Abstract
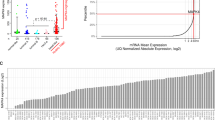
Triple-negative breast cancers (TNBCs) have poor prognosis and lack targeted therapies. Here we identified increased copy number and expression of the PIM1 proto-oncogene in genomic data sets of patients with TNBC. TNBC cells, but not nonmalignant mammary epithelial cells, were dependent on PIM1 for proliferation and protection from apoptosis. PIM1 knockdown reduced expression of the anti-apoptotic factor BCL2, and dynamic BH3 profiling of apoptotic priming revealed that PIM1 prevents mitochondrial-mediated apoptosis in TNBC cell lines. In TNBC tumors and their cellular models, PIM1 expression was associated with several transcriptional signatures involving the transcription factor MYC, and PIM1 depletion in TNBC cell lines decreased, in a MYC-dependent manner, cell population growth and expression of the MYC target gene MCL1. Treatment with the pan–PIM kinase inhibitor AZD1208 impaired the growth of both cell line and patient-derived xenografts and sensitized them to standard-of-care chemotherapy. This work identifies PIM1 as a malignant-cell-selective target in TNBC and the potential use of PIM1 inhibitors for sensitizing TNBC to chemotherapy-induced apoptotic cell death.






Similar content being viewed by others
Accession codes
Change history
26 January 2017
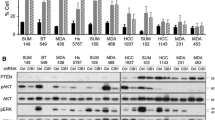
In the version of this article initially published, the western blot in Figure 2a for the BT474 cell line was incorrect. In addition, one of the accession codes for gene expression data for the Guy's Hospital TNBC-enriched cohort was incorrect. The errors have been corrected in the HTML and PDF versions of the article.
References
Reis-Filho, J.S. & Tutt, A.N. Triple-negative tumors: a critical review. Histopathology 52, 108–118 (2008).
Symmans, W.F. et al. Measurement of residual breast cancer burden to predict survival after neoadjuvant chemotherapy. J. Clin. Oncol. 25, 4414–4422 (2007).
Masuda, H. et al. Differential response to neoadjuvant chemotherapy among seven triple-negative breast cancer molecular subtypes. Clin. Cancer Res. 19, 5533–5540 (2013).
Balko, J.M. et al. Molecular profiling of the residual disease of triple-negative breast cancers after neoadjuvant chemotherapy identifies actionable therapeutic targets. Cancer Discov. 4, 232–245 (2014).
Labisso, W.L. et al. MYC directs transcription of MCL1 and EIF4E genes to control sensitivity of gastric cancer cells toward HDAC inhibitors. Cell Cycle 11, 1593–1602 (2012).
Andre, F. et al. Molecular characterization of breast cancer with high-resolution oligonucleotide comparative genomic hybridization array. Clin. Cancer Res. 15, 441–451 (2009).
Bergamaschi, A. et al. Distinct patterns of DNA copy-number alteration are associated with different clinicopathological features and gene expression subtypes of breast cancer. Genes Chromosom. Cancer 45, 1033–1040 (2006).
Turner, N. et al. Integrative molecular profiling of triple-negative breast cancers identifies amplicon drivers and potential therapeutic targets. Oncogene 29, 2013–2023 (2010).
Nawijn, M.C., Alendar, A. & Berns, A. For better or for worse: the role of Pim oncogenes in tumorigenesis. Nat. Rev. Cancer 11, 23–34 (2011).
Wang, J. et al. Pim1 kinase synergizes with c-MYC to induce advanced prostate carcinoma. Oncogene 29, 2477–2487 (2010).
Keeton, E.K. et al. AZD1208, a potent and selective pan–Pim kinase inhibitor, demonstrates efficacy in preclinical models of acute myeloid leukemia. Blood 123, 905–913 (2014).
Laird, P.W. et al. In vivo analysis of Pim1 deficiency. Nucleic Acids Res. 21, 4750–4755 (1993).
Magnuson, N.S., Wang, Z., Ding, G. & Reeves, R. Why target PIM1 for cancer diagnosis and treatment? Future Oncol. 6, 1461–1478 (2010).
Narlik-Grassow, M., Blanco-Aparicio, C. & Carnero, A. The PIM family of serine–threonine kinases in cancer. Med. Res. Rev. 34, 136–159 (2014).
Aceto, N. et al. Tyrosine phosphatase SHP2 promotes breast cancer progression and maintains tumor-initiating cells via activation of key transcription factors and a positive-feedback signaling loop. Nat. Med. 18, 529–537 (2012).
