
Prescher, J.A. & Bertozzi, C.R. Chemistry in living systems. Nat. Chem. Biol. 1, 13–21 (2005).
Kikugawa, K., Iizuka, K. & Ichino, M. Platelet aggregation inhibitors. 4. N6-substituted adenosines. J. Med. Chem. 16, 358–364 (1973).
Holzenberger, M. et al. Cre-mediated germline mosaicism: a method allowing rapid generation of several alleles of a target gene. Nucleic Acids Res. 28, E92 (2000).
Acknowledgements
We thank D. Randle and D. Morgan for wild-type Cdc5, D. Kenski for performing GRK2 kinase assays, H. Luecke for helpful suggestions, D. Maly for helpful comments on the manuscript, and V. Ohman for excellent administrative assistance. Erk1−/− mice were generously provided by G. Pages and J. Pouyssegur (Centre National de la Recherche Scientifique, Nice, France). HeLa cells were provided by the National Cell Culture Center. This work was supported by the US National Institutes of Health (NIH) National Institute of Biomedical Imaging and BioEngineering R01 EB001987, and in part by NIH grant AA013588 and funds provided by the State of California for medical research on alcohol and substance abuse through the University of California at San Francisco (to R.O.M.). J.J.A. was funded by an Achievement Rewards for College Scientists fellowship, Genentech, and the Sandler Foundation. Mass spectrometry was made possible by NIH grants NCRR RR015804 and NCRR RR001614.
Author information
Authors and Affiliations
Contributions
J.J.A. and K.M.S. wrote the manuscript. J.J.A. performed all experiments except those mentioned below. M.L. performed Erk2 substrate labeling reactions, C.S.B. performed mass spectrometry of modified peptides, J.L.P. expressed Cdc5, D.W. provided technical assistance for PKCδ experiments, A.H. constructed JNK1 plasmids, C.D.K. constructed the AS Erk2 mouse, W.C. constructed PKCδ plasmids, A.L.B. advised C.S.B., R.O.M. advised D.W. and W.C., and assisted in manuscript revision, R.J.D. advised A.H., S.M.H. advised M.L. and C.D.K. and K.M.S. advised J.J.A.
Corresponding author
Ethics declarations
Competing interests
Antibody 51-8 was made for a fee paid by grant NIH EB001987 to Epitomics Inc., under an academic-institution agreement with Epitomics Inc.. University of California San Francisco is presently negotiating with Epitomics to commercialize this antibody along with the necessary reagents for labeling kinase substrates.
Supplementary information
Supplementary Fig. 1
Western blot analysis of α-hapten–IgY specificity and full length western blots for main text figures. (PDF 545 kb)
Supplementary Fig. 2
Screen for thiophosphate ester specific IgG. (PDF 397 kb)
Supplementary Fig. 3
Specificity and yield of PNBM alkylation. (PDF 99 kb)
Supplementary Fig. 4
DELFIA for thiophosphorylation kinetics. (PDF 92 kb)
Supplementary Fig. 5
Synthesis of hapten and thioether control. (PDF 51 kb)
Supplementary Fig. 6
Synthesis and purification of A*TPγS analogs. (PDF 150 kb)
Supplementary Table 1
Generality of AS kinase engineering. (PDF 70 kb)
Rights and permissions
About this article
Cite this article
Allen, J., Li, M., Brinkworth, C. et al. A semisynthetic epitope for kinase substrates. Nat Methods 4, 511–516 (2007). https://doi.org/10.1038/nmeth1048
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth1048
- Springer Nature America, Inc.
This article is cited by
-
Innate immune and proinflammatory signals activate the Hippo pathway via a Tak1-STRIPAK-Tao axis
Nature Communications (2024)
-
The LRR receptor-like kinase ALR1 is a plant aluminum ion sensor
Cell Research (2024)
-
In vitro kinase assay reveals ADP-heptose-dependent ALPK1 autophosphorylation and altered kinase activity of disease-associated ALPK1 mutants
Scientific Reports (2023)
-
Enhanced SREBP2-driven cholesterol biosynthesis by PKCλ/ι deficiency in intestinal epithelial cells promotes aggressive serrated tumorigenesis
Nature Communications (2023)
-
The Hippo signaling component LATS2 enhances innate immunity to inhibit HIV-1 infection through PQBP1-cGAS pathway
Cell Death & Differentiation (2022)