Abstract
Here, we present liquid extraction surface analysis (LESA) coupled with electron-induced dissociation (EID) mass spectrometry in a Fourier-transform ion cyclotron resonance mass spectrometer for the analysis of small organic pharmaceutical compounds directly from dosed tissue. First, the direct infusion electrospray ionisation EID and collision-induced dissociation (CID) behaviour of erlotinib, moxifloxacin, clozapine and olanzapine standards were compared. EID mass spectra were also compared with experimental or reference electron impact ionisation mass spectra. The results show that (with the exception of erlotinib) EID and CID result in complementary fragment ions. Subsequently, we performed LESA EID MS/MS and LESA CID MS/MS on singly charged ions of moxifloxacin and erlotinib extracted from a thin tissue section of rat kidney from a cassette-dosed animal. Both techniques provided structural information, with the majority of peaks observed for the drug standards also observed for the tissue-extracted species. Overall, these results demonstrate the feasibility of LESA EID MS/MS of drug compounds from dosed tissue and extend the number of molecular structures for which EID behaviour has been determined.

ᅟ
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
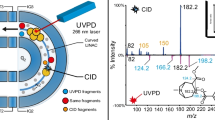
Tandem mass spectrometry, in which a precursor ion is characterised according to its fragments, is well-established as a tool for structural elucidation. Although collision-based methods remain the method of choice for the most part, there has been a growing reliance on electron-mediated methods over the past 20 years following the introduction of electron capture dissociation (ECD) [1]. ECD and the related method of electron transfer dissociation (ETD) [2] involve the capture/transfer of low-energy electrons by multiply charged cations. Electron-induced dissociation (EID) involves irradiation of singly charged ions with higher energy electrons causing further ionisation and/or excitation followed by dissociation of the precursor. EID is used here as a generic term to include methods which have been previously termed as electron-induced excitation of ions from organics (EIEIO) [3], electron excitation dissociation (EED) [3], and electron ionisation dissociation (also EID) [4]. EID has been applied to the structural characterisation of peptides [4,5,6,7,8], polyketides [9], metabolites [10], lipids [11], betaine dimers [12], amino acids [5], phosphorylated nucleotides [13], glycoconjugates (including glycopeptides, glycoalkaloids and glycolipids) [14], oxo-centered trinuclear carboxylate-bridged iron complexes [15] and octaethylporphoryin and its iron complex [16]. Mosely et al. [17] compared the EID mass spectra of 33 small singly charged organic molecules of pharmaceutical interest with their corresponding collision-induced dissociation (CID) mass spectra. They found that EID resulted in complementary fragments to CID, with very little overlap in observed product ions, suggesting that structural characterisation of such species would benefit from inclusion of both fragmentation techniques.
Recent work in our laboratory has focused on in situ analysis of biomolecules directly from biological substrates by use of liquid extraction surface analysis (LESA) mass spectrometry. LESA [18] has been demonstrated as a useful tool in the spatial profiling (imaging) of proteins [19, 20], lipids [21], and small molecule drugs [22, 23] in thin tissue sections. Here, we present LESA EID mass spectrometry of pharmaceutical compounds from kidney tissue from a cassette-dosed animal. Initial experiments focused on establishing the EID behaviour of erlotinib, olanzapine, moxifloxacin and clozapine small molecule drug standards (see Table 1), and comparing EID with CID behaviour. EID MS/MS spectra were also compared with electron impact ionisation mass spectra, either measured EI mass spectra or that reported in the NIST database (http://webbook.nist.gov/chemistry) [24,25,26]. Subsequently, we performed LESA mass spectrometry on a thin tissue section of rat kidney cassette-dosed with erlotinib, moxifloxacin, olanzapine and terfenadine. (The EID behaviour of terfenadine has previously been reported by Mosely et al. [17]). Peaks corresponding to singly charged erlotinib and moxifloxacin were observed in the mass spectrum. EID MS/MS and CID MS/MS were performed on singly charged ions of erlotinib and moxifloxacin. Despite the lower signal-to-noise of the LESA extracted precursor ions, the results show that structural information can be obtained following LESA EID, with the majority of fragments observed for the standard also observed for the tissue-extracted species. Nevertheless, the loss of the methyl ether group, which was unique to EID of the moxifloxacin standard, was not observed in the tissue-extracted species. Overall, these results show that LESA EID MS/MS of drug compounds from dosed tissue is feasible, expand the group of small organic molecules for which EID behaviour has been determined, and confirm that, in general, EID and CID provide complementary structural information.
