Abstract
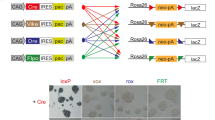
We have developed dual recombinase-mediated cassette exchange (dRMCE) to efficiently re-engineer the thousands of available conditional alleles in mouse embryonic stem cells. dRMCE takes advantage of the wild-type loxP and FRT sites present in these conditional alleles and in many gene-trap lines. dRMCE is a scalable, flexible tool to introduce tags, reporters and mutant coding regions into an endogenous locus of interest in an easy and highly efficient manner.


Similar content being viewed by others
References
Glaser, S., Anastassiadis, K. & Stewart, A.F. Nat. Genet. 37, 1187–1193 (2005).
Branda, C.S. & Dymecki, S.M. Dev. Cell 6, 7–28 (2004).
Lauth, M., Spreafico, F., Dethleffsen, K. & Meyer, M. Nucleic Acids Res. 30, e115 (2002).
Collins, F.S., Rossant, J. & Wurst, W. Cell 128, 9–13 (2007).
Shimshek, D.R. et al. Genesis 32, 19–26 (2002).
Raymond, C.S. & Soriano, P. PLoS ONE 2, e162 (2007).
Kanda, T., Sullivan, K.F. & Wahl, G.M. Curr. Biol. 8, 377–385 (1998).
Galli, A. et al. PLoS Genet. 6, e1000901 (2010).
Nagy, A. Manipulating the Mouse Embryo: A Laboratory Manual 3rd edn. (Cold Spring Harbor Laboratory Press, Cold Spring Harbor, New York, USA, 2003).
Pettitt, S.J. et al. Nat. Methods 6, 493–495 (2009).
Anastassiadis, K. et al. Dis. Model, Mech. 2, 508–515 (2009).
Singla, V. et al. Nat. Methods 7, 50–52 (2010).
Skarnes, W.C. et al. Nat. Genet. 36, 543–544 (2004).
Szymczak, A.L. et al. Nat. Biotechnol. 22, 589–594 (2004).
Chu, G.C., Dunn, N.R., Anderson, D.C., Oxburgh, L. & Robertson, E.J. Development 131, 3501–3512 (2004).
Acknowledgements
We are grateful to D. Klewe-Nebenius and T. Hennek (University of Basel) for generating the chimeric mice. We thank I. Verma (Salk Institute, San Diego, California, USA), B. Sauer (Stowers Institute, Kansas City, Missouri, USA), R. Jaenisch (Whitehead Institute, Cambridge, Massachusetts, USA) and P. Soriano (Mount Sinai School of Medicine, New York, New York, USA) for plasmids obtained via Addgene and A.-K. Hadjantonakis (Sloan-Kettering Institute, New York, New York, USA) for suggesting the use of the H2B-Venus fusion protein. We thank A. Schauerte and P. Lorentz for expert technical assistance and are indebted to P. Bovolenta, G. Nusspaumer, A. Zuniga and members of our research groups for helpful input and discussions. Our research is supported by the Swiss National Science Foundation (to R.Z.), by a Marie Curie Intra-European Fellowship and a European Reintegration Grant (to J.L.-R.) and by both cantons of Basel, Switzerland.
Author information
Authors and Affiliations
Contributions
M.O., J.L.-R. and R.Z. conceived and designed the experiments. M.O., J.L.-R. and B.R. designed and constructed the dRMCE tool-kit vectors. B.R. and W.C.S. provided the IKMC mouse embryonic stem cell lines. M.O., A.G., J.L.-R. and B.R. performed the experiments. J.L.-R., W.C.S. and R.Z. wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
A dRMCE patent is pending.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–5, Supplementary Tables 1 and 2 (PDF 8023 kb)
Supplementary Data
Vector sequences in GenBank format. (ZIP 30 kb)
Rights and permissions
About this article
Cite this article
Osterwalder, M., Galli, A., Rosen, B. et al. Dual RMCE for efficient re-engineering of mouse mutant alleles. Nat Methods 7, 893–895 (2010). https://doi.org/10.1038/nmeth.1521
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.1521
- Springer Nature America, Inc.