Abstract
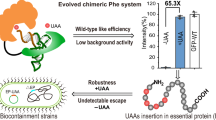
Expansion of the genetic code with nonstandard amino acids (nsAAs) has enabled biosynthesis of proteins with diverse new chemistries. However, this technology has been largely restricted to proteins containing a single or few nsAA instances. Here we describe an in vivo evolution approach in a genomically recoded Escherichia coli strain for the selection of orthogonal translation systems capable of multi-site nsAA incorporation. We evolved chromosomal aminoacyl-tRNA synthetases (aaRSs) with up to 25-fold increased protein production for p-acetyl-L-phenylalanine and p-azido-L-phenylalanine (pAzF). We also evolved aaRSs with tunable specificities for 14 nsAAs, including an enzyme that efficiently charges pAzF while excluding 237 other nsAAs. These variants enabled production of elastin-like-polypeptides with 30 nsAA residues at high yields (∼50 mg/L) and high accuracy of incorporation (>95%). This approach to aaRS evolution should accelerate and expand our ability to produce functionalized proteins and sequence-defined polymers with diverse chemistries.





Similar content being viewed by others
References
Seitchik, J.L. et al. Genetically encoded tetrazine amino acid directs rapid site-specific in vivo bioorthogonal ligation with trans-cyclooctenes. J. Am. Chem. Soc. 134, 2898–2901 (2012).
Chin, J.W. Expanding and reprogramming the genetic code of cells and animals. Annu. Rev. Biochem. 83, 379–408 (2014).
Link, A.J., Mock, M.L. & Tirrell, D.A. Non-canonical amino acids in protein engineering. Curr. Opin. Biotechnol. 14, 603–609 (2003).
Liu, C.C. & Schultz, P.G. Adding new chemistries to the genetic code. Annu. Rev. Biochem. 79, 413–444 (2010).
O'Donoghue, P., Ling, J., Wang, Y.S. & Söll, D. Upgrading protein synthesis for synthetic biology. Nat. Chem. Biol. 9, 594–598 (2013).
Tian, F. et al. A general approach to site-specific antibody drug conjugates. Proc. Natl. Acad. Sci. USA 111, 1766–1771 (2014).
Furman, J.L. et al. A genetically encoded aza-Michael acceptor for covalent cross-linking of protein-receptor complexes. J. Am. Chem. Soc. 136, 8411–8417 (2014).
Kang, M. et al. Evolution of iron(II)-finger peptides by using a bipyridyl amino acid. ChemBioChem 15, 822–825 (2014).
Wang, F., Niu, W., Guo, J. & Schultz, P.G. Unnatural amino acid mutagenesis of fluorescent proteins. Angew. Chem. Int. Edn Engl. 51, 10132–10135 (2012).
Li, X. & Liu, C.C. Biological applications of expanded genetic codes. ChemBioChem 15, 2335–2341 (2014).
Dieterich, D.C., Link, A.J., Graumann, J., Tirrell, D.A. & Schuman, E.M. Selective identification of newly synthesized proteins in mammalian cells using bioorthogonal noncanonical amino acid tagging (BONCAT). Proc. Natl. Acad. Sci. USA 103, 9482–9487 (2006).
Yuet, K.P. & Tirrell, D.A. Chemical tools for temporally and spatially resolved mass spectrometry-based proteomics. Ann. Biomed. Eng. 42, 299–311 (2014).
Nishi, Y. et al. Different effects of 4-hydroxyproline and 4-fluoroproline on the stability of collagen triple helix. Biochemistry 44, 6034–6042 (2005).
Kothakota, S., Mason, T.L., Tirrell, D.A. & Fournier, M.J. Biosynthesis of a periodic protein containing 3-thienylalanine - a step toward genetically-engineered conducting polymers. J. Am. Chem. Soc. 117, 536–537 (1995).
Bae, J.H. et al. Incorporation of beta-selenolo[3,2-b]pyrrolyl-alanine into proteins for phase determination in protein X-ray crystallography. J. Mol. Biol. 309, 925–936 (2001).
Kirshenbaum, K., Carrico, I.S. & Tirrell, D.A. Biosynthesis of proteins incorporating a versatile set of phenylalanine analogs. ChemBioChem 3, 235–237 (2002).
Young, D.D. et al. An evolved aminoacyl-tRNA synthetase with atypical polysubstrate specificity. Biochemistry 50, 1894–1900 (2011).
Park, H.S. et al. Expanding the genetic code of Escherichia coli with phosphoserine. Science 333, 1151–1154 (2011).