Experimental
Materials
Stock solutions of erlotinib, moxifloxacin, and olanzapine at 10 mM were supplied by AstraZeneca (Cambridge, U.K). Stock solutions were diluted in methanol, water (50:50) to a final concentration of ~ 0.1 μM for EID/CID experiments. Ten microlitres of stock solution of erlotinib and moxifloxacin were diluted at around ~ 5 μM concentration for EI experiments. Methanol (LC-MS grade), acetonitrile (LC-MS grade) and water (LC-MS grade) were purchased from Fisher Scientific (Leicestershire, U.K.). One milligram of clozapine (VWR International Lutterworth, Leicester, U.K.) was dissolved in 1 mL of acetonitrile, water, 0.1% of formic acid (49.95:49.95:0.1) and further diluted to ~ 0.1 μM.
Dosing and Scheduling
Kidney tissue from orally dosed adult male Hans Wistar rats was used as described by Swales et al. [27]. The drugs were administered as a cassette containing erlotinib, moxifloxacin, olanzapine and terfenadine (at 10, 25, 10 and 25 mg/kg, respectively) and were supplied by AstraZeneca. The animal was euthanized 2 h post-dose. All tissue dissection was performed by trained AstraZeneca staff (project licence 40/3484, procedure number 10). The kidney was snap frozen and kept at − 80 °C until tissue processing. The kidney was cryosectioned into 10 μm sections using a cryostat CM1850 (Leica Microsystems, Wetzlar, Germany) and thaw mounted into glass slides (Fisher Scientific, Loughborough, UK) for LESA.
Electron-Induced Dissociation/Collision-Induced Dissociation
Each of the molecules presented in Table 1 were subjected to individual fragmentation and analysis using a 7 T solariX-XR (Bruker Daltonics, Bremen, Germany). Each molecule was ionised using nanospray ionisation (nESI) in positive ion mode using a capillary voltage of 800 V, with a temperature of 120 °C. Nanospray ionisation was achieved using pulled glass capillaries (P-97 Flaming/Brown micropipette puller tip Sutter Instrument Company, Novato, CA, USA) placed in front of the capillary spray shield. The singly charged species generated by electrospray were then isolated in the quadrupole (3 m/z isolation width) prior to CID and EID fragmentation. CID experiments were performed in the collision cell using argon as the collision gas and a voltage in the range 15 to 23 V (depending on the precursor ion). Three mass spectra were acquired at different voltages to aid in the identification of fragment ions. Each mass spectrum comprises 200 co-added scans. EID was performed in the ion cyclotron resonance cell (ParaCell) [28]. Electrons were supplied by a heated cathode at a current of 1.60 A, a cathode bias of 24.1 V, and lens potential of 0 V. EID pulse length was varied, according to each precursor ion, between 0.04 s and 0.06 s. Each EID mass spectrum comprises 800 co-added scans. MS experiments were performed in the broadband mode (m/z 80–500) with the following settings: 4-M data size, 1-s ion accumulation time, 0.9-ms time of flight and sweep width of 14.3 kHz. EID and CID spectra were externally calibrated with sodium trifluoroacetic acid clusters (mass accuracy of 5 ppm). EID background spectra were acquired by leaving all the parameters the same, but with the capillary spray voltage set to zero. EID background signal was subtracted from each EID spectra using in-house software. Only ions with a signal-to-noise ratio above 10 are reported (Supplementary Tables S-1 to S-8). Data analysis was performed with Data Analysis 4.2 software (Bruker Daltonics), in-house software, and validated by manual analysis.
Electron Ionisation
Erlotinib and moxifloxacin were ionised using a Waters solids probe (Waters Corp., Milford, MA, USA). Samples were left to dry at ambient temperature on a glass capillary inserted into the solids probe before it was placed inside the mass spectrometer Waters GTC Premier micromass (Waters Corp., Milford, MA, USA). The temperature was gradually increased in 10 K steps until fragment ions were observed from the molecular ion. Data acquisition and processing were carried out using MassLynx (ver. 4.1) software (Waters Corporation, Manchester, UK). The instrument was calibrated using heptacosafluorotributylamine to a mass range of m/z 49.996 to m/z 613.694. EI spectra of olanzapine, and clozapine, were obtained from the NIST Chemistry WebBook (http://webbook.nist.gov/chemistry) and manually assigned by the authors.
