Abstract
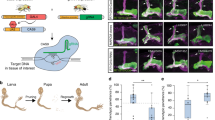
Informational recoding by adenosine-to-inosine RNA editing diversifies neuronal proteomes by chemically modifying structured mRNAs. However, techniques for analyzing editing activity on substrates in defined neurons in vivo are lacking. Guided by comparative genomics, here we reverse-engineered a fluorescent reporter sensitive to Drosophila melanogaster adenosine deaminase that acts on RNA (dADAR) activity and alterations in dADAR autoregulation. Using this artificial dADAR substrate, we visualized variable patterns of RNA-editing activity in the Drosophila nervous system between individuals. Our results demonstrate the feasibility of structurally mimicking ADAR substrates as a method to regulate protein expression and, potentially, therapeutically repair mutant mRNAs. Our data suggest variable RNA editing as a credible molecular mechanism for mediating individual-to-individual variation in neuronal physiology and behavior.





Similar content being viewed by others
References
Nishikura, K. Functions and regulation of RNA editing by ADAR deaminases. Annu. Rev. Biochem. 79, 321–349 (2010).
Hoopengardner, B., Bhalla, T., Staber, C. & Reenan, R. Nervous system targets of RNA editing identified by comparative genomics. Science 301, 832–836 (2003).
Seeburg, P.H. & Hartner, J. Regulation of ion channel/neurotransmitter receptor function by RNA editing. Curr. Opin. Neurobiol. 13, 279–283 (2003).
Higuchi, M. et al. Point mutation in an AMPA receptor gene rescues lethality in mice deficient in the RNA-editing enzyme ADAR2. Nature 406, 78–81 (2000).
Palladino, M.J., Keegan, L.P., O'Connell, M.A. & Reenan, R.A. A-to-I pre-mRNA editing in Drosophila is primarily involved in adult nervous system function and integrity. Cell 102, 437–449 (2000).
Tonkin, L.A. et al. RNA editing by ADARs is important for normal behavior in Caenorhabditis elegans. EMBO J. 21, 6025–6035 (2002).
Higuchi, M. et al. RNA editing of AMPA receptor subunit GluR-B: a base-paired intron-exon structure determines position and efficiency. Cell 75, 1361–1370 (1993).
Reenan, R.A. Molecular determinants and guided evolution of species-specific RNA editing. Nature 434, 409–413 (2005).
Gommans, W.M., McCane, J., Nacarelli, G.S. & Maas, S. A mammalian reporter system for fast and quantitative detection of intracellular A-to-I RNA editing levels. Anal. Biochem. 399, 230–236 (2010).
Herbert, A., Wagner, S. & Nickerson, J.A. Induction of protein translation by ADAR1 within living cell nuclei is not dependent on RNA editing. Mol. Cell 10, 1235–1246 (2002).
Pokharel, S. & Beal, P.A. High-throughput screening for functional adenosine to inosine RNA editing systems. ACS Chem. Biol. 1, 761–765 (2006).
Keegan, L.P. et al. Tuning of RNA editing by ADAR is required in Drosophila. EMBO J. 24, 2183–2193 (2005).
Palladino, M.J., Keegan, L.P., O'Connell, M.A. & Reenan, R.A. dADAR, a Drosophila double-stranded RNA-specific adenosine deaminase is highly developmentally regulated and is itself a target for RNA editing. RNA 6, 1004–1018 (2000).
Jepson, J.E. & Reenan, R.A. Adenosine-to-inosine genetic recoding is required in the adult stage nervous system for coordinated behavior in Drosophila. J. Biol. Chem. 284, 31391–31400 (2009).
Jepson, J.E. et al. Engineered alterations in RNA editing modulate complex behavior in Drosophila: regulatory diversity of adenosine deaminase acting on RNA (ADAR) targets. J. Biol. Chem. 286, 8325–8337 (2011).
Keene, A.C. & Waddell, S. Drosophila olfactory memory: single genes to complex neural circuits. Nat. Rev. Neurosci. 8, 341–354 (2007).
