Abstract
Background
Traditional adverse event (AE) reporting systems have been slow in adapting to online AE reporting from patients, relying instead on gatekeepers, such as clinicians and drug safety groups, to verify each potential event. In the meantime, increasing numbers of patients have turned to social media to share their experiences with drugs, medical devices, and vaccines.
Objective
The aim of the study was to evaluate the level of concordance between Twitter posts mentioning AE-like reactions and spontaneous reports received by a regulatory agency.
Methods
We collected public English-language Twitter posts mentioning 23 medical products from 1 November 2012 through 31 May 2013. Data were filtered using a semi-automated process to identify posts with resemblance to AEs (Proto-AEs). A dictionary was developed to translate Internet vernacular to a standardized regulatory ontology for analysis (MedDRA®). Aggregated frequency of identified product-event pairs was then compared with data from the public FDA Adverse Event Reporting System (FAERS) by System Organ Class (SOC).
Results
Of the 6.9 million Twitter posts collected, 4,401 Proto-AEs were identified out of 60,000 examined. Automated, dictionary-based symptom classification had 72 % recall and 86 % precision. Similar overall distribution profiles were observed, with Spearman rank correlation rho of 0.75 (p < 0.0001) between Proto-AEs reported in Twitter and FAERS by SOC.
Conclusion
Patients reporting AEs on Twitter showed a range of sophistication when describing their experience. Despite the public availability of these data, their appropriate role in pharmacovigilance has not been established. Additional work is needed to improve data acquisition and automation.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Existing post-marketing adverse event surveillance systems suffer from under-reporting and data processing lags. |
Social media services such as Twitter are seeing increasing adoption, and patients are using them to describe adverse experiences with medical products. |
An analysis of 4,401 of these ‘posts with resemblance to adverse events’ (‘Proto-AEs’) from Twitter found concordance with consumer-reported FDA Adverse Event Reporting System reports at the System Organ Class level. |
1 Introduction
Pharmacovigilance systems rely on spontaneous adverse event (AE) reports received by regulatory authorities, mostly after passing through third-party gatekeepers (biopharmaceutical industry, lawyers, clinicians, and pharmacists), generating possible bottlenecks where information may be lost or misinterpreted. In the USA, 80 % of drug [1], 35 % of vaccine [2], and 98 % of device [3] AE reports received by the US FDA come from the biopharmaceutical industry. In the clinical context, while the true extent of under-reporting is unknown, a recent study of Medicare enrollees in a hospital found that 86 % of AEs went unreported [4]. Meanwhile, the government currently releases the data approximately 1 year after receipt [5]. This systemic friction results in an important reporting and information gap that has persisted for decades. Social media data may provide additional insight when combined with additional information sources in modern pharmacovigilance systems, including those that can provide a denominator for rate calculations, such as electronic health records and administrative claims databases.
Regulatory authorities have recognized the importance of listening to the patient’s voice at some level, requiring the collection of patient-reported AEs from online sources [6] and providing guidance on using patient-reported outcomes [7]. The most explicit direction in this area to date has come from the Association of the British Pharmaceutical Industry, which issued voluntary guidelines for how to approach AEs reported in third-party social media sites [8]. Their recommendation is “if a company chooses to ‘listen in’ at non-company-sponsored sites, it is recommended that the relevant pages of the site should be monitored for AE/PC (product complaint) for the period of the listening activity only.”
Despite these initial efforts, the information gap also extends in the opposite direction. As medical product safety concerns emerge, data collected through health institutions and official reporting structures may not be available for months or even years, hindering timely pharmacoepidemiological assessment and public awareness. In the meantime, most consumers are unlikely to be aware that they can and should report AEs.
