Abstract
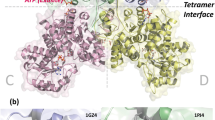
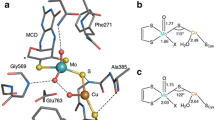
Malic enzymes are widely distributed in nature and have many biological functions. The crystal structure of human mitochondrial NAD(P)+-dependent malic enzyme in a quaternary complex with NAD+, Mn++ and oxalate has been determined at 2.2 Å resolution. The structures of the quaternary complex with NAD+, Mg++, tartronate or ketomalonate have been determined at 2.6 Å resolution. The structures show the enzyme in a closed form in these complexes and reveal the binding modes of the cation and the inhibitors. The divalent cation is coordinated in an octahedral fashion by six ligating oxygens, two from the substrate/inhibitor, three from Glu 255, Asp 256 and Asp 279 of the enzyme, and one from a water molecule. The structural information has significant implications for the catalytic mechanism of malic enzymes and identifies Tyr 112 and Lys 183 as possible catalytic residues. Changes in tetramer organization of the enzyme are also observed in these complexes, which might be relevant for its cooperative behavior and allosteric control.




Similar content being viewed by others
References
Loeber, G., Infante, A.A., Maurer-Fogy, I., Krystek, E. & Dworkin, M.B. Human NAD+-dependent mitochondrial malic enzyme. J. Biol. Chem. 266, 3016– 3021 (1991).
Mallick, S., Harris, B.G. & Cook, P.F. Kinetic mechanism of NAD:malic enzyme from Ascaris suum in the direction of reductive carboxylation. J. Biol. Chem. 266, 2732–2738 ( 1991).
Chou, W.-Y., Huang, S.-M., Liu, Y. H. & Chang, G.-G. Cloning and expression of pigeon liver cytosolic NADP+-dependent malic enzyme cDNA and some of its abortive mutants. Arch. Biochem. Biophys. 310, 158–166 (1994).
Urbauer, J.L., Bradshaw, D.E. & Cleland, W.W. Determination of the kinetic and chemical mechanism of malic enzyme using (2R,3R)-erythro-fluoromalate as a slow alternative substrate . Biochemistry 37, 18026– 18031 (1998).
Xu, Y.W., Bhargava, G., Wu, H., Loeber, G. & Tong, L. Crystal structure of human mitochondrial NAD(P)+-dependent malic enzyme: a new class of oxidative decarboxylases. Structure 7, 877–889 ( 1999).
Loeber, G., Dworkin, M.B., Infante, A. & Ahorn, H. Characterization of cytosolic malic enzyme in human tumor cells. FEBS Lett. 344, 181–186 (1994).
Baggetto, L.G. Deviant energetic metabolism of glycolytic cancer cells. Biochimie 74, 959–974 ( 1992).
McKeehan, W.L. Glycolysis, glutaminolysis and cell proliferation. Cell Biol. Int. Rep. 6, 635–650 ( 1982).
Sauer, L.A. An NAD- and NADP-dependent malic enzyme with regulatory properties. Biochem. Biophys. Res. Commun. 50, 524– 531 (1973).
Wierenga, R.K., Terpstra, P. & Hol, W.G.J. Prediction of the occurrence of the ADP-binding bab-fold in proteins, using an amino acid sequence fingerprint. J. Mol. Biol. 187, 101–107 ( 1986).
Wei, C.-H., Chou, W.-Y., Huang, S.-M., Lin, C.-C. & Chang, G.-G. Affinity cleavage at the putative metal-binding site of pigeon liver malic enzyme by the Fe2+-ascorbate system . Biochemistry 33, 7931– 7936 (1994).
Wei, C.-H., Chou, W.-Y. & Chang, G.-G. Identification of Asp258 as the metal coordinate of pigeon liver malic enzyme by site-specific mutagenesis. Biochemistry 34, 7949–7954 ( 1995).
Satterlee, J. & Hsu, R.Y. Duck liver malic enzyme: sequence of a tryptic peptide containing the cysteine residue labeled by the substrate analog bromopyruvate. Biochim. Biophys. Acta 1079, 247–252 (1991).
Vernon, C.M. & Hsu, R.Y. Pigeon liver malic enzyme: involvement of an arginyl residue at the binding site for malate and its analogs. Arch. Biochem. Biophys. 225, 296–305 (1983).
