Abstract
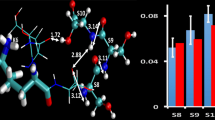
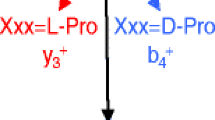
The dynamical behavior of model peptides was evaluated with respect to their ability to form internal proton donor-acceptor pairs using molecular dynamics simulations. The proton donor-acceptor pairs are postulated to be prerequisites for peptide bond cleavage resulting in formation of b and y ions during low-energy collision-induced dissociation in tandem mass spectrometry (MS/MS). The simulations for the polyalanine pentamer Ala5H+ were compared with experimental data from energy-resolved surface induced dissociation (SID) studies. The results of the simulation are insightful into the events that likely lead up to the fragmentation of peptides. Nine-mer polyalanine-based model peptides were used to examine the dynamical effect of each of the 20 common amino acids on the probability to form donor-acceptor pairs at labile peptide bonds. A range of probabilities was observed as a function of the substituted amino acid. However, the location of the peptide bond involved in the donor-acceptor pair plays a critical role in the dynamical behavior. This influence of position on the probability of forming a donor-acceptor pair would be hard to predict from statistical analyses on experimental spectra of aggregate, diverse peptides. In addition, the inclusion of basic side chains in the model peptides alters the probability of forming donor-acceptor pairs across the entire backbone. In this case, there are still more ionizing protons than basic residues, but the side chains of the basic amino acids form stable hydrogen bond networks with the peptide carbonyl oxygens and thus act to prevent free access of “mobile protons” to labile peptide bonds. It is clear from the work that the identification of peptides from low-energy CID using automated computational methods should consider the location of the fragmenting bond as well as the amino acid composition.
Similar content being viewed by others
References
Aebersold, R.; Mann, M. Mass Spectrometry-Based Proteomics. Nature 2003, 422(6928), 198–207.
Smith, R. D.; Anderson, G. A.; Lipton, M. S.; Pasa-Tolic, L.; Shen, Y.; Conrads, T. P.; Veenstra, T. D.; Udseth, H. R. An Accurate Mass Tag Strategy for Quantitative and High-Throughput Proteome Measurements. Proteomics 2002, 2(5), 513–523.
Wolters, D. A.; Washburn, M. P.; Yates, J. R. III. An Automated Multidimensional Protein Identification Technology for Shotgun Proteomics. Anal. Chem. 2001, 73(23), 5683–5690.
Cannon, W. R.; Jarman, K. H.; Webb-Robertson, B.-J.; Baxter, D. J.; Oehmen, C. S.; Jarman, K. D.; Heredia-Langner, A.; Auberry, K. J.; Anderson, D. C. A Comparison of Probability and Likelihood Models for Peptide Identification from Tandem Mass Spectrometry Data. J. Proteome Res. 2005, 4(5), 1687–1698.
Colinge, J.; Masselot, A.; Giron, M.; Dessingy, T.; Magnin, J. OLAV: Towards High-Throughput Tandem Mass Spectrometry Data Identification. Proteomics 2003, 3(8), 1454–1463.
Dancik, V.; Addona, T. A.; Clauser, K. R.; Vath, J. E.; Pevzner, P. A. De Novo Peptide Sequencing Via Tandem Mass Spectrometry. J. Comput. Biol. 1999, 6(3/4), 327–342.
Elias, J. E.; Gibbons, F. D.; King, O. D.; Roth, F. P.; Gygi, S. P. Intensity-Based Protein Identification by Machine Learning from a Library of Tandem Mass Spectra. Nat. Biotechnol. 2004, 22(2), 214–219.
Havilio, M.; Haddad, Y.; Smilansky, Z. Intensity-Based Statistical Scorer for Tandem Mass Spectrometry. Anal. Chem. 2003, 75(3), 435–444.
Sadygov, R.; Wohlschlegel, J.; Park, S. K.; Xu, T.; Yates, J. R. III. Central Limit Theorem as an Approximation for Intensity-Based Scoring Function. Anal. Chem. 2006, 78(1), 89–95.
Sadygov, R. G.; Yates, J. R. A Hypergeometric Probability Model for Protein Identification and Validation Using Tandem Mass Spectral Data and Protein Sequence Databases. Anal. Chem. 2003, 75(15), 3792–3798.
Zhang, Z. Q. Prediction of Low-Energy Collision-Induced Dissociation Spectra of Peptides with Three or More Charges. Anal. Chem. 2005, 77(19), 6364–6373.
Zhang, Z. Q. Prediction of Low-Energy Collision-Induced Dissociation Spectra of Peptides. Anal. Chem. 2004, 76(14), 3908–3922.
Kapp, E. A.; Schutz, F.; Reid, G. E.; Eddes, J. S.; Moritz, R. L.; O’Hair, R. A. J.; Speed, T. P.; Simpson, R. J. Mining a Tandem Mass Spectrometry Database to Determine the Trends and Global Factors Influencing Peptide Fragmentation. Anal. Chem. 2003, 75(22), 6251–6264.