At the same time that there is significant under-reporting of AEs through official channels, new Internet services have given voice to patients who routinely share information in public forums, including their experiences with medical products. Approximately 25 % of Facebook [9] profiles and 90 % of Twitter [10] feeds are fully public, and a broad range of health-focused forums support public discussions. Previous studies have suggested the potential of high-quality data generated by online social networks at low cost [11–13]. Even users’ search engine query histories have also been used to identify AEs [14]. Many users also report AEs publicly, often expecting that someone is paying attention, as evidenced by hashtags for regulatory agencies (e.g., #FDA), manufacturers (#Pfizer, #GSK), and specific products (#accutaneprobz, @EssureProblems). (The hashtag [e.g., #accutaneprobz] placed in the body of a post is a way to categorize or tag the post to allow for quick retrieval via subsequent searching, similar conceptually to an email folder; the ‘at’ sign [@] placed in front of a Twitter username constitutes a ‘mention’, directing the message to the username in question, similar conceptually to an email address.) However, these data have not yet been used for routine safety surveillance and careful consideration must be given to how to process the information.
In order to assess the feasibility and reliability of harnessing social media data for AE surveillance, we analyzed data from the micro-blogging site Twitter. With posts limited to 140 characters, we expect that micro-blogging data are the smallest units of text in which events can be detected at present. We compared posts mentioning AEs detected in Twitter with public data derived from the US FDA Adverse Event Reporting System (FAERS).
2 Methods
Our approach was to first collect all English-language Twitter posts mentioning medical products. We applied manual and semi-automated techniques to identify posts with resemblance to AEs. Colloquial language was then mapped to a standard regulatory dictionary. We then compared the aggregate frequency of identified product-event pairs with FAERS at the organ system level. A data collection schematic is shown in Fig. 1.
Data collection scheme for both Twitter and FAERS reports. API application programming interface, FAERS FDA Adverse Event Reporting System, FDA Food and Drug Administration, MedDRA Medical Dictionary for Regulatory Activities, NLP natural language processing, Proto-AEs posts with resemblance to adverse events
2.1 Data Sources
Twitter data were collected from 1 November 2012 through 31 May 2013, and consisted of public posts acquired through the general-use streaming application programming interface (API). We chose this data source because it contains a large volume of publicly available posts about medical products. Data were stored in databases using Amazon Web Services (AWS) cloud services. Since the vast majority of the over 400 million daily Twitter posts have no relevance to AE reporting, we created a list of medical product names and used them as search term inputs to the Twitter API. Though this approach may remove posts that contain misspellings, slang terms, and other oblique references, it allowed us to start from a manageable data corpus. In order to avoid confusion with regulatory definitions of an ‘adverse event’ report, the term ‘Proto-AE’ was coined to signify ‘posts with resemblance to AEs’, designating posts containing discussion of AEs identified in social media sources. The labeled subset was chosen through a combination of review of the data in sequence as collected from the API for convenient time periods, as well as by searching the unlabeled data for specific product names and symptom terms.
Public FAERS data were obtained from the FDA website in text format for the time period concurrent with the collection of Twitter data, the fourth quarter of 2012 and first quarter of 2013.
2.2 Product Selection
In conjunction with the FDA, a priori selected 23 prescription and over-the-counter drug products in diverse therapeutic areas were selected for quantitative analysis, representing new and old medicines, as well as widely used products and more specialized ones: acetaminophen, adalimumab, alprazolam, citalopram, duloxetine, gabapentin, ibuprofen, isotretinoin, lamotrigine, levonorgestrel, metformin, methotrexate, naproxen, oxycodone, paroxetine, prednisone, pregabalin, sertraline, tramadol, varenicline, venlafaxine, warfarin, and zolpidem. We also selected vaccines for influenza, human papillomavirus (HPV), hepatitis B, and the combined tetanus/diphtheria/pertussis (Tdap) vaccine. Whenever possible, we identified brand and generic names for each product using the DailyMed site from the National Library of Medicine and the FDA’s Orange Book.
2.3 Adverse Event (AE) Identification in Twitter
The next step was classification of the information, which includes filtering the corpus to remove items irrelevant to AEs. To determine whether or not a given post constitutes an AE report, we established guidelines for human annotators to consistently identify AE reports. We proceeded under the general guidance of the four statutorily required data elements for AE reporting in the USA: an identifiable medical product, an identifiable reporter, an identifiable individual, and mention of a negative outcome, though we did not automatically exclude posts if they failed to meet one of these criteria. We also considered Twitter accounts as sufficient to meet the requirement for an identifiable reporter, though this standard is not the current regulatory expectation for mandatory reporting.