Rao, S.R., Kamath, B.G. & Bhagwat, A.S. Chemical modification of the functional arginine residue(s) of malic enzyme from Zea mays. Phytochem. 30 , 431–435 (1991).
You, K.S. Stereospecificity for nicotinamide nucleotides in enzymatic and chemical hydride transfer reactions. CRC Crit. Rev. Biochem. 17, 313–451 (1985).
Hall, M.D. & Banaszak, L.J. Crystal structure of a ternary complex of Escherichia coli malate dehydrogenase citrate and NAD at 1.9 Å resolution. J. Mol. Biol. 232, 213–222 (1993).
Hsu, R.Y., Mildvan, A.S., Chang, G.-G. & Fung, C.-H. Mechanism of malic enzyme from pigeon liver. Magnetic resonance and kinetic studies of the role of Mn2+. J. Biol. Chem. 251, 6574–6583 (1976).
Tipton, P.A., Quinn, T.P., Peisach, J. & Cook, P.F. Role of the divalent metal ion in the NAD:malic enzyme reaction: An ESEEM determination of the ground state conformation of malate in the E:Mn:malate complex. Prot. Sci. 5, 1648–1654 (1996).
Hurley, J.H., Dean, A.M., Koshland, D.E. Jr & Stroud, R.M. Catalytic mechanism of NADP+-dependent isocitrate dehydrogenase: Implications from the structures of Magnesium-isocitrate and NADP+ complexes. Biochemistry 30, 8671–8678 (1991).
Lee, M.E., et al. Mutational analysis of the catalytic residue Lysine 230 and Tyrosine 160 in the NADP+-dependent isocitrate dehydrogenase from Escherichia coli. Biochemistry 34, 378– 384 (1995).
Scrutton, N.S., Berry, A. & Perham, R.N. Redesign of the coenzyme specificity of a dehydrogenase by protein engineering. Nature 343, 38– 43 (1990).
Bhargava, G., et al. Preliminary crystallographic studies of human mitochondrial NAD(P)+-dependent malic enzyme. J. Struct. Biol. 127 , 72–75 (1999).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Method Enzymol. 276, 307–326 ( 1997).
Tong, L. Replace: A suite of computer programs for molecular-replacement calculations . J. Appl. Cryst. 26, 748– 751 (1993).
Brunger, A.T. The X-PLOR manual (Yale University, New Haven, Connecticut, 1992).
Collaborative computational project, Number 4. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D50, 760–763 ( 1994).
Jones, T.A., Zou, J.Y., Cowan, S.W. & Kjeldgaard, M. Improved methods for building protein models in electron density maps and the location of errors in these models. Acta Crystallogr. A47, 110–119 (1991).
Carson, M. Ribbon models of macromolecules. J. Mol. Graphics 5 , 103–106 (1987).
Nicholls, A., Sharp, K.A. & Honig, B. Protein folding and association: insights from the interfacial and thermodynamic properties of hydrocarbons. Proteins 11, 281–296 (1991).
Acknowledgements
We thank C. Ogata for setting up the beam line, Y. Xu and R. Khayat for help with data collection at NSLS, W. W. Cleland for many helpful discussions, and H. Wu for comments on the manuscript. This work was supported by a grant from the National Science Foundation to L.T.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Yang, Z., Floyd, D., Loeber, G. et al. Structure of a closed form of human malic enzyme and implications for catalytic mechanism. Nat Struct Mol Biol 7, 251–257 (2000). https://doi.org/10.1038/73378
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/73378
- Springer Nature America, Inc.
This article is cited by
-
Protein engineering and iterative multimodule optimization for vitamin B6 production in Escherichia coli
Nature Communications (2023)
-
A rotary mechanism for allostery in bacterial hybrid malic enzymes
Nature Communications (2021)
-
Molecular adaptations of NADP-malic enzyme for its function in C4 photosynthesis in grasses
Nature Plants (2019)
-
Functional Roles of Metabolic Intermediates in Regulating the Human Mitochondrial NAD(P)+-Dependent Malic Enzyme
Scientific Reports (2019)
-
dissectHMMER: a HMMER-based score dissection framework that statistically evaluates fold-critical sequence segments for domain fold similarity
Biology Direct (2015)