Abstract
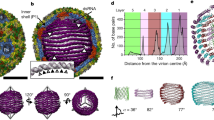
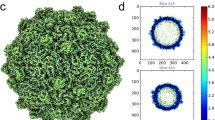
The 3.0 Å resolution crystal structure of Pariacoto virus (PaV) reveals extensive interactions between portions of the viral RNA genome and the icosahedral capsid. Under the protein shell of the T = 3 quasi equivalent capsid lies a dodecahedral cage composed of RNA duplex that accounts for ∼35% of the single-stranded RNA genome. The highly basic N-terminal regions (residues 7–54) of the subunits, forming pentamers (A subunits) are clearly visible in the density map and make numerous interactions with the RNA cage. The C-terminal segments (residues 394–401) of the A subunits lie in channels near the quasi three-fold axes. Electron cryo-microscopy and image reconstruction of PaV particles clearly show the dodecahedral RNA cage.






Similar content being viewed by others
Accession codes
References
Zeddam, J.-L., Rodriguez, J.L., Ravallec, M. & Lagnaoui, A. A noda-like virus isolated from the sweet potato pest Spodoptera eridania (Cramer) (Lep.; Noctuidae). J. Inv. Pathol. 74, 267–274 (1999).
Johnson, K.N., Zeddam, J.-L. & Ball, L.A. Characterization and construction of functional cDNA clones of Pariacoto virus, the first alphanodavirus isolated outside Australasia. J. Virol. 74, 5123–5132 (2000).
Schneemann, A., Reddy, V. & Johnson, J.E. The structure and function of nodavirus particles: a paradigm for understanding chemical biology. Adv. Virus Res. 50, 381–446 (1998).
Schneemann, A., Zhong, W., Gallagher, T.M. & Rueckert, R.R. Maturation cleavage required for infectivity of a nodavirus. J. Virol. 66, 6728–6734 (1992).
Cheng, R.H. et al. Functional implications of quasi-equivalence in a T=3 icosahedral animal virus established by cryo-electron microscopy and X-ray crystallography. Structure 2, 271–282 (1994).
Bong, D.T., Steinem, C., Janshoff, A., Johnson, J.E. & Ghadiri, M.R. A highly membrane-active peptide in Flock House virus: implications for the mechanism of nodavirus infection. Chem. Biol. 6, 473–481 (1999).
Fisher, A.J. & Johnson, J.E. Ordered duplex RNA controls capsid architecture in an icosahedral animal virus. Nature 361, 176–179 (1993).
Wery, J.-P., Reddy, V.S., Hosur, M.V. & Johnson, J.E. The refined three-dimensional structure of an insect virus at 2.8 Å resolution. J. Mol. Biol. 235, 565–586 (1994).
Dasgupta, R., Selling, B. & Rueckert, R. Flock House virus: a simple model for studying persistent infection in cultured Drosophila cells. Arch. Virol. (Suppl.) 9, 121–132 (1994).
Abad-Zapatero, C. et al. Structure of southern bean mosaic virus at 2.8 Å Resolution. Nature 286, 33–39 (1980).
Harrison, S.C., Olson, A.J., Schutt, C.E., Winkler, F.K. & Bricogne, G. Tomato bushy stunt virus at 2.9 Å resolution. Nature 276, 368–373 (1978).
Dong, X.F., Natarajan, P., Tihova, M., Johnson, J.E. & Schneemann, A. Particle polymorphism caused by deletion of a peptide molecular switch in a quasiequivalent icosahedral virus. J. Virol. 72, 6024–6033 (1998).
Schneemann, A. & Marshall D. Specific encapsidation of nodavirus RNAs is mediated through the C terminus of capsid precursor protein alpha. J. Virol. 72, 8738–8746 (1998).
Reddy, V.S. et al. Energetics of quasiequivalence: computational analysis of protein-protein interactions in icosahedral viruses. Biophys. J. 74, 546–558 (1998).
Zhong, W., Dasgupta, R. & Rueckert, R. Evidence that the packaging signal for nodaviral RNA2 is a bulged stem-loop. Proc. Natl. Acad. Sci. USA 89, 11146–11150 (1992).
Chow, M. et al. Myristylation of picornavirus capsid protein VP4 and its structural significance. Nature 327, 482–486 (1987).