Liquid Extraction Surface Analysis
LESA of kidney tissue sections was performed by use of a Triversa NanoMate (Advion Biosciences, Ithaca, NY, USA), with a nanospray chip with 2.5-um ID nozzles. LESA Points software (Advion Biosciences, Ithaca, NY, USA) was used to select the precise location of the tissue to be sampled. The sample was loaded onto a universal adapter plate and placed in the Triversa Nanomate. The LESA extraction solvent comprised acetonitrile, water, isopropanol and 1% formic acid (47:47:5:1). Five microlitres of extraction solvent was aspirated from the solvent well. The robotic arm relocated to a position above the tissue and descended to a height 0.6 mm above the surface of the sample. Two microlitres of the solution was dispensed onto the tissue, forming a liquid microjunction. The liquid microjunction was maintained for 60 s, with one mixing step; then 2.5 μL were reaspirated into the pipette tip. The Triversa Nanomate was coupled with the solariX-XR (Bruker Daltonics, Bremen, Germany). Samples were introduced into the mass spectrometer via the Triversa Nanomate, with gas pressure 0.45 psi, a tip voltage of 2.0 kV and a capillary temperature of 150 °C. MS experiments were performed in broadband mode (m/z 80–500) with the following settings: 512 K data size, 1 s ion accumulation time, 0.9 ms time of flight and sweep width of 14.3 kHz. The mass spectra were calibrated by single-point correction with diethyhexylphalate (m/z 391.28429), which has an m/z close to that of erlotinib (m/z 394.17613) and moxifloxacin (m/z 402.18181). EID and CID parameters were as above with the exception that CID was performed at a voltage of 18 V only. Mass spectra comprise 20 co-added scans.
Results and Discussion
Electron ionisation, electron-, and collision-induced dissociation
Figure 1a, b shows the EID and CID mass spectra obtained from erlotinib. Similar cleavage was observed for the two techniques (Fig. 1c and Supplemental Table S-1) therefore either CID or EID can be used to obtain structural information for erlotinib. Two of the fragment ions (290.092 m/z and 292.108 m/z) show different relative abundances by EID and CID. [C17H14N3O2]+ (290.092 m/z, shown in blue in Fig. 1a, b) is less abundant following CID compared to EID, whereas [C17H14N3O2]+ (292.108 m/z, shown in red in Fig. 1a, b), is more abundant following CID than EID. Nevertheless, there is no apparent advantage, in terms of structural confirmation of erlotinib, of one fragmentation type over the other. Electron impact ionisation of erlotinib (Fig. 1d) results in a similar cleavage to both EID MS/MS and CID MS/MS (Fig. 1e and Supplemental Table S-2).
EID and CID mass spectra obtained from olanzapine are shown in Fig. 2a, b. Again, similar fragmentation patterns are observed for the two techniques (Fig. 2c and Supplemental Table S-3). The only additional fragment observed following EID is the ion [C13H12N3S]+. The EI mass spectrum for olanzapine (obtained from the NIST Chemistry WebBook, Supplementary Fig. S-4) is similar to the EID MS/MS spectrum, including the observation of the [C13H12N3S]+ fragment, although in the case of EI that fragment corresponds to the base peak in the mass spectrum.
Figure 3a shows the EID and CID mass spectra obtained from clozapine. The fragments observed are detailed in Supplementary Table S-5. Three fragment ions were observed following EID that were not observed following CID (highlighted in red in Fig. 3a). One fragment ion was observed in the CID spectrum of clozapine, corresponding to [C16H12ClN3]+ (highlighted in blue in Fig. 3b) which was not observed in the EID mass spectrum. Fragment ions shown in green (Fig. 3) correspond to the cleavage of the same bond in the methylpiperazine moiety and highlights the radical fragmentation of the EID experiments compared to CID (Fig. 3c). The overall bond cleavage obtained by EID and CID is similar to that observed in the EI mass spectrum of clozapine shown in the Supplemental Fig. S-6. (Abundant low m/z fragment ions observed following EI, but not EID or CID, are shown in red).