Tabb, D. L.; Smith, L. L.; Breci, L. A.; Wysocki, V. H.; Lin, D.; Yates, J. R., III. Statistical Characterization of Ion Trap Mass Spectra from Doubly Charged Tryptic Peptides. Anal. Chem. 2003, 75(5), 1155–1163.
Huang, Y. Y.; Triscari, J. M.; Tseng, G. C.; Pasa-Tolic, L.; Lipton, M. S.; Smith, R. D.; Wysocki, V. H. Statistical Characterization of the Charge State and Residue Dependence of Low-Energy CID Peptide Dissociation Patterns. Anal. Chem. 2005, 77(18), 5800–5813.
O’Hair, R. A. J. Commentary—The Role of Nucleophile-Electrophile Interactions in the Unimolecular and Bimolecular Gas-Phase Ion Chemistry of Peptides and Related Systems. J. Mass Spectrom. 2000, 35(12), 1377–1381.
Paizs, B.; Suhai, S. Fragmentation Pathways of Protonated Peptides. Mass Spectrom. Rev. 2005, 24(4), 508–548.
Polce, M. J.; Ren, D.; Wesdemiotis, C. Special Feature: Commentary—Dissociation of the Peptide Bond in Protonated Peptides. J. Mass Spectrom. 2000, 35(12), 1391–1398.
Schlosser, A.; Lehmann, W. D. Special Feature: Commentary—Five-Membered Ring Formation in Unimolecular Reactions of Peptides: A Key Structural Element Controlling Low-Energy Collision-Induced Dissociation of Peptides. J. Mass Spectrom. 2000, 35(12), 1382–1390.
Wysocki, V. H.; Tsaprailis, G.; Smith, L. L.; Breci, L. A. Special Feature: Commentary—Mobile and Localized Protons: A Framework for Understanding Peptide Dissociation. J. Mass Spectrom. 2000, 35(12), 1399–1406.
Dongre, A. R.; Jones, J. L.; Somogyi, A.; Wysocki, V. H. Influence of Peptide Composition, Gas-Phase Basicity, and Chemical Modification on Fragmentation Efficiency: Evidence for the Mobile Proton Model. J. Am. Chem. Soc. 1996, 118(35), 8365–8374.
Bailey, T. H.; Laskin, J.; Futrell, J. H. Energetics of Selective Cleavage at Acidic Residues Studied by Time- and Energy-Resolved Surface-Induced Dissociation in FT-ICR MS. Int. J. Mass Spectrom. 2003, 222(1/3), 313–327.
Laskin, J.; Bailey, T. H.; Futrell, J. H. Fragmentation Energetics for Angiotensin II and I Analogs from Time- and Energy-Resolved Surface-Induced Dissociation Studies. Int. J. Mass Spectrom. 2004, 234(1/3), 89–99.
Harrison, A. G.; Yalcin, T. Proton Mobility in Protonated Amino Acids and Peptides. Int. J. Mass Spectrom. 1997, 165, 339–347.
Johnson, R. S.; Krylov, D.; Walsh, K. A. Proton Mobility within Electrosprayed Peptide Ions. J. Mass Spectrom. 1995, 30(2), 386–387.
Mueller, D. R.; Eckersley, M.; Richter, W. J. Hydrogen Transfer-Reactions in the Formation of Y + 2 Sequence Ions from Protonated Peptides. Org. Mass Spectrom. 1988, 23(3), 217–222.
Tsang, C. W.; Harrison, A. G. Chemical Ionization of Amino-Acids. J. Am. Chem. Soc. 1976, 98(6), 1301–1308.
Black, G., Daily, J., Didier, B., Elsethagen, T., Feller, D., Gracio, D., Hackler, M., Havre, S., Jones, D., Jurrus, E., Keller, T., Lansing, C., Matsumoto, S., Palmer, B., Peterson, M., Schuchardt, K., Stephan, E., Sun, L., Swanson, K., Taylor, H., Thomas, G., Vorpagel, E., Windus, T., Winters, C. ECCE, a Problem Solving Environment for Computational Chemistry, Software Version 4.0.2; Pacific Northwest National Laboratory, Richland, Washington 99352-0999, USA. 2006.
Kendall, R.; Apra, E.; Bernholdt, D.; Bylaska, E.; Dupuis, M.; Fann, G.; Harrison, R.; Ju, J.; Nichols, J.; Nieplocha, J.; Straatsma, T.; Windus, T.; Wong, A. High Performance Computational Chemistry: An Overview of NWChem a Distributed Parallel Application. Comput. Phys. Commun. 2000, 128(1/2), 260–283.
Weiner, S. J.; Kollman, P. A.; Nguyen, D. T.; Case, D. A. An All Atom Force-Field for Simulations of Proteins and Nucleic-Acids. J. Comput. Chem. 1986, 7(2), 230–252.
Hunter, E. P. L.; Lias, S. G. Evaluated Gas Phase Basicities and Proton Affinities of Molecules: An Update. J. Phys. Chem. Ref. Data 1998, 27(3), 413–656.