Umehara, T. et al. N-acetyl lysyl-tRNA synthetases evolved by a CcdB-based selection possess N-acetyl lysine specificity in vitro and in vivo. FEBS Lett. 586, 729–733 (2012).
Johnson, D.B. et al. RF1 knockout allows ribosomal incorporation of unnatural amino acids at multiple sites. Nat. Chem. Biol. 7, 779–786 (2011).
Lajoie, M.J. et al. Genomically recoded organisms expand biological functions. Science 342, 357–360 (2013).
Heinemann, I.U. et al. Enhanced phosphoserine insertion during Escherichia coli protein synthesis via partial UAG codon reassignment and release factor 1 deletion. FEBS Lett. 586, 3716–3722 (2012).
Mukai, T. et al. Codon reassignment in the Escherichia coli genetic code. Nucleic Acids Res. 38, 8188–8195 (2010).
Isaacs, F.J. et al. Precise manipulation of chromosomes in vivo enables genome-wide codon replacement. Science 333, 348–353 (2011).
Wiltschi, B., Wenger, W., Nehring, S. & Budisa, N. Expanding the genetic code of Saccharomyces cerevisiae with methionine analogs. Yeast 25, 775–786 (2008).
Nehring, S., Budisa, N. & Wiltschi, B. Performance analysis of orthogonal pairs designed for an expanded eukaryotic genetic code. PLoS One 7, e31992 (2012).
Zaher, H.S. & Green, R. Fidelity at the molecular level: lessons from protein synthesis. Cell 136, 746–762 (2009).
Odoi, K.A., Huang, Y., Rezenom, Y.H. & Liu, W.R. Nonsense and sense suppression abilities of original and derivative Methanosarcina mazei pyrrolysyl-tRNA synthetase-tRNAPyl pairs in the Escherichia coli BL21(DE3) cell strain. PLoS One 8, e57035 (2013).
Wang, H.H. et al. Programming cells by multiplex genome engineering and accelerated evolution. Nature 460, 894–898 (2009).
Gallagher, R.R., Li, Z., Lewis, A.O. & Isaacs, F.J. Rapid editing and evolution of bacterial genomes using libraries of synthetic DNA. Nat. Protoc. 9, 2301–2316 (2014).
MacEwan, S.R. & Chilkoti, A. Elastin-like polypeptides: biomedical applications of tunable biopolymers. Biopolymers 94, 60–77 (2010).
Young, T.S., Ahmad, I., Yin, J.A. & Schultz, P.G. An enhanced system for unnatural amino acid mutagenesis in E. coli. J. Mol. Biol. 395, 361–374 (2010).
Pédelacq, J.D., Cabantous, S., Tran, T., Terwilliger, T.C. & Waldo, G.S. Engineering and characterization of a superfolder green fluorescent protein. Nat. Biotechnol. 24, 79–88 (2006).
Sharan, S.K., Thomason, L.C., Kuznetsov, S.G. & Court, D.L. Recombineering: a homologous recombination-based method of genetic engineering. Nat. Protoc. 4, 206–223 (2009).
DeVito, J.A. Recombineering with tolC as a selectable/counter-selectable marker: remodeling the rRNA operons of Escherichia coli. Nucleic Acids Res. 36 (1), e4 (2008).
Kobayashi, T. et al. Structural basis of nonnatural amino acid recognition by an engineered aminoacyl-tRNA synthetase for genetic code expansion (vol. 102, pages 1366, 2005). Proc. Natl. Acad. Sci. USA 102, 1366–1371 (2005).
Schultz, K.C. et al. A genetically encoded infrared probe. J. Am. Chem. Soc. 128, 13984–13985 (2006).
Wang, L., Zhang, Z., Brock, A. & Schultz, P.G. Addition of the keto functional group to the genetic code of Escherichia coli. Proc. Natl. Acad. Sci. USA 100, 56–61 (2003).
Cooley, R.B. et al. Structural basis of improved second-generation 3-nitro-tyrosine tRNA synthetases. Biochemistry 53, 1916–1924 (2014).
Stokes, A.L. et al. Enhancing the utility of unnatural amino acid synthetases by manipulating broad substrate specificity. Mol. Biosyst. 5, 1032–1038 (2009).
Ko, J.H. et al. Pyrrolysyl-tRNA synthetase variants reveal ancestral aminoacylation function. FEBS Lett. 587, 3243–3248 (2013).
Guo, L.T. et al. Polyspecific pyrrolysyl-tRNA synthetases from directed evolution. Proc. Natl. Acad. Sci. USA 111, 16724–16729 (2014).
