Abstract
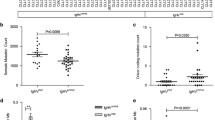
To understand the genetic mechanisms driving variant and IGHV4-34–expressing hairy-cell leukemias, we performed whole-exome sequencing of leukemia samples from ten affected individuals, including six with matched normal samples. We identified activating mutations in the MAP2K1 gene (encoding MEK1) in 5 of these 10 samples and in 10 of 21 samples in a validation set (overall frequency of 15/31), suggesting potential new strategies for treating individuals with these diseases.

Similar content being viewed by others
Accession codes
References
Arons, E. & Kreitman, R.J. Leuk. Lymphoma 52 (suppl. 2), 99–102 (2011).
Tiacci, E. et al. N. Engl. J. Med. 364, 2305–2315 (2011).
Dietrich, S. et al. N. Engl. J. Med. 366, 2038–2040 (2012).
Tiacci, E. et al. Blood 119, 192–195 (2012).
Xi, L. et al. Blood 119, 3330–3332 (2012).
Laurini, J.A., Aoun, P., Iqbal, J., Chan, W. & Greiner, T.C. Am. J. Clin. Pathol. 138, 877–883 (2012).
Bromberg-White, J.L., Andersen, N.J. & Duesbery, N.S. Brief. Funct. Genomics 11, 300–310 (2012).
Rossi, D. et al. J. Exp. Med. 209, 1537–1551 (2012).
Wagle, N. et al. J. Clin. Oncol. 29, 3085–3096 (2011).
Emery, C.M. et al. Proc. Natl. Acad. Sci. USA 106, 20411–20416 (2009).
Marks, J.L. et al. Cancer Res. 68, 5524–5528 (2008).
Nikolaev, S.I. et al. Nat. Genet. 44, 133–139 (2012).
Fischmann, T.O. et al. Biochemistry 48, 2661–2674 (2009).
Mansour, S.J. et al. Science 265, 966–970 (1994).
Flaherty, K.T. et al. N. Engl. J. Med. 367, 1694–1703 (2012).
Hockley, S.L. et al. Leukemia 25, 1189–1192 (2011).
Hahn, C.N. & Scott, H.S. Nat. Genet. 44, 9–10 (2012).
Imielinski, M. et al. Cell 150, 1107–1120 (2012).
Wu, J.N. & Roberts, C.W.M. Cancer Discov. 3, 35–43 (2013).
Arons, E., Suntum, T., Stetler-Stevenson, M. & Kreitman, R.J. Blood 114, 4687–4695 (2009).
Acknowledgements
This work was supported by the Intramural Research Program of the US NIH, NCI, CCR (P.S.M. and R.J.K.) and the Hairy Cell Leukemia Research Foundation (R.J.K.).
Author information
Authors and Affiliations
Contributions
J.J.W., E.A., R.J.K. and P.S.M. conceived the study and supervised analyses. J.J.W. designed and performed analyses. E.A. and R.J.K. provided patient materials. L.R. extracted DNA and processed patient samples. R.L.W. performed exome sequencing. M.P. conducted Sanger sequencing and TaqMan analysis. J.K.K. designed and supervised TaqMan assays. S.R.D. and O.D.A. provided computational scripts. J.J.W., R.J.K. and P.S.M. wrote the manuscript with contributions from all other authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Note, Supplementary Tables 1–3 and 5, and Supplementary Figure 1 (PDF 970 kb)
Supplementary Table 4
Candidate driver mutations from whole-exome sequencing (XLS 82 kb)
Rights and permissions
About this article
Cite this article
Waterfall, J., Arons, E., Walker, R. et al. High prevalence of MAP2K1 mutations in variant and IGHV4-34–expressing hairy-cell leukemias. Nat Genet 46, 8–10 (2014). https://doi.org/10.1038/ng.2828
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.2828
- Springer Nature America, Inc.
This article is cited by
-
A comparison of the International Consensus and 5th World Health Organization classifications of mature B-cell lymphomas
Leukemia (2023)
-
Comparison of histological and molecular features of pediatric-type follicular lymphoma and pediatric nodal marginal zone lymphoma
Virchows Archiv (2023)
-
The many faces of nodal and splenic marginal zone lymphomas. A report of the 2022 EA4HP/SH lymphoma workshop
Virchows Archiv (2023)
-
Genetic Variations in Hairy Cell Leukemia with Brain Involvement: Case Report
SN Comprehensive Clinical Medicine (2023)
-
The 2022 classifications of lymphoid neoplasms
Die Pathologie (2023)