Abstract
Purpose
Hepatocellular carcinoma (HCC) is one of the most common human malignancies. It has frequently been associated with metabolic perturbations and liver damages. Various members of the family of acyl-CoA synthetases are known to be involved in the production of bioactive fatty acids, and altered expression of its encoding genes has been found to be involved in metabolic perturbations. For the development of novel diagnostic and therapeutic HCC options, a fundamental understanding of the mechanisms associated with the deregulation of candidate genes involved in metabolic perturbation is required.
Methods
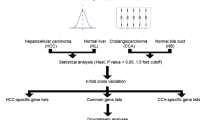
A meta-analysis of multiple HCC mRNA profiles was performed to identify consistently deregulated genes. Expression of the acyl-CoA synthetase medium chain family member 3 (ACSM3) gene was subsequently assessed in different HCC tumor stages and correlated with various clinicopathological features. Transcription regulation, survival and pathway-associated features of the ACSM3 gene were investigated using integrative functional genomic and molecular cell biological methods.
Results
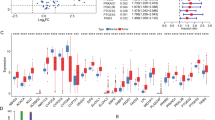
We found that expression of the ACSM3 gene was significantly reduced in HCC tissues and was frequently downregulated in patients exhibiting high alpha-fetoprotein (AFP) levels, high alanine aminotransferase (ALT) levels, multiple nodules and large tumors. Loss of ACSM3 expression was found to correlate with advanced HCC stages and a poor survival. In addition, HNF4α was found to positively regulate the expression of the ACSM3 gene, while PPARγ was found to transcriptionally repress it. Downregulation of ACSM3 expression was perceived upon activation of the TGFβ, WNT, AKT and MYC signalling pathways. In addition, we found that ACSM3 expression correlates with fatty acid oxidation in HCC.
Conclusion
Our data provide evidence for a differential expression and regulation of the ACSM3 gene in HCC, and may lay a foundation for therapeutically targeting fatty acid metabolism in these tumors.








Similar content being viewed by others
References
A.G. Singal, A. Pillai, J. Tiro, Early detection, curative treatment, and survival rates for hepatocellular carcinoma surveillance in patients with cirrhosis: A meta-analysis. PLoS Med 11, e1001624 (2014). doi:10.1371/journal.pmed.1001624
J. Liu, X. Wei, Y. Wu, Y. Wang, Y. Qiu, J. Shi, H. Zhou, Z. Lu, M. Shao, L. Yu, L. Tong, Giganteaside D induces ROS-mediated apoptosis in human hepatocellular carcinoma cells through the MAPK pathway. Cell Oncol 39, 333–342 (2016). doi:10.1007/s13402-016-0273-9
V. Ramesh, K. Selvarasu, J. Pandian, S. Myilsamy, C. Shanmugasundaram, K. Ganesan, NFkappaB activation demarcates a subset of hepatocellular carcinoma patients for targeted therapy. Cell Oncol 39, 523–536 (2016). doi:10.1007/s13402-016-0294-4
J.M. Llovet, J. Bustamante, A. Castells, R. Vilana, C. Ayuso Mdel, M. Sala, C. Bru, J. Rodes, J. Bruix, Natural history of untreated nonsurgical hepatocellular carcinoma: Rationale for the design and evaluation of therapeutic trials. Hepatology 29, 62–67 (1999). doi:10.1002/hep.510290145
K.T. Padhya, J.A. Marrero, A.G. Singal, J.K. Choi, J.Y. Choi, D.G. Kim, D.W. Choi, B.Y. Kim, K.H. Lee, Y.I. Yeom, H.S. Yoo, O.J. Yoo, S. Kim, Recent advances in the treatment of hepatocellular carcinoma. Curr Opin Gastroenterol 29, 285–292 (2013). doi:10.1097/MOG.0b013e32835ff1cf