We applied a tree-based dictionary-matching algorithm to identify both product and symptom mentions. It consists of three components. First, we loaded the dictionary from a multi-user editable spreadsheet form into the tree structure in memory. Two separate product and symptom dictionaries were superimposed into a single tree, allowing only a single pass over the input for both product and symptom matches. Second, for the extraction step, a tokenizer stripped punctuation and split the input into a series of tokens, typically corresponding to words. Finally, we processed the tokens one at a time, matching each against the tree and traversing the tree as matches occurred. If we reached a leaf in the tree, then a positive match was established and we returned the identifier for the appropriate concept (product or symptom). Because the concordance analysis is based on the output of the algorithm, its performance characteristics are important. We assessed the performance of the symptom classifier by manually examining a random sample of 10 % of the Proto-AEs and comparing the algorithmically identified symptoms with symptoms that a rater (CCF) determined to be attributed to a product.
We created a curation tool for reviewing and labeling posts. Two trained raters (CCF, CMM) classified a convenience sample of 61,402 posts. Discrepancies between raters were adjudicated by three of the authors (CCF, CMM, ND). Agreement on overlapped subsets increased from 97.9 to 98.4 % (Cohen’s kappa: 0.97) over successive rounds of iterative protocol development and classification. The convenience sample was selected as a training dataset for further development of an automated Bayesian classifier, but the classifier was not used in the analysis presented in this study. The sample was enriched to include posts that contained AEs based on preliminary data review.
2.4 Coding of AEs in Twitter
Further natural language processing was required to identify the event in each post. Starting with the subset of posts identified to contain AEs, we developed a dictionary to convert Internet vernacular to a standardized regulatory dictionary, namely Medical Dictionary for Regulatory Activities (MedDRA®) version 16 in English. MedDRA®, the Medical Dictionary for Regulatory Activities, terminology is the international medical terminology developed under the auspices of the International Conference on Harmonization of Technical Requirements for Registration of Pharmaceuticals for Human Use (ICH). MedDRA® trademark is owned by the International Federation of Pharmaceutical Manufacturers and Associations (IFPMA) on behalf of ICH. The ontology matches Internet vernacular to the closest relevant MedDRA preferred term, but allows for less specific higher-level terms to be used when not enough detail is available for matching to a preferred term. The empirically derived dictionary currently contains over 4,800 terms spread across 257 symptom categories. For example, the post “So much for me going to sleep at 12. I am wide awake thanks prednisone and albuterol” (30 Sep 2013), would be coded to the MedDRA preferred term ‘insomnia’, by identifying ‘wide awake’ as the outcome, and both prednisone and albuterol as the drugs involved. Multiple vernacular phrases could be mapped to the same MedDRA preferred term, such as ‘can’t sleep’ and ‘tossing and turning’ in the previous example. As noted above and detailed in Freifeld et al. [15], we used a tree-based text-matching algorithm to match the raw text from the posts to the vernacular dictionary. Preferred terms were aggregated up to the System Organ Class (SOC), the broadest hierarchical category in MedDRA.
2.5 AE Identification in FAERS
AEs were identified from public FAERS data for the products of interest using exact name matching for brand and generic names. Un-duplication was conducted using the FDA case identification number, date of event, country of occurrence, age, and gender. Reports submitted by consumers were identified using reporter field. All roles (primary suspect, secondary suspect, etc.) were considered; preliminary analysis suggested that limiting to primary suspect medicines did not alter results meaningfully (data not shown).
2.6 Data Analysis
We analyzed vaccines and drugs separately, since vaccine AE data from the US Vaccine Adverse Event Reporting System (VAERS) [16] were not available for this analysis. We compared prescription and over-the-counter drug AEs identified in Twitter posts with corresponding FAERS data for those products, at the SOC level. This approach is intended to identify gross patterns, and not to assess the ability to detect rare but serious AEs.
We assessed the precision and recall of the symptom dictionary-matching algorithm by manual classification of a random sample of 437 Twitter posts (10 % of the full sample). A single post can contain multiple symptom mentions; a symptom match was considered a true positive only if the symptom was considered an AE of one of the mentioned products.