Otwinowski, Z. Data collection and processing. In Proceedings of the CCP4 study weekend: data collection and processing (eds Sawyer, L., Isaacs, N. & Baily, S.) 56–62 (Science and Engineering Research Council, Daresbury Laboratory, Daresbury, UK; 1993).
Tong, L. & Rossmann, M.G. The locked rotation function. Acta Crystallogr. A 46, 783–792 (1990).
Jones, T.A. A set of averaging programs. In Proceedings of the CCP4 study weekend: molecular replacement (eds Dodson, E.J., Glover, S. & Wolf, W.) 91–105 (Science and Engineering Research Council, Daresbury Laboratory, Daresbury, UK; 1992).
Collaborative Computational Project, Number 4. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D 50, 760–763 (1994).
Kleywegt, G.J. & Jones, T.A. Halloween . . . masks and bones. In Proceedings of the CCP4 study weekend: from first map to final model (eds Bailey, S., Hubbard, R. & Waller, D.) 56–66 (Science and Engineering Research Council, Daresbury Laboratory, Daresbury, UK; 1994).
Jones, T.A., Zou, J.-Y., Cowan, S.W. & Kjeldgaard, M. Improved methods for building protein models in electron density maps and the location of errors in these models. Acta Crystallogr. A 47, 110–119 (1991).
Brünger, A.T. X-PLOR version 3.0 (Yale University, New Haven, Connecticut; 1992).
Engh, R.A. & Huber, R. Accurate bond and angle parameters for X-ray protein structure refinement. Acta Crystallog. A 47, 392–400 (1991).
Parkinson, G., Vojtechovsky, J., Clowney, L., Brünger, A.T. & Berman, H.M. New parameters for the refinement of nucleic acid-containing structures. Acta Crystallog. D 52, 57–64 (1996).
Frank, J. et al. SPIDER and WEB: processing and visualization of images in 3D electron microscopy and related fields. J. Struct. Biol. 116, 190–199 (1996).
Kraulis, P.J. MOLSCRIPT: a program to produce both detailed and schematic plots of protein structures. J. Appl. Crystallogr. 24, 946–950 (1991).
Esnouf, R.M. An extensively modified version of MolScript that includes greatly enhanced coloring capacities. J. Mol. Graph. Model. 15, 132–134, 112–113 (1997).
Merritt, E.A. & Murphy, M.E.P. Raster3D version 2.0. A program for photorealistic molecular graphics. Acta Crystallogr. D 50, 869–873 (1994).
Nicholls, A., Sharp, K.A. & Honig, B. Protein folding and association: insights from the interfacial and thermodynamic properties of hydrocarbons. Proteins 11, 281–296 (1991).
Acknowledgements
We thank the staff at The Stanford Synchrotron Radiation Laboratory (SSRL) and at the BioCARS sector of The Advanced Photon Source (APS) for help in data collection, M. Tihova and K. Dryden for help in electron microscopy, V.S. Reddy for stimulating discussion, and A. Palmenberg and J.-Y. Sgro for RNA secondary structure predictions. This work was supported by NIH grants to L.A.B., J.E.J. and M.Y. During this work M.Y. was an Established Investigator of the American Heart Association and Bristol-Myers Squibb and is now a recipient of a Clinical Scientist Award in Translational Research from the Burroughs Wellcome Fund.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Tang, L., Johnson, K., Ball, L. et al. The structure of Pariacoto virus reveals a dodecahedral cage of duplex RNA. Nat Struct Mol Biol 8, 77–83 (2001). https://doi.org/10.1038/83089
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/83089
- Springer Nature America, Inc.
This article is cited by
-
Physics of viral dynamics
Nature Reviews Physics (2021)
-
Chimeric virus-like particles (VLPs) designed from shrimp nodavirus (MrNV) capsid protein specifically target EGFR-positive human colorectal cancer cells
Scientific Reports (2021)
-
Reassessing the mechanism of genome packaging in plant viruses with lessons from ATPase fold
Australasian Plant Pathology (2021)
-
Structural puzzles in virology solved with an overarching icosahedral design principle
Nature Communications (2019)
-
Supramolecular Kandinsky circles with high antibacterial activity
Nature Communications (2018)