J.K. Choi, J.Y. Choi, D.G. Kim, D.W. Choi, B.Y. Kim, K.H. Lee, Y.I. Yeom, H.S. Yoo, O.J. Yoo, S. Kim, Integrative analysis of multiple gene expression profiles applied to liver cancer study. FEBS Lett 565, 93–100 (2004). doi:10.1016/j.febslet.2004.03.081
S.K. Chan, O.L. Griffith, I.T. Tai, S.J. Jones, R. Elkon, C. Linhart, R. Sharan, R. Shamir, Y. Shiloh, Meta-analysis of colorectal cancer gene expression profiling studies identifies consistently reported candidate biomarkers. Cancer Epidemiol Biomark Prev 17, 543–552 (2008). doi:10.1158/1055-9965.EPI-07-2615
M. Giulietti, G. Occhipinti, G. Principato, F. Piva, Weighted gene co-expression network analysis reveals key genes involved in pancreatic ductal adenocarcinoma development. Cell Oncol 39, 379–388 (2016). doi:10.1007/s13402-016-0283-7
R. Elkon, C. Linhart, R. Sharan, R. Shamir, Y. Shiloh, Genome-wide in silico identification of transcriptional regulators controlling the cell cycle in human cells. Genome Res 13, 773–780 (2003). doi:10.1101/gr.947203
Y. Zhao, E.B. Butler, M. Tan, Targeting cellular metabolism to improve cancer therapeutics. Cell Death Dis 4, e532 (2013). doi:10.1038/cddis.2013.60
E. Currie, A. Schulze, R. Zechner, T.C. Walther, R.V. Farese Jr., Cellular fatty acid metabolism and cancer. Cell Metab 18, 153–161 (2013). doi:10.1016/j.cmet.2013.05.017
P.A. Watkins, D. Maiguel, Z. Jia, J. Pevsner, Evidence for 26 distinct acyl-coenzyme a synthetase genes in the human genome. J Lipid Res 48, 2736–2750 (2007). doi:10.1194/jlr.M700378-JLR200
H. Cai, H. Chen, T. Yi, C.M. Daimon, J.P. Boyle, C. Peers, S. Maudsley, B. Martin, VennPlex--a novel Venn diagram program for comparing and visualizing datasets with differentially regulated datapoints. PLoS One 8, e53388 (2013). doi:10.1371/journal.pone.0053388
C. Li, W.H. Wong, Model-based analysis of oligonucleotide arrays: Expression index computation and outlier detection. Proc Natl Acad Sci U S A 98, 31–36 (2001). doi:10.1073/pnas.011404098
J.T. Chang, J.R. Nevins, GATHER: A systems approach to interpreting genomic signatures. Bioinformatics 22, 2926–2933 (2006). doi:10.1093/bioinformatics/btl483
V.D. Marinescu, I.S. Kohane, A. Riva, The MAPPER database: A multi-genome catalog of putative transcription factor binding sites. Nucleic Acids Res 33, D91–D97 (2005). doi:10.1093/nar/gki103
A. Subramanian, P. Tamayo, V.K. Mootha, S. Mukherjee, B.L. Ebert, M.A. Gillette, A. Paulovich, S.L. Pomeroy, T.R. Golub, E.S. Lander, J.P. Mesirov, Gene set enrichment analysis: A knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci U S A 102, 15545–15550 (2005). doi:10.1073/pnas.0506580102
K. Kandasamy, S.S. Mohan, R. Raju, S. Keerthikumar, G.S. Kumar, A.K. Venugopal, D. Telikicherla, J.D. Navarro, S. Mathivanan, C. Pecquet, S.K. Gollapudi, S.G. Tattikota, S. Mohan, H. Padhukasahasram, Y. Subbannayya, R. Goel, H.K. Jacob, J. Zhong, R. Sekhar, V. Nanjappa, L. Balakrishnan, R. Subbaiah, Y.L. Ramachandra, B.A. Rahiman, T.S. Prasad, J.X. Lin, J.C. Houtman, S. Desiderio, J.C. Renauld, S.N. Constantinescu, O. Ohara, T. Hirano, M. Kubo, S. Singh, P. Khatri, S. Draghici, G.D. Bader, C. Sander, W.J. Leonard, A. Pandey, NetPath: a public resource of curated signal transduction pathways. Genome Biol 11, R3 (2010). doi:10.1186/gb-2010-11-1-r3
M. Muthuswami, V. Ramesh, S. Banerjee, S. Viveka Thangaraj, J. Periasamy, D. Bhaskar Rao, G.D. Barnabas, S. Raghavan, K. Ganesan, B.W. Dyer, F.A. Ferrer, D.K. Klinedinst and R. Rodriguez. Breast tumors with elevated expression of 1q candidate genes confer poor clinical outcome and sensitivity to Ras/PI3K inhibition. e77553 (2013). doi:10.1371/journal.pone.0077553