Rakov, V. S.; Denisov, E. V.; Futrell, J. H.; Ridge, D. P. Surface Induced Dissociation of Chromium Hexacarbonyl Ion Fluorinated Alkanethiolate Surface in Ion Cyclotron Resonance Mass Spectrometer: Studies of Energetics of the Process Using Recursive Internal Energy Distribution Search Method. Int. J. Mass Spectrom. 2002, 213(1), 25–44.
Laskin, J.; Denisov, E.; Futrell, J. Comparative Study of Collision-Induced and Surface-Induced Dissociation. 2: Fragmentation of Small Alanine-Containing Peptides in FT-ICR MS. J. Phys. Chem. B 2001, 105(9), 1895–1900.
Laskin, J.; Denisov, E.; Futrell, J. A Comparative Study of Collision-Induced and Surface-Iinduced Dissociation. 1: Fragmentation of Protonated Dialanine. J. Am. Chem. Soc. 2000, 122(40), 9703–9714.
Laskin, J.; Denisov, E.; Futrell, J. H. Fragmentation Energetics of Small Peptides from Multiple-Collision Activation and Surface-Induced Dissociation in FT-ICR MS. Int. J. Mass Spectrom. 2002, 219(1), 189–201.
Paizs, B.; Suhai, S. Towards Understanding the Tandem Mass Spectra of Protonated Oligopeptides. 1: Mechanism of Amide Bond Cleavage. J. Am. Soc. Mass Spectrom. 2004, 15(1), 103–113.
Henry, B.; Tekely, P.; Delpuech, J. J. pH and pK Determinations by High-Resolution Solid-State C-13 NMR: Acid-Base and Tautomeric Equilibria of Lyophilized L-Histidine. J. Am. Chem. Soc. 2002, 124(9), 2025–2034.
Ivanov, I.; Klein, M. L. Deprotonation of a Histidine Residue in Aqueous Solution Using Constrained ab Initio Molecular Dynamics. J. Am. Chem. Soc. 2002, 124(45), 13380–13381.
Huang, Y.; Triscari, J. M.; Tseng, G. C.; Pasa-Tolic, L.; Lipton, M. S.; Smith, R. D.; Wysocki, V. H. Statistical Characterization of the Charge State and Residue Dependence of Low-Energy CID Peptide Dissociation Patterns. Anal. Chem. 2005, 77(18), 5800–5813.
Breci, L. A.; Tabb, D. L.; Yates, J. R. III; Wysocki, V. H. Cleavage N-terminal to Proline: Analysis of a Database of Peptide Tandem Mass Spectra. Anal. Chem. 2003, 75(9), 1963–1971.
Huang, Y.; Triscari, J.; Pasa-Tolic, L.; Anderson, G.; Lipton, M.; Smith, R.; Wysocki, V. Dissociation Behavior of Doubly-Charged Tryptic Peptides: Correlation of Gas-Phase Cleavage Abundance with Ramachandran Plots. J Am. Chem. Soc. 2004, 126(10), 3034–3035.
Bouchoux, G.; Salpin, J. Y. Gas-Phase Basicity of Glycine, Alanine, Proline, Serine, Lysine, Histidine, and Some of Their Peptides by the Thermokinetic Method. Eur. J. Mass Spectrom. 2003, 9(4), 391–402.
Tsaprailis, G.; Nair, H.; Zhong, W.; Kuppannan, K.; Futrell, J. H.; Wysocki, V. H. A Mechanistic Investigation of the Enhanced Cleavage at Histidine in the Gas-Phase Dissociation of Protonated Peptides. Anal. Chem. 2004, 76(7), 2083–2094.
Craig, R.; Cortens, J. C.; Fenyo, D.; Beavis, R. C. Using Annotated Peptide Mass Spectrum Libraries for Protein Identification. J. Proteome Res. 2006, 5(8), 1843–1849.
Frewen, B. E.; Merrihew, G. E.; Wu, C. C.; Noble, W. S.; MacCoss, M. J. Analysis of Peptide MS/MS Spectra from Large-Scale Proteomics Experiments Using Spectrum Libraries. Anal. Chem. 2006, 78(16), 5678–5684.
Lam, H.; Deutsch, E.; Eddes, J. S.; Eng, J. K.; King, N.; Stein, S.; Aebersold, R. Development and Validation of a Spectral Library Searching Method for Human Peptide Identification from Tandem Mass Spectrometry. Mol. Cell. Proteom. 2006, 5(10), S361-S361.
Author information
Authors and Affiliations
Corresponding author
Additional information
Published online June 20, 2007
Rights and permissions
About this article
Cite this article
Cannon, W.R., Taasevigen, D., Baxter, D.J. et al. Evaluation of the influence of amino acid composition on the propensity for collision-induced dissociation of model peptides using molecular dynamics simulations. J Am Soc Mass Spectrom 18, 1625–1637 (2007). https://doi.org/10.1016/j.jasms.2007.06.005
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1016/j.jasms.2007.06.005