Aerni, H.R., Shifman, M.A., Rogulina, S., O'Donoghue, P. & Rinehart, J. Revealing the amino acid composition of proteins within an expanded genetic code. Nucleic Acids Res. 43, e8 (2015).
Chin, J.W. et al. Addition of p-azido-L-phenylalanine to the genetic code of Escherichia coli. J. Am. Chem. Soc. 124, 9026–9027 (2002).
Wu, I.L. et al. Multiple site-selective insertions of noncanonical amino acids into sequence-repetitive polypeptides. ChemBioChem 14, 968–978 (2013).
Meyer, D.E. & Chilkoti, A. Quantification of the effects of chain length and concentration on the thermal behavior of elastin-like polypeptides. Biomacromolecules 5, 846–851 (2004).
Tinberg, C.E. et al. Computational design of ligand-binding proteins with high affinity and selectivity. Nature 501, 212–216 (2013).
Rovner, A.J. et al. Recoded organisms engineered to depend on synthetic amino acids. Nature 518, 89–93 (2015).
Mandell, D.J. et al. Biocontainment of genetically modified organisms by synthetic protein design. Nature 518, 55–60 (2015).
Lajoie, M.J. et al. Probing the limits of genetic recoding in essential genes. Science 342, 361–363 (2013).
Gibson, D.G. et al. Enzymatic assembly of DNA molecules up to several hundred kilobases. Nat. Methods 6, 343–345 (2009).
Murphy, K.C. Use of bacteriophage lambda recombination functions to promote gene replacement in Escherichia coli. J. Bacteriol. 180, 2063–2071 (1998).
Meyer, D.E. & Chilkoti, A. Genetically encoded synthesis of protein-based polymers with precisely specified molecular weight and sequence by recursive directional ligation: examples from the elastin-like polypeptide system. Biomacromolecules 3, 357–367 (2002).
Acknowledgements
We thank K. Bilguvar and J. Knight (Yale Center for Genome Analysis) for conducting next-generation sequencing experiments; T. Wu (Yale West Campus Analytical Core facility) for conducting intact MS experiments; B. Gassaway for assistance with shotgun MS experiments. We are grateful to members of the Isaacs laboratory, J. Ling and G. Church for critical discussions and feedback. This work was supported by the Defense Advanced Research Projects Agency contracts N66001-12-C-4020 and N66001-12-C-4211 to (F.J.I., J.R., D.S., and M.C.J.), U.S. Department of Energy (DE-FG02-02ER63445 to F.J.I.) grants GM22854 to D.S. and GM67193 to N.L.K. from the National Institute for General Medical Sciences, T32GM007205 and 1F30CA196191 (A.D.H.), Army Research Office (W911NF- 11-1-0445 to M.C.J.), the David and Lucile Packard Foundation (M.C.J.), the Camille Dreyfus Teacher-Scholar Program (M.C.J.), DuPont, Inc. (F.J.I.) and the Arnold and Mabel Beckman Foundation (F.J.I.).
Author information
Authors and Affiliations
Contributions
M.A. designed ELP constructs, and conducted and interpreted multi-site nsAA incorporation experiments. M.A., A.D.H. and F.J.I. designed, conducted and interpreted synthetase evolution experiments. H.-R.A. and J.R. conducted and interpreted MS experiments. I.N. and N.L.K. performed and interpreted top-down MS experiments. A.D.H, A.L.G. and F.J.I. analyzed NextGen sequencing experiments. C.F., D.W.M. and D.S. conducted and interpreted biochemical experiments. Y.-S.W. executed nsAA screens. S.H.H. tested ELP expression in vitro. M.A., A.D.H. and A.J.R. performed crystal structure analysis and target selection. N.J.M. constructed and characterized the tolC variant used for negative selection. F.J.I., J.R. and D.S. directed the studies and interpreted data. M.A. and F.J.I. wrote the paper with assistance from A.D.H., N.J.M., D.S., M.C.J., I.N., N.L.K. and J.R.
Corresponding authors
Ethics declarations
Competing interests
M.A., A.D.H. and F.J.I. have filed a provisional application with the US Patent and Trademark Office on this work.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–16, Supplementary Tables 1–7 and Supplementary Notes 1–7 (PDF 8120 kb)
Rights and permissions
About this article
Cite this article
Amiram, M., Haimovich, A., Fan, C. et al. Evolution of translation machinery in recoded bacteria enables multi-site incorporation of nonstandard amino acids. Nat Biotechnol 33, 1272–1279 (2015). https://doi.org/10.1038/nbt.3372
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.3372
- Springer Nature America, Inc.