Clozapine: (a) EID spectrum (inset: expanded m/z region, vertical expansion × 1200); (b) CID spectrum (inset: expanded m/z region, vertical expansion × 10) and (c) schematic of bonds cleaved during EID and CID. Fragment ions highlighted in red are unique to EID, and in blue are unique to CID. Fragment ions shown in green highlight the radical fragmentation of the EID experiments
Figure 4a, b shows the EID and CID mass spectra obtained from moxifloxacin. The fragments observed are summarised in Supplemental Table S-7. Five fragment ions are common to both, mostly losses of functional groups around the core structure, for instance loss of water, carbon dioxide and hydrogen fluoride. There is a fragment ion at m/z 371.16220 (loss of ●CH3O from the precursor ion) which is observed in the EID mass spectrum but not the CID mass spectrum, highlighted red in Fig. 4a. Figure 4c shows an expanded m/z region from the CID mass spectra (obtained with different collision voltages), EID mass spectrum and EI mass spectrum. (The full EI mass spectrum is shown in Supplemental Fig. S-8 and assignment of the fragment ions is presented in Supplemental Table S-9). Loss of the radical methyl ether moiety is observed only in the EID and EI mass spectra of moxifloxacin.
Moxifloxacin: (a) EID spectrum; (b) CID spectrum; (c) enhanced m/z region (vertical expansion: EID = × 1200, CID = × 40, respectively), with the fragment ions at three different CID potentials; including EID and EI spectra with the loss of the methyl ether radical group ([M – ●CH3O]+); (d) schematic of bonds cleaved during EID and CID. Fragment ion highlighted in red is unique to EID
The results described above suggest that, in most cases, complementary fragments are observed following CID and EID of these small molecule drug compounds. These findings support earlier work by Mosely et al. [17] and suggest that structural elucidation of small singly charged molecules will benefit from inclusion of both fragmentation approaches. The EID fragmentation efficiency, calculated as the sum of the intensities of the fragment ions (excluding the precursor ion and the charge-reduced species) divided by the total ion abundance in the EID spectrum [29], is low for all the drug compounds when compared to CID fragmentation (see Supplementary Table S-10). This finding is in agreement with Prakash et al. [30].
LESA EID/CID
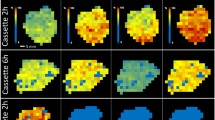
Following the experiments on small molecule drug standards, we subsequently applied EID and CID MS/MS to pharmaceutical compounds sampled directly from drug-dosed tissues by LESA. Figure 5a shows a photograph of a thin tissue section of kidney, from an animal dosed with erlotonib, moxifloxacin, olanzapine and terfenadine, mounted on a glass slide. The corresponding LESA mass spectrum of the sampled area is shown in Fig. 5b. The expanded m/z region (m/z 393–404) (Fig. 5c) reveals the presence of peaks at m/z 394.17543 and m/z 402.18632, corresponding to the singly charged ions of erlotinib (Δ 1.77 ppm) and moxifloxacin (Δ 2.36 ppm), respectively. Peaks corresponding to olanzapine and terfenadine or their metabolites were not observed. (In previous work, these species were observed following LESA [27]; however, that work was performed on a Q-trap instrument offering higher sensitivity than the FT-ICR). Figure 5d shows the mass spectrum obtained following quadrupole isolation of m/z 394 (erlotinib), and Fig. 5e shows the same for m/z 402 (moxifloxacin). These isolated ions were fragmented by EID (Fig. 5f, j) and the results are summarised in Table 2. Six fragment ions were identified in the EID spectrum of erlotinib and five fragment ions were present in the EID spectrum of moxifloxacin. In contrast to the EID of the moxifloxacin standard, loss of the radical methyl ether moiety was not observed in the EID of the moxifloxacin extracted directly from the tissue. This observation is likely due to the low extraction efficiency of the LESA experiment, which did not provide sufficient signal-to-noise ratio for the radical fragment ion to be detected. Surprisingly, sufficient erlotinib and moxifloxacin were extracted from the tissue in a single sampling event to acquire 20 scans of CID followed by EID for both drug molecules, i.e., we were able to compare EID and CID fragmentation from the same sampling location (see Supplemental Fig. S-11).