We did not seek to verify each individual report as truthful, but rather to identify overall associations between Twitter and official spontaneous report data as a preliminary proof of concept. We calculated correlation using the Spearman correlation rank statistic (rho) by the 22 SOC categories, with a statistical alpha of <0.05.
3 Results
The resulting dataset contained a high volume of irrelevant information, but provided a useful starting point. Over the period from 1 November 2012 through 31 May 2013, we collected a total of 6.9 million Twitter posts (‘tweets’). Of these, we manually categorized 61,401 as Proto-AEs (4,401) or not (57,000). While the number of Proto-AEs represented 7.2 % of the 61,402 posts analyzed, we reiterate that this was a convenience sample enriched for Proto-AEs and should not be interpreted as the prevalence of AE-related reports in social media. There were 1,400 AEs reported to FAERS from 1 October 2012 through 31 March 2013 for the 23 active ingredients analyzed.
The number of Proto-AEs for each drug were as follows: acetaminophen (303), adalimumab (21), alprazolam (332), citalopram (16), duloxetine (49), gabapentin (32), ibuprofen (1,268), isotretinoin (75), lamotrigine (21), levonorgestrel (57), metformin (22), methotrexate (18), naproxen (85), oxycodone (102), paroxetine (19), prednisone (153), pregabalin (25), sertraline (67), tramadol (213), varenicline (29), venlafaxine (23), warfarin (16), and zolpidem (554).
The performance analysis of the symptom dictionary-matching algorithm based on a random sample of 10 % of AE posts found precision was 72 % (410/573), and recall was 86 % (410/475). The performance of the classifier validates the dictionary-matching approach itself, and also indicates that the symptom counts used in the comparison are close to what would have been derived by a purely manual process.
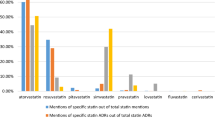
The five pain reliever drugs we analyzed represent multiple functional classes of compounds, over-the-counter (OTC) versus prescription status, and new versus old drugs: tramadol, oxycodone, naproxen, ibuprofen, and acetaminophen. Collectively, the medicines in this broad therapeutic area represent the most commonly mentioned regulated drugs in our Twitter dataset.
3.1 Consumer-Reported AEs in the FDA AE Reporting System (FAERS)
Exactly 1,400 events were reported by consumers to the FDA during the two quarters for the products of interest, corresponding to 1,478 unique drug–event pairs. For 59.4 % of events, the drugs were primary suspect agents; 33.1 % were concomitant, and 7.5 % secondary suspect agents.
3.2 Overall Association between Twitter Posts and FAERS
The preliminary analyses suggested that, when analyzed at preferred-term levels, the sample was too noisy to provide meaningful comparisons with FAERS due to the difference in specificity between Internet vernacular on Twitter and the depth of clinical distinctions in MedDRA. The overall distribution profiles for AEs reported in Twitter and FAERS for drugs were similar when analyzed by SOC (Fig. 2). The Spearman correlation coefficient was 0.75, p < 0.0001 (Fig. 3).
3.3 Vaccine AEs in Twitter and FAERS
We analyzed data for four common types of vaccines: influenza, HPV, Tdap, and hepatitis B. Posts with AEs mentioned in Twitter (posts 460, AEs 634, some posts contained multiple AEs) were similar across vaccines. Most vaccine AEs were associated with influenza vaccines (posts 398, AEs 557), in part because ‘flu shot’ was a search term, whereas other vaccines did not have a corresponding vernacular. The imbalance may also be due to the differences in Twitter use among the populations receiving this vaccine versus others, given that it is an annual vaccine indicated for nearly all individuals aged over 6 months. Other vaccines had many fewer events, as shown in Table 1. The most common AE by preferred term was injection site pain, followed by pain, urticaria, and malaise.