Speers, C. et al. Identification of novel kinase targets for the treatment of estrogen-receptor-negative breast cancer. Clin. Cancer Res. 15, 6327–6340 (2009).
Lehmann, B.D. et al. Identification of human triple-negative breast cancer subtypes and preclinical models for selection of targeted therapies. J. Clin. Invest. 121, 2750–2767 (2011).
de Rinaldis, E. et al. Integrated genomic analysis of triple-negative breast cancers reveals novel microRNAs associated with clinical and molecular phenotypes and sheds light on the pathways they control. BMC Genomics 14, 643 (2013).
Gazinska, P. et al. Comparison of basal-like triple-negative breast cancer defined by morphology, immunohistochemistry and transcriptional profiles. Mod. Pathol. 26, 955–966 (2013).
The Cancer Genome Atlas. Comprehensive molecular portraits of human breast tumors. Nature 490, 61–70 (2012).
Curtis, C. et al. The genomic and transcriptomic architecture of 2,000 breast tumors reveals novel subgroups. Nature 486, 346–352 (2012).
Parker, J.S. et al. Supervised risk predictor of breast cancer based on intrinsic subtypes. J. Clin. Oncol. 27, 1160–1167 (2009).
Neve, R.M. et al. A collection of breast cancer cell lines for the study of functionally distinct cancer subtypes. Cancer Cell 10, 515–527 (2006).
Canbay, E. Erb-B2 homodimerization inhibits MUC1 transcription in cultured human mammary epithelial cells. Cell Biol. Int. 27, 477–481 (2003).
Aho, T.L. et al. Pim1 kinase promotes inactivation of the pro-apoptotic Bad protein by phosphorylating it on the Ser112 gatekeeper site. FEBS Lett. 571, 43–49 (2004).
Lilly, M., Sandholm, J., Cooper, J.J., Koskinen, P.J. & Kraft, A. The PIM1 serine kinase prolongs survival and inhibits apoptosis-related mitochondrial dysfunction in part through a BCL-2-dependent pathway. Oncogene 18, 4022–4031 (1999).
Juin, P., Geneste, O., Gautier, F., Depil, S. & Campone, M. Decoding and unlocking the BCL-2 dependency of cancer cells. Nat. Rev. Cancer 13, 455–465 (2013).
Montero, J. et al. Drug-induced death signaling strategy rapidly predicts cancer response to chemotherapy. Cell 160, 977–989 (2015).
Verbeek, S. et al. Mice bearing the Eμ-Myc and Eμ-Pim1 transgenes develop pre-B-cell leukemia prenatally. Mol. Cell. Biol. 11, 1176–1179 (1991).
Zippo, A., De Robertis, A., Serafini, R. & Oliviero, S. PIM1-dependent phosphorylation of histone H3 at Ser10 is required for MYC-dependent transcriptional activation and oncogenic transformation. Nat. Cell Biol. 9, 932–944 (2007).
Sears, R. et al. Multiple Ras-dependent phosphorylation pathways regulate Myc protein stability. Genes Dev. 14, 2501–2514 (2000).
Noguchi, K. et al. Regulation of c-Myc through phosphorylation at Ser-62 and Ser-71 by c-Jun N-terminal kinase. J. Biol. Chem. 274, 32580–32587 (1999).
Wang, X. et al. Phosphorylation regulates c-Myc's oncogenic activity in the mammary gland. Cancer Res. 71, 925–936 (2011).
Chandriani, S. et al. A core MYC gene expression signature is prominent in basal-like breast cancer but only partially overlaps the core serum response. PLoS One 4, e6693 (2009).
Horiuchi, D. et al. MYC pathway activation in triple-negative breast cancer is synthetic lethal with CDK inhibition. J. Exp. Med. 209, 679–696 (2012).
Grigoriadis, A. et al. Molecular characterization of cell line models for triple-negative breast cancers. BMC Genomics 13, 619 (2012).
Morishita, D., Katayama, R., Sekimizu, K., Tsuruo, T. & Fujita, N. Pim kinases promote cell cycle progression by phosphorylating and down-regulating p27Kip1 at the transcriptional and posttranscriptional levels. Cancer Res. 68, 5076–5085 (2008).
Miura, K. et al. Involvement of EphA2-mediated tyrosine phosphorylation of Shp2 in Shp2-regulated activation of extracellular-signal-regulated kinase. Oncogene 32, 5292–5301 (2013).
Hudis, C.A. & Gianni, L. Triple-negative breast cancer: an unmet medical need. Oncologist 16 (Suppl. 1), 1–11 (2011).
