Abstract
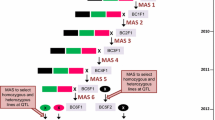
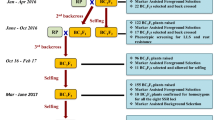
The aim of the present study was to transfer resistance to P. capsici alleles at four quantitative trait loci (QTLs) from a small fruited pepper into a bell pepper recipient line using markers. The marker-assisted selection program was initiated from a doubled-haploid line issued from the mapping population and involved three cycles of marker-assisted backcross (MAB). Two populations, derived by selfing the plants selected after the first selection cycle, were genotyped and evaluated phenotypically for their resistance level. The additive and epistatic effects of the four resistance factors were re-detected and validated in these populations, indicating that introgression of 4 QTLs in this MAB program was successful. A decrease of the effect for the moderate-effect QTLs and of the epistatic interaction was observed. Phenotypic evaluations of horticultural traits were performed on sample of each backcross generation. The results indicated an efficient return to the recipient phenotype using this MAB strategy.
Similar content being viewed by others
References
Basten C.J., Weir B.S., Zeng Z.B. 1997. QTL cartographer: a reference manual and tutorial for QTL mapping. Department of statistics, North Carolina state University, Raleigh, North Carolina, USA.
Ben Chaim A., Paran I., Grube R.C., Jahn M., van Wijk R., Peleman J. 2001. QTL mapping of fruit-related traits in pepper. Theor. Appl. Genet. 102: 1016–1028.
Bouchez A., Hospital F., Causse M., Gallais A., Charcosset A. 2002. Marker-assisted introgression of favorable alleles at quantitative trait loci between maize elite lines. Genetics 162: 1945–1959.
Clerjeau M., Pitrat M., Nourisseau J.G. 1976. La résistance du piment (Capsicum annuum( à Phytophthora capsici. IV. Etude de 19 l'agressivité de divers isolats au niveau des feuilles, des tiges et du collet des plantes résistantes et sensibles. Ann Phytopathol. 8: 411–423.
Dekkers J.C.M., Hospital F. 2002. The use of molecular genetics in the improvement of agricultural populations. Nature Reviews Genetics 3: 22–32.
Frisch M., Melchinger A.E. 2001. Marker-assisted backcrossing for introgression of a recessive gene. Crop. Sci. 41: 1485–1494.
Fulton T.M., Chunwongse J., Tanksley S.D. 1995. Microprep protocol for extraction of DNA of tomato and other herbaceous plants. Plant Mol. Biol. Report. 13(3): 207–209.
Han F., Romagosa I., Ullrich S.E., Jones B.L., Hayes P.M., Wesenberg D.M., 1997. Molecular marker-assisted selection for malting quality traits in barley. Mol. Breed. 3: 427–437.
Hospital F., Charcosset A. 1997. Marker-assisted introgression of quantitative trait loci. Genetics 147: 1469–1485.
Lander E., Green P., Abrahamson J., Barlow A., Daley M., Lincoln S., Newburg L. 1987. Mapmaker: an interactive package for constructing primary genetic linkage maps of experimental and natural populations. Genomics 1: 174–181.
Lawson D.M., Lunde C.F., Mutschler M.A. 1997. Marker-assisted transfer of acylsugar-mediated pest resistance from the wild tomato, Lycopersicon pennellii, to the cultivated tomato, Lycopersicon esculentum. Mol. Breed. 3: 307–317.
Lefebvre V, Palloix A. 1996. Both epistatic and additive effects of QTLs are involved in polygenic induced resistance to disease: a case study, the interaction pepper-Phytophthora capsici Leonian. Theor. Appl. Genet. 93: 503–511.
Lefebvre V., Palloix A., Rives M. 1993. Nuclear RFLP between pepper cultivars (Capsicum annuum L.). Euphytica 71: 189–199.
Lefebvre V., Palloix A., Caranta C., Pochard E. 1995. Construction of an intraspecific integrated linkage map of pepper using molecular markers and doubled-haploid progenies. Genome 38: 112–121.
Lefebvre V., Goffinet B., Chauvet J-C, Caromel B., Signoret P., Brand R., Palloix A. 2001. Evaluation of genetic distances between pepper inbred lines for cultivar protection purposes: comparison of AFLP, RFLP and phenotypic data. Theor. Appl. Genet. 102: 741–750.