4 Discussion
The recently passed FDA Safety and Innovation Act (FDASIA) legislation (2012) and the FDA-issued report Advancing Regulatory Science at FDA: A Strategic Plan (2011) have emphasized the importance of post-market safety surveillance and called for identification of new sources of post-market safety data. Accordingly, this analysis was intended to evaluate the potential value of burgeoning user-generated social media data in post-market safety surveillance of drugs and biologics.
There were nearly three times as many Proto-AEs found in Twitter data than reported to FDA by consumers, with rank correlation between them at the SOC level. Further, there was evidence that patients intend to passively report AEs in social media, as evidenced by hashtags and mentions such as #accutaneprobz and @EssureProblems. Even within 140 characters, some tweets demonstrate an understanding of basic concepts of causation in drug safety, such as alleviation of the AE after discontinuation of the drug: “I found that Savella made my blood sugar [sic] high, once I got off, my blood sugar returned to normal.”
One of the key aspects of this research was to develop an ontology that allowed translation between social media vernacular and MedDRA, yielding strong automated classification performance as noted above. As an illustrative example of this cross-ontology translation process, we consider the tweet “Humira never really worked for me. Orencia was good. Xeljanz was the best but ate a hole in my stomach. #RABlows.” In this example with rheumatoid arthritis medicines, the first product could be reported as ineffective, but there is also a more serious event. The last product mentioned what could be an exaggeration: “ate a hole in my stomach.” However, the medication guide for this product states “XELJANZ may cause serious side effects including: tears (perforation) in the stomach or intestines.” [17] The example illustrates that context is required to interpret the findings, a task that humans inherently perform better than machines. Initially, identifying that a “hole in the stomach” could be a serious event required previous knowledge of the labeled side effects [18]. We would likely have been less concerned if the post seemed to be complaining about routine gastrointestinal (GI) discomfort after taking aspirin. As such, we believe that this is a task for which humans are, for the time being, best suited. Advanced methods to apply label information may alleviate this particular issue, but we found that a human curation step, after the machine classified the posts, was the most efficient way to understand the nature of the problems reported.
Another illustrative example is paresthesia, often referred to as ‘brain zaps’ or ‘head zaps’. These events have been reported for rapid discontinuation of serotonergic antidepressants as a class [19]. We found ample evidence of the relatively new medicine vilazodone (marketed as Viibryd) from patients: “Second day off Effexor & on Viibryd here. Brain zaps are fun. And by fun, I mean horrendous and miserable.” Also: “Viibryd side effects—I’m having the awful head zaps too. They are mostly @ night when I’m laying down … .” The US prescribing information in the label states that “paresthesia, such as electric shock sensations” have been reported for the class of antidepressants, but during clinical trials only two patients in the treatment arm and one patient on placebo experienced these sensations [20]. While this association is not particularly unexpected, it demonstrates again how knowledge of the label can help put findings from the Internet in context and begins to suggest the rapidity with which information can begin to flow back to the public authorities and manufacturers if social media data are cleaned and curated in a thoughtful manner.
In addition to patients, clinicians may use Twitter to communicate about cases that may involve AEs. For example, the following post was made from the account of an internal medicine resident at a hospital in the USA: “PMHx includes DVT x2 on xaerlto [sic], recurrent tonsillitis, former smoker, denies EtOH, denies surgeries, is married.” The drug Xarelto (rivaroxaban) is indicated for deep vein thrombosis, but it is unclear if the recurrent tonsillitis is associated with the medicine. The US labeling suggests increased incidence of sinusitis in one clinical trial relative to placebo [21], and a post hoc analysis of a phase III randomized trial found increased incidence of respiratory tract or lung infections among participants with higher body mass index (BMI) relative to comparator (enoxaparin) [22]. Without emphasizing the particular drug–event pair, we provide this example to show how information from label studies, peer-reviewed research, and regulatory databases can be used in conjunction with social media to generate hypotheses for further testing. While it may be feasible to review all social media posts with Proto-AEs for lower volume drugs, there is likely to be a point at which the volume of posts, say for a widely used and established medicine, may overwhelm the capacity for human review, requiring further automated analysis. We also stress that, at this time, we do not recommend the wholesale import of individual social media posts into post-marketing safety databases. Rather, in parallel with other post-marketing sources, these data should be considered for idea generation, and reasonable hypotheses followed up with formal epidemiologic studies.