Kirschner, A.N. et al. PIM kinase inhibitor AZD1208 for treatment of MYC-driven prostate cancer. J. Natl. Cancer Inst. 107, dju407 (2014).
Turner, N.C. & Reis-Filho, J.S. Tackling the diversity of triple-negative breast cancer. Clin. Cancer Res. 19, 6380–6388 (2013).
Morton, J.P. & Sansom, O.J. MYC-y mice: from tumor initiation to therapeutic targeting of endogenous MYC. Mol. Oncol. 7, 248–258 (2013).
Horiuchi, D. et al. PIM kinase inhibition as a targeted therapy against triple-negative breast tumors with elevated MYC expression. Nat. Med. http://dx.doi.org/10.1038/nm.4213 (2016).
Petrocca, F. et al. A genome-wide siRNA screen identifies proteasome addiction as a vulnerability of basal-like triple-negative breast cancer cells. Cancer Cell 24, 182–196 (2013).
Garcia, P.D. et al. Pan–PIM kinase inhibition provides a novel therapy for treating hematologic cancers. Clin. Cancer Res. 20, 1834–1845 (2014).
Malinen, M. et al. Proto-oncogene PIM1 is a novel estrogen receptor target associating with high-grade breast tumors. Mol. Cell. Endocrinol. 365, 270–276 (2013).
Raab, M.S. et al. Phase 1 study of the novel pan–PIM kinase inhibitor LGH447 in patients with relapsed or refractory multiple myeloma. in 55th ASH Annual meeting and exposition (New Orleans, LA, 2013).
McEachern, K.A. et al. Preclinical and clinical pharmacodynamics of pan-PIM inhibition by AZD1208 in acute myeloid leukemia: assessment of PIM isoform dependency for BAD and 4EBP1 phosphorylation. in 56th ASH Annual meeting and exposition (San Francisco, CA, 2014).
Mori, S. et al. Utilization of pathway signatures to reveal distinct types of B lymphoma in the Eμ-myc model and human diffuse large-B-cell lymphoma. Cancer Res. 68, 8525–8534 (2008).
Bild, A.H. et al. Oncogenic pathway signatures in human cancers as a guide to targeted therapies. Nature 439, 353–357 (2006).
Zeller, K.I., Jegga, A.G., Aronow, B.J., O'Donnell, K.A. & Dang, C.V. An integrated database of genes responsive to the Myc oncogenic transcription factor: identification of direct genomic targets. Genome Biol. 4, R69 (2003).
O'Donnell, K.A. et al. Activation of transferrin receptor 1 by c-Myc enhances cellular proliferation and tumorigenesis. Mol. Cell. Biol. 26, 2373–2386 (2006).
Fernandez, P.C. et al. Genomic targets of the human c-Myc protein. Genes Dev. 17, 1115–1129 (2003).
Kim, Y.H. et al. Combined microarray analysis of small-cell lung cancer reveals altered apoptotic balance and distinct expression signatures of MYC family gene amplification. Oncogene 25, 130–138 (2006).
Lastowska, M. et al. Comprehensive genetic and histopathologic study reveals three types of neuroblastoma tumors. J. Clin. Oncol. 19, 3080–3090 (2001).
Lee, J.S. et al. Application of comparative functional genomics to identify best-fit mouse models to study human cancer. Nat. Genet. 36, 1306–1311 (2004).
Menssen, A. & Hermeking, H. Characterization of the c-MYC-regulated transcriptome by SAGE: identification and analysis of c-MYC target genes. Proc. Natl. Acad. Sci. USA 99, 6274–6279 (2002).
Schlosser, I. et al. Dissection of transcriptional programs in response to serum and c-Myc in a human B cell line. Oncogene 24, 520–524 (2005).
Schuhmacher, M. et al. The transcriptional program of a human B cell line in response to Myc. Nucleic Acids Res. 29, 397–406 (2001).
Yu, D., Cozma, D., Park, A. & Thomas-Tikhonenko, A. Functional validation of genes implicated in lymphomagenesis: an in vivo selection assay using a Myc-induced B cell tumor. Ann. NY Acad. Sci. 1059, 145–159 (2005).
Hammond, M.E. et al. American Society of Clinical Oncology–College Of American Pathologists guideline recommendations for immunohistochemical testing of estrogen and progesterone receptors in breast cancer. J. Clin. Oncol. 28, 2784–2795 (2010).
Van Loo, P. et al. Allele-specific copy-number analysis of tumors. Proc. Natl. Acad. Sci. USA 107, 16910–16915 (2010).