Lefebvre V., Pflieger S., Thabuis A., Caranta C., Blattes A., Chauvet J.C., Daubéze A.M., Palloix A. 2002. Towards the saturation of the pepper linkage map by alignment of three intraspecific maps including known-function genes. Genome 45: 839–854.
Melchinger A.E., Utz H.F., Schon C.C. 1998. Quantitative trait locus (QTL( mapping using different testers and independent population samples in maize reveals low power of QTL detection and large bias in estimates of QTL effects. Genetics 149: 383–403.
Palloix A., Daubéze A.M., Pochard E. 1988. Time sequences of root infection and resistance expression in an artificial inoculation method of pepper with Phytophthora capsici. J. Phytopathol. 123: 12–24.
Palloix A., Daubéze A.M., Phaly T., Pochard E. 1990. Breeding transgressive lines of pepper for resistance to Phytophthora capsici in a recurrent selection system. Euphytica 51: 141–150.
Palloix A., Daubéze A.M., Lefebvre V., Caranta C., Moury B., Pflieger S., Gebre-Selassie K., Marchoux G. 1997. Constructing resistant genotypes fitting cultivation conditions in pepper. C. R. Acad. Agric. Fr. 7: 87–98.
Pochard E., Daubéze A.M. 1980. Recherche et évaluation des composantes d'une résistance polygénique: la résistance du piment à Phytophthora capsici. Ann. Amélior. Plant. 26: 377–398.
Pochard E., Clerjeau M., Pitrat M. 1976. La résistance du piment, Capsicum annuum L., à Phytophthora capsici Leon. I. Mise en évidence d'une induction progressive de la résistance. Ann. Amélior. Plant. 26: 35–50.
Robert V.J.M., West M.A.L., Inai S., Caines A., Arntzen L., Smith J.K., St. Clair D.A. 2001. Marker-assisted introgression of blackmold resistance QTL alleles from wild Lycopersicon cheesmanii to cultivated tomato (L. esculentum)and evaluation of QTL phenotypic effects. Mol. Breed. 8: 217–233.
Romagosa I., Han F., Ullrich S.E., Hayes P.M., Wesenberg D.M., 1999. Verification of yield QTL through realized molecular marker-assisted selection responses in a barley cross. Mol. Breed.5: 143–152.
SAS Institute Inc., 1989. SAS/STAT user's guide, version 6, 4th edition. SAS Institute Inc, Cary, North Carolina, USA.
Servin B., Dillman C., Decoux G., Hospital F. 2002. MDM: a program to compute fully informative genotype frequencies in complex breeding schemes. J. Hered. 93(3): 227–228.
Thabuis A., Palloix A., Pflieger S., Daubéze A.M., Caranta C., Lefebvre V., 2003. Comparative mapping of Phytophthora resistance loci in pepper germplasm: evidence for conserved resistance loci across Solanaceae and for a large genetic diversity. Theor. Appl. Genet. 106: 1473–1485.
Toojinda T., Baird E., Booth A., Broers L., Hayes P., Powell W., Thomas W., Vivar H., Young G., 1998. Introgression of quantitative trait loci (QTLs( determining stripe rust resistance in barley: an example of marker-assisted line development. Theor. Appl. Genet. 96: 123–131.
Vos P., Hogers R., Bleeker M., Reijans M,. van de Lee T., Hornes M., Frijters A., Pot J., Peleman J., Kuiper M., Zabeau M. 1995. AFLP: a new technique for DNA fingerprinting. Nucleic. Acid. Research 23(21): 4407–4417.
Young N.D., 1999. A cautiously optimistic vision for marker-assisted breeding. Mol. Breed. 5: 505–510.
Zhu H., Briceno G., Dovel R., Hayes P.M., Liu B.H., Liu C.T., Ullrich S.E., 1999. Molecular breeding for grain yield in barley: an evaluation of QTL effects in a spring barley cross. Theor. Appl. Genet. 98: 772–779.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Thabuis, A., Palloix, A., Servin, B. et al. Marker-assisted introgression of 4 Phytophthora capsici resistance QTL alleles into a bell pepper line: validation of additive and epistatic effects. Molecular Breeding 14, 9–20 (2004). https://doi.org/10.1023/B:MOLB.0000037991.38278.82
Issue Date:
DOI: https://doi.org/10.1023/B:MOLB.0000037991.38278.82