LESA coupled with EID (a) ~ 1 mm2 area sampled from rat kidney tissue mounted in glass slides (animal was dosed with erlotinib, moxifloxacin, terfenadine and clozapine); (b) MS spectrum of the sampled area; (c) enhanced area of the MS spectrum (2 × 10); (d) erlotinib: quadrupole isolation spectrum; (e) EID spectrum from erlotinib standard; (f) EID spectrum of erlotinib extracted from tissue; (g) schematic of bonds cleaved during EID of erlotinib extracted from tissue; (h) moxifloxacin quadrupole isolation spectrum; (i) EID spectrum from moxifloxacin standard; (j) EID spectrum of moxifloxacin extracted from the tissue; (k) schematic of bonds cleaved during EID of moxifloxacin extracted from tissue
Table 3 shows the fragment ions observed following CID of singly charged erlotonib and moxifloxacin. A total of 89% of the fragment ions observed in the CID mass spectrum of erlotinib standard (Supplemental Table S-1) were also identified in the LESA CID of erlotonib in tissue, whereas 71% of the fragment ions observed for moxifloxacin standard were present in the LESA CID mass spectrum of moxifloxacin.
Conclusions
The EID fragmentation behaviour of singly charged ions of erlotinib, moxifloxacin, clozapine and olanzapine has been determined. With the exception of erlotinib, EID MS/MS results in complementary fragments when compared with those observed following CID MS/MS. Our findings support the earlier work of Mosely et al. [17], i.e. that greater structural information about small organic molecules of pharmaceutical interest can be obtained through inclusion of both techniques in their analysis. Moreover, we demonstrate the feasibility of LESA EID MS/MS of drug compounds extracted directly from dosed tissue. Despite the lower S/N of the tissue-extracted drugs, a natural consequence of the use of tissue from dosed animals coupled with LESA extraction efficiency, the results show that the majority of fragments observed for the drug standards are also observed from tissue.
References
Zubarev, R.A., Kelleher, N.L., McLafferty, F.W.: Electron capture dissociation of multiply charged protein cations. A nonergodic process. J. Am. Chem. Soc. 120, 3265–3266 (1998)
Syka, J.E.P., Coon, J.J., Schroeder, M.J., Shabanowitz, J., Hunt, D.F.: Peptide and protein sequence analysis by electron transfer dissociation mass spectrometry. Proc. Natl. Acad. Sci. U. S. A. 101, 9528–9533 (2004)
Nielsen, M.L., Budnik, B.A., Haselmann, K.F., Olsen, J.V., Zubarev, R.A.: Intramolecular hydrogen atom transfer in hydrogen-deficient polypeptide radical cations. Chem. Phys. Lett. 330, 558–562 (2000)
Fung, Y.M.E., Adams, C.M., Zubarev, R.A.: Electron ionization dissociation of singly and multiply charged peptides. J. Am. Chem. Soc. 131, 9977–9985 (2009)
Lioe, H., O’Hair, R.A.J.: Comparison of collision-induced dissociation and electron-induced dissociation of singly protonated aromatic amino acids, cystine and related simple peptides using a hybrid linear ion trap-FT-ICR mass spectrometer. Anal. Bioanal. Chem. 389, 1429–1437 (2007)
Ly, T., Yin, S., Loo, J.A., Julian, R.R.: Electron-induced dissociation of protonated peptides yields backbone fragmentation consistent with a hydrogen-deficient radical. Rapid Commun. Mass Spectrom. 23, 2099–2101 (2009)
Wills, R.H., O’Connor, P.B.: Structural characterization of actinomycin D using multiple ion isolation and electron induced dissociation. J. Am. Soc. Mass Spectrom. 25, 186–195 (2014)
Voinov, V.G., Bennett, S.E., Barofsky, D.F.: Electron-induced dissociation of peptides in a triple quadrupole mass spectrometer retrofitted with an electromagnetostatic cell. J. Am. Soc. Mass Spectrom. 26, 752–761 (2015)
Wills, R.H., Tosin, M., Connor, P.B.O.: Structural characterization of polyketides using high mass accuracy tandem mass spectrometry. Anal. Chem. 84, 8863–8870 (2012)
Yoo, H.J., Liu, H., Håkansson, K.: Infrared multiphoton dissociation and electron-induced dissociation as alternative MS/MS strategies for metabolite identification metabolite analysis is the identification and structural. Anal. Chem. 79, 7858–7866 (2007)
