Abstract
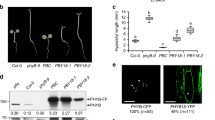
A plant modulates its developmental processes in response to light by several informational photoreceptors such as phytochrome. Phytochrome is a dimeric chromoprotein which regulates various aspects of plant development from seed germination to flowering1. Upon absorption of red light, phytochrome translocates from the cytoplasm to the nucleus2,3,4, and regulates gene expression through interaction with transcription factors such as PIF3 (refs 5–7). The phytochrome polypeptide has two domains1,8: the amino-terminal photosensory domain with a chromophore and the carboxy-terminal domain which contains signalling motifs such as a kinase domain9. The latter is widely believed to transduce the signal to downstream components1,5,8,9,10. Here we show that the C-terminal domain of Arabidopsis phytochrome B (phyB), which is known as the most important member of the phytochrome family1, is not directly involved in signal transduction. The N-terminal domain isolated from phyB, when dimerized and localized in the nucleus, triggered full phyB responses with much higher photosensitivity than the full-length phyB. These findings indicate that the C-terminal domain attenuates the activity of phyB rather than positively transducing the signal.





Similar content being viewed by others
References
Smith, H. Phytochromes and light signal perception by plants—an emerging synthesis. Nature 407, 585–591 (2000)
Yamaguchi, R., Nakamura, M., Mochizuki, N., Kay, S. A. & Nagatani, A. Light-dependent translocation of a phytochrome B:GFP fusion protein to the nucleus in transgenic Arabidopsis. J. Cell Biol. 145, 437–445 (1999)
Nagy, F., Kircher, S. & Schäfer, E. Nucleo-cytoplasmic partitioning of plant photoreceptors phytochromes. Semin. Cell Dev. Biol. 11, 505–510 (2000)
Kircher, S. et al. Nucleocytoplasmic partitioning of the plant photoreceptors phytochrome A, B, C, D, and E is regulated differentially by light and exhibits a diurnal rhythm. Plant Cell 14, 1541–1555 (2002)
Ni, M., Tepperman, J. M. & Quail, P. H. PIF3, a phytochrome-interacting factor necessary for normal photoinduced signal transduction, is a novel basic helix-loop-helix protein. Cell 95, 657–667 (1998)
Ni, M., Tepperman, J. M. & Quail, P. H. Binding of phytochrome B to its nuclear signalling partner PIF3 is reversibly induced by light. Nature 400, 781–784 (1999)
Martínez-García, J. F., Huq, E. & Quail, P. H. Direct targeting of light signals to a promoter element-bound transcription factor. Science 288, 859–863 (2000)
Park, C.-M., Bhoo, S.-H. & Song, P.-S. Inter-domain crosstalk in the phytochrome molecules. Semin. Cell Dev. Biol. 11, 449–456 (2000)
Yeh, K.-C. & Lagarias, J. C. Eukaryotic phytochromes: light-regulated serine/threonine protein kinases with histidine kinase ancestry. Proc. Natl Acad. Sci. USA 95, 13976–13981 (1998)
Wagner, D. & Quail, P. H. Mutational analysis of phytochrome B identifies a small COOH-terminal domain region critical for regulatory activity. Proc. Natl Acad. Sci. USA 92, 8596–8600 (1995)
Sakamoto, K. & Nagatani, A. Nuclear localization activity of phytochrome B. Plant J. 10, 859–868 (1996)
Reed, J. W., Nagpal, P., Poole, D. S., Furuya, M. & Chory, J. Mutations in the gene for the red/far-red light receptor phytochrome B alter cell elongation and physiological responses throughout Arabidopsis development. Plant Cell 5, 147–157 (1993)
Wagner, D., Koloszvari, M. & Quail, P. H. Two small spatially distinct regions of Phytochrome B are required for efficient signaling rates. Plant Cell 8, 859–871 (1996)
Grebenok, R. J. et al. Green-fluorescent protein fusions for efficient characterization of nuclear targeting. Plant J. 11, 573–586 (1997)
Kato, A., Hayashi, M. & Nishimura, M. Oligomeric proteins containing N-terminal targeting signals are imported into peroxisomes in transgenic Arabidopsis. Plant Cell Physiol. 40, 586–591 (1999)
Cherry, J. R. et al. Carboxy-terminal deletion analysis of oat phytochrome A reveals the presence of separate domains required for structure and biological activity. Plant Cell 5, 565–575 (1993)