Masse, N.Y., Turner, G.C. & Jefferis, G.S. Olfactory information processing in Drosophila. Curr. Biol. 19, R700–R713 (2009).
Yin, J.C. & Tully, T. CREB and the formation of long-term memory. Curr. Opin. Neurobiol. 6, 264–268 (1996).
Decher, N. et al. RNA editing modulates the binding of drugs and highly unsaturated fatty acids to the open pore of Kv potassium channels. EMBO J. 29, 2101–2113 (2010).
Ingleby, L., Maloney, R., Jepson, J., Horn, R. & Reenan, R. Regulated RNA editing and functional epistasis in Shaker potassium channels. J. Gen. Physiol. 133, 17–27 (2009).
Jones, A.K. et al. Splice-variant- and stage-specific RNA editing of the Drosophila GABA receptor modulates agonist potency. J. Neurosci. 29, 4287–4292 (2009).
Rosenthal, J.J. & Bezanilla, F. Extensive editing of mRNAs for the squid delayed rectifier K+ channel regulates subunit tetramerization. Neuron 34, 743–757 (2002).
Chou, Y.H. et al. Diversity and wiring variability of olfactory local interneurons in the Drosophila antennal lobe. Nat. Neurosci. 13, 439–449 (2010).
Murthy, M., Fiete, I. & Laurent, G. Testing odor response stereotypy in the Drosophila mushroom body. Neuron 59, 1009–1023 (2008).
Claridge-Chang, A. et al. Writing memories with light-addressable reinforcement circuitry. Cell 139, 405–415 (2009).
Dracheva, S. et al. Increased serotonin 2C receptor mRNA editing: a possible risk factor for suicide. Mol. Psychiatry 13, 1001–1010 (2008).
Gurevich, I. et al. Altered editing of serotonin 2C receptor pre-mRNA in the prefrontal cortex of depressed suicide victims. Neuron 34, 349–356 (2002).
Bhogal, B. et al. Modulation of dADAR-dependent RNA editing by the Drosophila fragile X mental retardation protein. Nat. Neurosci. 14, 1517–1524 (2011).
Acknowledgements
We thank C. Lawrence and members of the Reenan laboratory for helpful discussions, C. Staber and G. Williams for expert technical assistance, and K. Koh (Thomas Jefferson University) for reagents. This work was funded by an Ellison Medical Foundation Senior Scholar award.
Author information
Authors and Affiliations
Contributions
J.E.C.J. performed all imaging experiments and analyzed the data. Y.A.S. and J.E.C.J. performed RNA-editing analysis. R.A.R. designed the GFPedit reporter. K.A.J. and R.A.R. cloned the GFPedit and YFPsplice constructs and performed in vitro validation experiments. Y.A.S. and J.E.C.J. generated the recombinant dAdar alleles. J.E.C.J. and R.A.R. wrote the paper, with contributions from Y.A.S.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–8 and Supplementary Table 1 (PDF 1660 kb)
Rights and permissions
About this article
Cite this article
Jepson, J., Savva, Y., Jay, K. et al. Visualizing adenosine-to-inosine RNA editing in the Drosophila nervous system. Nat Methods 9, 189–194 (2012). https://doi.org/10.1038/nmeth.1827
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.1827
- Springer Nature America, Inc.
This article is cited by
-
Emerging strategies for the genetic dissection of gene functions, cell types, and neural circuits in the mammalian brain
Molecular Psychiatry (2022)
-
Deciphering the functions and regulation of brain-enriched A-to-I RNA editing
Nature Neuroscience (2013)
-
RNA editing regulates transposon-mediated heterochromatic gene silencing
Nature Communications (2013)
-
Auto-regulatory RNA editing fine-tunes mRNA re-coding and complex behaviour in Drosophila
Nature Communications (2012)
-
Making sense out of nonsense to visualize editing in the fly nervous system
Nature Methods (2012)