Not all patients reporting side effects on Twitter are capable of identifying them correctly, as seen in this example: “I got the flu shot yet somehow I’ve gotten the stomach flu twice in this month” where the vernacular “stomach flu” would need to be translated into a medical term, and the report was excluded from analysis because the reported ineffectiveness does not correspond to an indication or common off-label use for the vaccine. Other errors may be inadvertent, but also need to be reviewed by curators: “The act of kissing releases OxyContin in the brain—a hormone that strengthens the emotional bond between two people.” In this case, the likely intended reference was to oxytocin, but this post is not characterized as a Proto-AE because it did not include an adverse reaction.
While providing initial qualitative information on the identification of Proto-AEs in Twitter data, this study has limitations. First, the sample of posts involved was a convenience sample enriched for positive AEs. These data were assembled to define the Bayesian priors for an automated classification system, and cannot be considered to be representative of Twitter. We also did not fully de-duplicate the posts at this point; for the moment, we assume that, broadly aggregated, relative counts are stable even with some duplication.
Additional work will include the development of denominator-based pharmacoepidemiological methods to establish baseline and threshold levels of signal for each medical product such that when social media signals deviate from these ‘norms’, users can be passively notified of potential new safety signals. For now, this platform is viewed as a hypothesis-generating system where potential signal would be validated against more formal methods, but it could potentially become a more confirmatory resource as our methods and validation are refined.
Another key future direction for the work is in improved usage of the existing FAERS and other official FDA data sources. As mentioned above, currently FDA AE data are often delayed and difficult to use without extensive de-duplication and other pre-processing steps. OpenFDA (open.fda.gov) is a new initiative in the FDA Office of Informatics and Technology Innovation to offer open access data and highlight projects in both the public and private sector that use these data to further scientific research, educate the public, and improve health. It is expected to launch in fall 2014 and will provide API and raw download access to a number of high-value structured datasets, including open access AEs.
5 Conclusion
Proto-AEs identified in Twitter appear to have a similar profile by SOC to spontaneous reports received by the FDA. Some high-volume products had hashtags for reporting AEs (#accutaneprobz). Sample size for classified reports needs to be increased before causal associations can be made or signal identified for further investigation. Future directions for research include assessing severity of events, differentiating unlabeled events, time series modeling, incorporation of patient and health Web sites, and potentially search history data.
References
Moore TJ, Cohen MR, Furberg CD. Serious adverse drug events reported to the Food and Drug Administration, 1998–2005. Arch Intern Med. 2007;167:1752–9.
Haber P, Iskander J, Walton K, Campbell SR, Kohl KS. Internet-based reporting to the vaccine adverse event reporting system: a more timely and complete way for providers to support vaccine safety. Pediatrics. 2011;127(Suppl 1):S39–44.
Duggirala HJ, Herz ND, Caños DA, Sullivan RA, Schaaf R, Pinnow E, et al. Disproportionality analysis for signal detection of implantable cardioverter-defibrillator-related adverse events in the Food and Drug Administration Medical Device Reporting System. Pharmacoepidemiol Drug Saf. 2012;21:87–93.
Levinson D. Hospital incident reporting systems do not capture most patient harm (OEI-06-09-00091) 01-05-2012 [Internet]. Cited 17 Oct 2013. http://oig.hhs.gov/oei/reports/oei-06-09-00091.pdf.
Center for Drug Evaluation, Research. FDA adverse events reporting system (FAERS)—FDA adverse event reporting system (FAERS): latest quarterly data files. fda.gov. Center for drug evaluation and research.
European Medicines Agency. Guideline on good pharmacovigilance practices (GVP). EMA; London; 2012:1–84.
US Food and Drug Administration. Guidance for industry. Patient-reported outcome measures: use in medical product development to support labeling claims. US FDA; Silver Spring (MD); 2009:1–43.
Association of the British Pharmaceutical Industry. Pharmacovigilance Expert Network. Guidance notes on the management of adverse events and product complaints from digital media. 2nd ed. London; ABPI; 2013:1–14.