B.W. Dyer, F.A. Ferrer, D.K. Klinedinst, R. Rodriguez, A noncommercial dual luciferase enzyme assay system for reporter gene analysis. Anal Biochem 282, 158–161 (2000). doi:10.1006/abio.2000.4605
J.S. Lee, J. Taminau, C. Lazar, S. Meganck, A. Nowe, J. Sakamoto, H. Kimura, S. Moriyama, H. Odaka, Y. Momose, Y. Sugiyama, H. Sawada, Genomic profiling of liver cancer. Genomics Inform 11, 180–185 (2013). doi:10.5808/GI.2013.11.4.180
J. Taminau, C. Lazar, S. Meganck, A. Nowe, Comparison of merging and meta-analysis as alternative approaches for integrative gene expression analysis. ISRN Bioinform 2014, 345106 (2014). doi:10.1155/2014/345106
I. Boomgaarden, C. Vock, M. Klapper, F. Doring, Y. Hoshida, S.M. Nijman, M. Kobayashi, J.A. Chan, J.P. Brunet, D.Y. Chiang, A. Villanueva, P. Newell, K. Ikeda, M. Hashimoto, G. Watanabe, S. Gabriel, S.L. Friedman, H. Kumada, J.M. Llovet, T.R. Golub, J.W. Kim, Q. Ye, M. Forgues, Y. Chen, A. Budhu, J. Sime, L.J. Hofseth, R. Kaul, X.W. Wang, Comparative analyses of disease risk genes belonging to the acyl-CoA synthetase medium-chain (ACSM) family in human liver and cell lines. Biochem Genet 47, 739–748 (2009). doi:10.1007/s10528–009-9273-z
J. Sakamoto, H. Kimura, S. Moriyama, H. Odaka, Y. Momose, Y. Sugiyama, H. Sawada, Activation of human peroxisome proliferator-activated receptor (PPAR) subtypes by pioglitazone. Biochem Biophys Res Commun 278, 704–711 (2000). doi:10.1006/bbrc.2000.3868
H.S. Camp, O. Li, S.C. Wise, Y.H. Hong, C.L. Frankowski, X. Shen, R. Vanbogelen, T. Leff, Differential activation of peroxisome proliferator-activated receptor-gamma by troglitazone and rosiglitazone. Diabetes 49, 539–547 (2000)
J.M. Seargent, E.A. Yates, J.H. Gill, GW9662, a potent antagonist of PPARgamma, inhibits growth of breast tumour cells and promotes the anticancer effects of the PPARgamma agonist rosiglitazone, independently of PPARgamma activation. Br J Pharmacol 143, 933–937 (2004). doi:10.1038/sj.bjp.0705973
D.J. Adamson, D. Frew, R. Tatoud, C.R. Wolf, C.N. Palmer, Diclofenac antagonizes peroxisome proliferator-activated receptor-gamma signaling. Mol Pharmacol 61, 7–12 (2002)
C.P. Martinez-Jimenez, I. Kyrmizi, P. Cardot, F.J. Gonzalez, I. Talianidis, S. Yu, K. Matsusue, P. Kashireddy, W.Q. Cao, V. Yeldandi, A.V. Yeldandi, M.S. Rao, J.K. Reddy, Hepatocyte nuclear factor 4alpha coordinates a transcription factor network regulating hepatic fatty acid metabolism. Mol Cell Biol 30, 565–577 (2010). doi:10.1128/MCB.00927-09
S. Yu, K. Matsusue, P. Kashireddy, W.Q. Cao, V. Yeldandi, A.V. Yeldandi, M.S. Rao, F.J. Gonzalez, J.K. Reddy, Adipocyte-specific gene expression and adipogenic steatosis in the mouse liver due to peroxisome proliferator-activated receptor gamma1 (PPARgamma1) overexpression. J Biol Chem 278, 498–505 (2003). doi:10.1074/jbc.M210062200
M. Lehrke, M.A. Lazar, The many faces of PPARgamma. Cell 123, 993–999 (2005). doi:10.1016/j.cell.2005.11.026
Y. Hoshida, S.M. Nijman, M. Kobayashi, J.A. Chan, J.P. Brunet, D.Y. Chiang, A. Villanueva, P. Newell, K. Ikeda, M. Hashimoto, G. Watanabe, S. Gabriel, S.L. Friedman, H. Kumada, J.M. Llovet, T.R. Golub, N. Iwai, T. Mannami, H. Tomoike, K. Ono, Y. Iwanaga, Integrative transcriptome analysis reveals common molecular subclasses of human hepatocellular carcinoma. Cancer Res 69, 7385–7392 (2009). doi:10.1158/0008-5472.CAN-09-1089
N. Iwai, T. Mannami, H. Tomoike, K. Ono, Y. Iwanaga, An acyl-CoA synthetase gene family in chromosome 16p12 may contribute to multiple risk factors. Hypertension 41, 1041–1046 (2003). doi:10.1161/01.HYP.0000064944.60569.87