Wagner, D., Fairchild, C. D., Kuhn, R. M. & Quail, P. H. Chromophore-bearing NH2-terminal domains of phytochromes A and B determine their photosensory specificity and differential light lability. Proc. Natl Acad. Sci. USA 93, 4011–4015 (1996)
Clough, R. C. et al. Sequences within both the N- and C-terminal domains of phytochrome A are required for PFR ubiquitination and degradation. Plant J. 17, 155–167 (1999)
Shimizu-Sato, S., Huq, E., Tepperman, J. M. & Quail, P. H. A light-switchable gene promoter system. Nature Biotechnol. 10, 1041–1044 (2002)
Stockhaus, J. et al. Serine-to-alanine substitutions at the amino-terminal region of phytochrome A result in an increase in biological activity. Genes Dev. 6, 2364–2372 (1992)
Liu, X. L., Covington, M. F., Fankhauser, C., Chory, J. & Wagner, D. R. ELF3 encodes a circadian clock-regulated nuclear protein that functions in an Arabidopsis PHYB signal transduction pathway. Plant Cell 13, 1293–1304 (2001)
Somers, D. E., Schultz, T. F., Milnamow, M. & Kay, S. A. ZEITLUPE encodes a novel clock-associated PAS protein from Arabidopsis. Cell 101, 319–329 (2000)
Jarillo, J. A. et al. An Arabidopsis circadian clock component interacts with both CRY1 and phyB. Nature 410, 487–490 (2001)
Fankhauser, C. et al. PKS1, a substrate phosphorylated by phytochrome that modulates light signaling in Arabidopsis. Science 284, 1539–1541 (1999)
Shacklock, P. S., Read, N. D. & Trewavas, A. J. Cytosolic free calcium mediates red light-induced photomorphogenesis. Nature 358, 753–755 (1992)
Mochizuki, N., Brusslan, J. A., Larkin, R., Nagatani, A. & Chory, J. Arabidopsis genomes uncoupled 5 (GUN5) mutant reveals the involvement of Mg-chelatase H subunit in plastid-to-nucleus signal transduction. Proc. Natl Acad. Sci. USA 98, 2053–2058 (2001)
Kalderon, D., Roberts, B. L., Richardson, W. D. & Smith, A. E. A short amino acid sequence able to specify nuclear location. Cell 39, 499–509 (1984)
Wen, W., Meinkoth, J. L., Tsien, R. Y. & Taylor, S. S. Identification of a signal for rapid export of proteins from the nucleus. Cell 82, 463–473 (1995)
Clough, S. J. & Bent, A. F. Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J. 16, 735–743 (1998)
Shinomura, T. et al. Action spectra for phytochrome A- and B-specific photoinduction of seed germination in Arabidopsis thaliana. Proc. Natl Acad. Sci. USA 93, 8129–8133 (1996)
Acknowledgements
We thank S. Kaihara, R. W. Ridge and T. Takeyama for critical reading of the manuscript; R. Tsugeki for providing GUS clone; and laboratory members for support and discussion. This work was supported by grants from the Program for Promotion of Basic Research Activities for Innovative Biosciences and from Special Coordination Funds for Promoting Science and Technology from the Science and Technology Agency and by Grants-in-Aid for Scientific Research (B) and for Scientific Research on Priority Areas from the Ministry of Education, Science, Sports and Culture of Japan.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare that they have no competing financial interests.
Rights and permissions
About this article
Cite this article
Matsushita, T., Mochizuki, N. & Nagatani, A. Dimers of the N-terminal domain of phytochrome B are functional in the nucleus. Nature 424, 571–574 (2003). https://doi.org/10.1038/nature01837
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature01837
- Springer Nature Limited
This article is cited by
-
Distinguishing individual photobodies using Oligopaints reveals thermo-sensitive and -insensitive phytochrome B condensation at distinct subnuclear locations
Nature Communications (2024)
-
Stomatal regulators are co-opted for seta development in the astomatous liverwort Marchantia polymorpha
Nature Plants (2023)
-
A base substitution in OsphyC disturbs its Interaction with OsphyB and affects flowering time and chlorophyll synthesis in rice
BMC Plant Biology (2022)
-
Plant phytochrome B is an asymmetric dimer with unique signalling potential
Nature (2022)
-
Direct photoresponsive inhibition of a p53-like transcription activation domain in PIF3 by Arabidopsis phytochrome B
Nature Communications (2021)