Rasmussen, M. et al. Allele-specific copy-number analysis of tumor samples with aneuploidy and tumor heterogeneity. Genome Biol. 12, R108 (2011).
Meerbrey, K.L. et al. The pINDUCER lentiviral toolkit for inducible RNA interference in vitro and in vivo. Proc. Natl. Acad. Sci. USA 108, 3665–3670 (2011).
Franken, N.A., Rodermond, H.M., Stap, J., Haveman, J. & van Bree, C. Clonogenic assay of cells in vitro. Nat. Protoc. 1, 2315–2319 (2006).
Acknowledgements
This research was supported by the Breast Cancer Now funding (A.N.T.) at King's College London and the Institute of Cancer Research London, the National Institute for Health Research Biomedical Research Centre based at Guy's and St Thomas' NHS Foundation Trust and King's College London, the US National Institutes of Health grant P30 CA008748 (M.S.) and the Breast Cancer Research Foundation (M.S.). The PDX studies were supported by a 'GHD-pink' research support via the FERO Foundation (V.S.). Patient samples and data were provided by King's Health Partners Cancer Biobank, London, UK, which is supported by the Experimental Cancer Medicine Centre at King's College London and the Department of Health via the National Institute for Health Research comprehensive Biomedical Research Centre award. We thank S. Swift, S. Utting and M. Ferrao for technical and administrative assistance, AstraZeneca for providing AZD1208, D. Huszar (AstraZeneca) for providing information on the use of AZD1208, H. Mirza (King's College London) for his support with the bioinformatics analysis, J. Hurst (The Institute of Cancer Research London) for helping with the NanoString nCounter PanCancer Pathway analysis, M. Dowsett (Institute of Cancer Research and Royal Marsden Hospital National Institute for Health Research comprehensive Biomedical Research Centre London) for providing staff and infrastructure support for the NanoString NCounter work, T. Tenev (Institute of Cancer Research) for providing MDA-MB-231 containing the BCL2 pTIPZ overexpression vector and the Guy's Hospital Pharmacy for providing the chemotherapeutic drugs.
Author information
Authors and Affiliations
Contributions
A.N.T., A.G., F.B.-M., S.F. and P. Marra conceived and designed the research; F.B.-M. and S.F. performed the in vitro experiments with crucial help from S.C., E.N., S.M., M. Shafat, N.P. and A.P.-R.; J.W. and J.Q. performed the bioinformatics analyses; R.M., E.F.-D., D.A.P. and L.Z. performed the cell line xenograft experiments; V.S. and A.G.-O. performed the PDX experiments; P.G., E.F.-D., L.Z. and F.N. performed IHC stainings; K.M. provided AZD1208 and expertise, and assisted in the design of in vitro and in vivo AZD1208 experiments; G.L. and P. Meier performed dynamic BH3 profiling analysis; R.B. performed the NanoString nCounter experiments; M.C.U.C. performed the statistical analysis of the gene expression analysis; P.C. and M. Scaltriti performed western blot analysis with the PIM1 NOV22-39-5 antibody, and F.B.-M. analyzed most of the results; F.B.-M. and A.N.T wrote the manuscript; A.N.T. coordinated this work; and all authors helped with data interpretation and manuscript editing.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text, Figures and Tables
Supplementary Figures 1–16 and Supplementary Tables 1–3 (PDF 28936 kb)
Rights and permissions
About this article
Cite this article
Brasó-Maristany, F., Filosto, S., Catchpole, S. et al. PIM1 kinase regulates cell death, tumor growth and chemotherapy response in triple-negative breast cancer. Nat Med 22, 1303–1313 (2016). https://doi.org/10.1038/nm.4198
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nm.4198
- Springer Nature America, Inc.
This article is cited by
-
Synthesis and biological evaluation of chromone derivatives against triple-negative breast cancer cells
Medicinal Chemistry Research (2023)
-
Integrated analysis of ceRNA network reveals potential prognostic Hint1-related lncRNAs involved in hepatocellular carcinoma progression
World Journal of Surgical Oncology (2022)
-
Antitumor activity of the protein kinase inhibitor 1-(β-D-2′-deoxyribofuranosyl)-4,5,6,7-tetrabromo- 1H-benzimidazole in breast cancer cell lines
BMC Cancer (2022)
-
Exploring the mechanism and experimental verification of puerarin in the treatment of endometrial carcinoma based on network pharmacology and bioinformatics analysis
BMC Complementary Medicine and Therapies (2022)
-
PIM1 promotes hepatic conversion by suppressing reprogramming-induced ferroptosis and cell cycle arrest
Nature Communications (2022)