52 Cool Facts About Social Media—2012 Edition #socialmedia—Danny Brown [Internet]. dannybrown.me. Cited 25 Nov 2013. http://dannybrown.me/2012/06/06/52-cool-facts-social-media-2012/.
An exhaustive study of twitter users across the world—social media analytics beevolve [Internet]. temp.beevolve.com. Cited 25 Nov 2013. http://temp.beevolve.com/twitter-statistics/#c1.
Kass-Hout TA, Alhinnawi H. Social media in public health. Brit Med Bull. 2013;108:5–24.
Knezevic MZ, Bivolarevic IC, Peric TS, Jankovic SM. Using Facebook to increase spontaneous reporting of adverse drug reactions. Drug Saf. 2011;34:351–2.
Edwards IR, Lindquist M. Social media and networks in pharmacovigilance: boon or bane? Drug Saf. 2011;34:267–71.
White RW, Tatonetti NP, Shah NH, Altman RB, Horvitz E. Web-scale pharmacovigilance: listening to signals from the crowd. J Am Med Inform Assoc. 2013;20(3):404-8.
Freifeld CC, Mandl KD, Reis BY, Brownstein JS. HealthMap: global infectious disease monitoring through automated classification and visualization of internet media reports. J Am Med Inform Assoc. 2008;15:150–7.
CDC-VAERS-vaccine safety [Internet]. cdc.gov. Cited 25 Nov 2013. http://www.cdc.gov/vaccinesafety/Activities/vaers.html.
Pfizer. XELJANZ-tofacitinib citrate tablet, film coated [Internet]. labeling.pfizer.com. Cited 18 Oct 2013. http://labeling.pfizer.com/ShowLabeling.aspx?id=959§ion=MedGuide.
Duke JD, Friedlin J. ADESSA: a real-time decision support service for delivery of semantically coded adverse drug event data. AMIA Annu Symp Proc. 2010;2010:177–81.
dailymed.nlm.nih.gov [Internet]. Cited 18 Oct 2013. http://dailymed.nlm.nih.gov.
Viibryd US prescribing information [Internet]. Cited 12 March 2014. http://dailymed.nlm.nih.gov/dailymed/lookup.cfm?setid=4c55ccfb-c4cf-11df-851a-0800200c9a66.
Xarelto (Rivaroxaban) tablet, film coated [Janssen Pharmaceuticals, Inc.] [Internet]. dailymed.nlm.nih.gov. Cited 18 Oct 2013. http://dailymed.nlm.nih.gov/dailymed/lookup.cfm?setid=10db92f9-2300-4a80-836b-673e1ae91610#nlm34084-4.
Friedman RJ, Hess S, Berkowitz SD, Homering M. Complication rates after hip or knee arthroplasty in morbidly obese patients. Clin Orthop Relat Res. 2013;471:3358–66.
Acknowledgments
The authors gratefully acknowledge our collaborators at the FDA, including Jesse Goodman, who was at the Office of the Chief Scientist at the time of writing, and Mark Walderhaug and Taxiarchis Botsis at the Center for Biologics Evaluation and Research, for their guidance and expertise. The work was funded by the FDA Office of the Chief Scientist; the funders had input into the study design. We thank two anonymous peer reviewers for helpful comments on an earlier version of the manuscript.
Disclosures
Authors CCF, JSB, WB, and ND own shares in and are employed as consultants to Epidemico, Inc; CMM is employed as a consultant to Epidemico, Inc, a company working to commercialize aspects of the research discussed in this article. Clark C. Freifeld, John S. Brownstein, Christopher M. Menone, Wenjie Bao, Taha Kass-Hout, Ross Filice, and Nabarun Dasgupta have no other conflicts of interest that are directly relevant to the content of this study.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution Noncommercial License which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and the source are credited.
About this article
Cite this article
Freifeld, C.C., Brownstein, J.S., Menone, C.M. et al. Digital Drug Safety Surveillance: Monitoring Pharmaceutical Products in Twitter. Drug Saf 37, 343–350 (2014). https://doi.org/10.1007/s40264-014-0155-x
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40264-014-0155-x