J.W. Kim, Q. Ye, M. Forgues, Y. Chen, A. Budhu, J. Sime, L.J. Hofseth, R. Kaul, X.W. Wang, Cancer-associated molecular signature in the tissue samples of patients with cirrhosis. Hepatology 39, 518–527 (2004). doi:10.1002/hep.20053
A. Budhu, M. Forgues, Q.H. Ye, H.L. Jia, P. He, K.A. Zanetti, U.S. Kammula, Y. Chen, L.X. Qin, Z.Y. Tang, X.W. Wang, Prediction of venous metastases, recurrence, and prognosis in hepatocellular carcinoma based on a unique immune response signature of the liver microenvironment. Cancer Cell 10, 99–111 (2006). doi:10.1016/j.ccr.2006.06.016
H.L. Jia, Q.H. Ye, L.X. Qin, A. Budhu, M. Forgues, Y. Chen, Y.K. Liu, H.C. Sun, L. Wang, H.Z. Lu, F. Shen, Z.Y. Tang, X.W. Wang, Gene expression profiling reveals potential biomarkers of human hepatocellular carcinoma. Clin Cancer Res 13, 1133–1139 (2007). doi:10.1158/1078-0432.CCR-06-1025
Q.H. Ye, L.X. Qin, M. Forgues, P. He, J.W. Kim, A.C. Peng, R. Simon, Y. Li, A.I. Robles, Y. Chen, Z.C. Ma, Z.Q. Wu, S.L. Ye, Y.K. Liu, Z.Y. Tang, X.W. Wang, Predicting hepatitis B virus-positive metastatic hepatocellular carcinomas using gene expression profiling and supervised machine learning. Nat Med 9, 416–423 (2003). doi:10.1038/nm843
N. Iizuka, M. Oka, H. Yamada-Okabe, M. Nishida, Y. Maeda, N. Mori, T. Takao, T. Tamesa, A. Tangoku, H. Tabuchi, K. Hamada, H. Nakayama, H. Ishitsuka, T. Miyamoto, A. Hirabayashi, S. Uchimura, Y. Hamamoto, Oligonucleotide microarray for prediction of early intrahepatic recurrence of hepatocellular carcinoma after curative resection. Lancet 361, 923–929 (2003). doi:10.1016/S0140-6736(03)12775-4
Y. Hoshida, A. Villanueva, M. Kobayashi, J. Peix, D.Y. Chiang, A. Camargo, S. Gupta, J. Moore, M.J. Wrobel, J. Lerner, M. Reich, J.A. Chan, J.N. Glickman, K. Ikeda, M. Hashimoto, G. Watanabe, M.G. Daidone, S. Roayaie, M. Schwartz, S. Thung, H.B. Salvesen, S. Gabriel, V. Mazzaferro, J. Bruix, S.L. Friedman, H. Kumada, J.M. Llovet, T.R. Golub, M.E. Monaco, C.J. Creighton, P. Lee, X. Zou, M.K. Topham, D.M. Stafforini, Gene expression in fixed tissues and outcome in hepatocellular carcinoma. N Engl J Med 359, 1995–2004 (2008). doi:10.1056/NEJMoa0804525
M.E. Monaco, C.J. Creighton, P. Lee, X. Zou, M.K. Topham, D.M. Stafforini, Expression of long-chain fatty acyl-CoA Synthetase 4 in breast and prostate cancers is associated with sex steroid hormone receptor negativity. Transl Oncol 3, 91–98 (2010)
X. Wu, Y. Li, J. Wang, X. Wen, M.T. Marcus, G. Daniels, D.Y. Zhang, F. Ye, L.H. Wang, X. Du, S. Adams, B. Singh, J. Zavadil, P. Lee, M.E. Monaco, Long chain fatty acyl-CoA synthetase 4 is a biomarker for and mediator of hormone resistance in human breast cancer. PLoS One 8, e77060 (2013). doi:10.1371/journal.pone.0077060
T. Mashima, S. Sato, S. Okabe, S. Miyata, M. Matsuura, Y. Sugimoto, T. Tsuruo, H. Seimiya, Acyl-CoA synthetase as a cancer survival factor: Its inhibition enhances the efficacy of etoposide. Cancer Sci 100, 1556–1562 (2009). doi:10.1111/j.1349-7006.2009.01203.x
Z. Pei, P. Fraisl, X. Shi, E. Gabrielson, S. Forss-Petter, J. Berger, P.A. Watkins, Very long-chain acyl-CoA synthetase 3: Overexpression and growth dependence in lung cancer. PLoS One 8, e69392 (2013). doi:10.1371/journal.pone.0069392
J.Y. Chiang, J.A. Bonzo, C.H. Ferry, T. Matsubara, J.H. Kim, F.J. Gonzalez, Hepatocyte nuclear factor 4alpha regulation of bile acid and drug metabolism. Expert Opin Drug Metab Toxicol 5, 137–147 (2009). doi:10.1517/17425250802707342
