Abstract
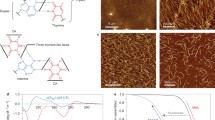
There are many examples of proteins which interact with specific DNA sequences (see references cited in refs 1 and 2). It is not clear whether specificity is derived from proteins sensing directly the base sequence of the DNA or is a result of unique secondary or tertiary structural features induced by certain DNA sequences. Recent studies have indicated the occurrence of several types of symmetries and sequence irregularities in DNA regions which interact with regulatory proteins, but they do not provide information on three-dimensional structure3. The Watson–Crick base-paired double helix can be observed in different conformations4–6, depending on humidity and salt concentration. Within the limited resolution of X-ray fibre diffraction at high relative humidity and in the appropriate salt conditions, studies indicate that natural DNA as well as synthetic repeating polynucleotide duplexes adopt essentially the same average secondary structure, namely the B conformation7. This implies that all residues adopt a single backbone conformation. Recently, however, a high-resolution proton NMR study8 has demonstrated that a change from a regular conformation may occur at the junction region of block copolymers. 31P NMR studies strongly suggest that alternating polynucleotides can have alternating backbone conformations in solution9,10. Klug et al.11 have proposed an alternating structure for poly(dAdT)·poly(dAdT), based on a single crystal X-ray diffraction study of the tetranucleotide, dA-dT-dA-dT (ref. 12). We report here structural 31P NMR studies which show the existence of such variation in the fibre form of a synthetic DNA, poly(dAdT)·poly(dAdT). This confirms our earlier conclusion13 that natural DNA has a non-uniform backbone conformation.
Similar content being viewed by others
References
Wells, R. D. et al. CRC Crit. Rev. Biochem. 4, 305–340 (1977).
Davidson, E. H. & Britten, R. J. Science 204, 1052–1059 (1979).
Dykes, G., Bambara, R., Marians, K. & Wu, R. Nucleic Acids Res. 2, 327–345 (1975).
Langridge, R. et al. J. molec. Biol. 2, 38–64 (1960).
Marvin, D. A., Spencer, M., Wilkins, M. H. F. & Hamilton, L. D. J. molec. Biol. 3, 547–565 (1961).
Fuller, W., Wilkins, M. H. F., Wilson, H. R. & Hamilton, L. D. J. molec. Biol. 12, 60–76 (1965).
Arnott, S., Chandrasekaran, R. & Selsing, E. Structure and Conformation of Nucleic Acids and Protein-Nucleic Acid Interactions (eds Sundralingam, M. & Rao, S. T.) 577–496 (University Park Press, Baltimore, 1975).
Early, T. A., Kearns, D. R., Burd, J. F., Larson, J. E. & Wells, R. D. Biochemistry 16, 541–551 (1977).
Patel, D. J., Canuel, L. L. & Pohl, F. M. Proc. natn. Acad. Sci. U.S.A. 76, 2508–2511 (1979).
Shindo, H., Simpson, R. T. & Cohen, J. S. J. biol. Chem. 254, 8125–8128 (1979).
Klug, A. et al. J. molec. Biol. 131, 669–680 (1979).
Viswamitra, M. A. et al. Nature 273, 687–688 (1978).
Shindo, H., Wooten, J. B., Pheiffer, B. H. & Zimmerman, S. B. Biochemistry (in the press).
Davies, D. R. & Baldwin, R. L. J. molec. Biol. 6, 251–255 (1963).
Lin, S. & Riggs, A. D. Nature 228, 1184–1186 (1970).
Riggs, A. D., Lin, S. & Wells, R. D. Proc. natn. Acad. Sci. U.S.A. 69, 761–764 (1972).
Lin, S. & Riggs, A. D. J. molec. Biol. 72, 671–690 (1972).
Kelsey, D. E., Rounds, T. C. & York, S. S. Proc. natn. Acad. Sci. U.S.A. 2649–2653 (1979).
Scheffler, I. E., Elson, E. L. & Baldwin, R. L. J. molec. Biol. 36, 291–304 (1968).
Zimmerman, S. B. & Pheiffer, B. H. Proc. natn. Acad. Sci. U.S.A. 76, 2703–2707 (1979).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Shindo, H., Zimmerman, S. Sequence-dependent variations in the backbone geometry of a synthetic DNA fibre. Nature 283, 690–691 (1980). https://doi.org/10.1038/283690a0
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/283690a0
- Springer Nature Limited