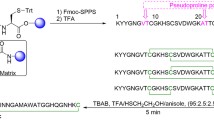
Abstract
IN a previous communication1, from the analysis of a partial acid hydrolysate, the structure of polymyxin E1 was shown to be limited to two alternatives, the so-called 7 α and 8γ formulæ. By applying the enzyme method of hydrolysis as devised by Suzuki et al. in their determinations of the structures of polymyxin B1 (ref. 2) and colistin A (ref. 3), the problem has now been resolved. We have isolated the peptides  and
and  from the hydrolysate of polymyxin E1 with the enzyme Nagarse (supplied by Teikoku Chemical Industry Co., Ltd., Osaka, Japan). The presence in particular of the second peptide eliminates the 8γ variant, and the structure of polymyxin E1 is therefore concluded to be (I; R=(+) 6-methyloctanoyl, X = D-Leu) and is therefore identical with that of colistin A (refs. 3 and 4).
from the hydrolysate of polymyxin E1 with the enzyme Nagarse (supplied by Teikoku Chemical Industry Co., Ltd., Osaka, Japan). The presence in particular of the second peptide eliminates the 8γ variant, and the structure of polymyxin E1 is therefore concluded to be (I; R=(+) 6-methyloctanoyl, X = D-Leu) and is therefore identical with that of colistin A (refs. 3 and 4). 
Similar content being viewed by others
References
Wilkinson, S., and Lowe, L. A., Nature, 204, 185 (1964).
Suzuki, T., Hayashi, K., Fujikawa, K., and Tsukamoto, K., J. Biochem. (Japan), 54, 555 (1963).
Suzuki, T., Hayashi, K., and Fujikawa, K., J. Biochem. (Japan), 54, 412 (1963).
Suzuki, T., Inouye, H., Fujikawa, K., and Nagasawa, S., J. Biochem. (Japan), 54, 173 (1963).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
WILKINSON, S., LOWE, L. Structures of Polymyxin B2 and Polymyxin E1. Nature 204, 993–994 (1964). https://doi.org/10.1038/204993a0
Issue Date:
DOI: https://doi.org/10.1038/204993a0
- Springer Nature Limited
This article is cited by
-
A study of the mechanism of polymyxin M inactivation
Experientia (1968)
-
Structures of the Polymyxins A and the Question of Identity with the Polymyxins M
Nature (1966)