J.A. Bonzo, C.H. Ferry, T. Matsubara, J.H. Kim, F.J. Gonzalez, Suppression of hepatocyte proliferation by hepatocyte nuclear factor 4alpha in adult mice. J Biol Chem 287, 7345–7356 (2012). doi:10.1074/jbc.M111.334599
L. Michalik, B. Desvergne, W. Wahli, Peroxisome-proliferator-activated receptors and cancers: Complex stories. Nat Rev Cancer 4, 61–70 (2004). doi:10.1038/nrc1254
J. Rieusset, F. Touri, L. Michalik, P. Escher, B. Desvergne, E. Niesor, W. Wahli, A new selective peroxisome proliferator-activated receptor gamma antagonist with antiobesity and antidiabetic activity. Mol Endocrinol 16, 2628–2644 (2002). doi:10.1210/me.2002-0036
K.R. Kim, H.N. Choi, H.J. Lee, H.A. Baek, H.S. Park, K.Y. Jang, M.J. Chung, W.S. Moon, A peroxisome proliferator-activated receptor gamma antagonist induces vimentin cleavage and inhibits invasion in high-grade hepatocellular carcinoma. Oncol Rep 18, 825–832 (2007)
G. Martin, K. Schoonjans, A.M. Lefebvre, B. Staels, J. Auwerx, Coordinate regulation of the expression of the fatty acid transport protein and acyl-CoA synthetase genes by PPARalpha and PPARgamma activators. J Biol Chem 272, 28210–28217 (1997)
M. Yang, S.N. Li, K.M. Anjum, L.X. Gui, S.S. Zhu, J. Liu, J.K. Chen, Q.F. Liu, G.D. Ye, W.J. Wang, J.F. Wu, W.Y. Cai, G.B. Sun, Y.J. Liu, R.F. Liu, Z.M. Zhang, B.A. Li, A. Sanchez, A.M. Alvarez, J.M. Lopez Pedrosa, C. Roncero, M. Benito, I. Fabregat, A double-negative feedback loop between Wnt-beta-catenin signaling and HNF4alpha regulates epithelial-mesenchymal transition in hepatocellular carcinoma. J Cell Sci 126, 5692–5703 (2013). doi:10.1242/jcs.135053
A. Sanchez, A.M. Alvarez, J.M. Lopez Pedrosa, C. Roncero, M. Benito, I. Fabregat, Apoptotic response to TGF-beta in fetal hepatocytes depends upon their state of differentiation. Exp Cell Res 252, 281–291 (1999). doi:10.1006/excr.1999.4624
S. Lucas Sd, J.M. Lopez-Alcorocho, J. Bartolome, V. Carreno, Nitric oxide and TGF-beta1 inhibit HNF-4alpha function in HEPG2 cells. Biochem Biophys Res Commun 321, 688–694 (2004). doi:10.1016/j.bbrc.2004.07.025
D. Becker, I. Sfakianakis, M. Krupp, F. Staib, A. Gerhold-Ay, A. Victor, H. Binder, M. Blettner, T. Maass, S. Thorgeirsson, P.R. Galle, A. Teufel, Genetic signatures shared in embryonic liver development and liver cancer define prognostically relevant subgroups in HCC. Mol Cancer 11, 55 (2012). doi:10.1186/1476-4598-11-55
Acknowledgements
This study was supported by grants from the Department of Atomic Energy (DAE), Government of India (Grant No.6/6/2008/R&D-II-230R) and the Department of Biotechnology (DBT), Government of India (MKU-DBT–IPLS programme, No.BT/PR 14553/INF/22/124/2010). Instrumentation support from UGC-CEGS, DBT-IPLS, UGC-NRCBS, UGC-CAS, and the DST-PURSE programme-supported central facilities of SBS, MKU are greatly acknowledged.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
Conflict of Interest for all authors – None.
Additional information
Key Points
• Expression of ACSM3 negatively correlates with aggressiveness in HCC
• HNF4α and PPARγ are the transcriptional regulators of ACSM3
• Activated TGFβ & AKT signalling inhibits ACSM3 expression in HCC
Rights and permissions
About this article
Cite this article
Gopal, R., Selvarasu, K., Pandian, P.P. et al. Integrative transcriptome analysis of liver cancer profiles identifies upstream regulators and clinical significance of ACSM3 gene expression. Cell Oncol. 40, 219–233 (2017). https://doi.org/10.1007/s13402-017-0321-0
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13402-017-0321-0