Jones, J.W., Thompson, C.J., Carter, C.L., K.M.: Electron-induced dissociation (EID) for structure characterization of glycerophosphatidylcholine: determination of double bond positions and localization of acyl chains. J. Mass Spectrom. 50, 11327–11339 (2015)
Feketeová, L., Khairallah, G.N., O’Hair, R.A.: Intercluster chemistry of protonated and sodiated betaine dimers upon collision induced dissociation and electron induced dissociation. Eur. J. Mass Spectrom. 14, 107–110 (2008)
Ball, A.T., Prakash, A.S., Bristow, A.W., Sims, M., Mosely, J.: Characterisation of phosphorylated nucleotides by collisional and electron-based tandem mass spectrometry. Rapid Commun. Mass Spectrom. 30, 2155–2163 (2016)
Wong, Y.L.E., Chen, X., Wu, R., Hung, Y.L.W., Chan, T.W.D.: Structural characterization of intact glycoconjugates by tandem mass spectrometry using electron-induced dissociation. Anal. Chem. 89, 10111–10117 (2017)
Kaczorowska, M.A., Cooper, H.J.: Electron induced dissociation: a mass spectrometry technique for the structural analysis of trinuclear oxo-centred carboxylate-bridged iron complexes. J. Am. Soc. Mass Spectrom. 21, 1398–1403 (2010)
Kaczorowska, M.A., Cooper, H.J.: 2011: Electron induced dissociation (EID) tandem mass spectrometry of octaethylporphyrin and its iron (III) complex. Chem. Commun. 47, 418–420 (2011)
Mosely, J.A., Smith, M.J.P., Prakash, A.S., Sims, M., Bristow, A.W.T.: Electron-induced dissociation of singly charged organic cations molecules. Anal. Chem. 83, 4068–4075 (2011)
Kertesz, V., Van Berkel, G.J.: Fully automated liquid extraction-based surface sampling and ionization using a chip-based robotic nanoelectrospray platform. J. Mass Spectrom. 45, 252–260 (2010)
Griffiths, R.L., Creese, A.J., Race, A.M., Bunch, J., Cooper, H.J.: LESA FAIMS mass spectrometry for the spatial profiling of proteins from tissue. Anal. Chem. 88, 6758–6766 (2016)
Griffiths, R.L., Sisley, E.K., Lopez-Clavijo, A.F., Simmonds, A.L., Styles, I.B., Cooper, H.J.: Native mass spectrometry imaging of intact proteins and protein complexes in thin tissue sections. Int. J. Mass Spectrom. (2017). https://doi.org/10.1016/j.ijms.2017.10.009
Randall, E.C., Race, A.M., Cooper, H.J., Bunch, J.: MALDI imaging of liquid extraction surface analysis sampled tissue. Anal. Chem. 88, 8433–8440 (2016)
Swales, J.G., Tucker, J.W., Strittmatter, N., Nilsson, A., Cobice, D., Clench, M.R., Mackay, C.L., Andren, P.E., Takáts, Z., Webborn, P.J.H., Goodwin, R.J.A.: Mass spectrometry imaging of cassette-dosed drugs for higher throughput pharmacokinetic and biodistribution analysis. Anal. Chem. 86, 8473–8480 (2014)
Swales, J.G., Tucker, J.W., Spreadborough, M.J., Iverson, S.L., Clench, M.R., Webborn, P.J.H., Goodwin, R.J.A.: Mapping drug distribution in brain tissue using liquid extraction surface analysis mass spectrometry imaging. Anal. Chem. 87, 10146–10152 (2015)
Lotz, W.: Electron-impact ionization cross-sections for atoms up to Z=108. Z. Phys. 232, 101–107 (1970)
Wang, B., Mclafferty, F.W.: Electron impact excitation of ions from larger organic molecules. Org. Mass Spectrom. 25, 554–556 (1990)
F.W. Mclafferty, F. Tureček, McLafertty Interpretation Of Mass Spectral, 4th edn. (1993)
J.G. Swales, J.W. Tucker, N. Strittmatter, A. Nilsson, D. Cobice, M.R. Clench, C.L. Mackay, P.E. Andren, Mass Spectrometry Imaging of Cassette-Dosed Drugs for Higher Throughput Pharmacokinetic and Biodistribution Analysis. (2014)
Nikolaev, E.N., Jertz, R., Grigoryev, A., Baykut, G.: Fine structure in isotopic peak distributions measured using a dynamically harmonized Fourier transform ion cyclotron resonance cell at 7 T. Anal. Chem. 84, 2275–2283 (2012)
Iavarone, A.T., Paech, K., Williams, E.R.: Effects of charge state and cationizing agent on the electron capture dissociation of a peptide. Anal. Chem. 76(8), 2231–2238 (2004)
Prakash, A.S., Smith, M.J.P., Kaabia, Z., Hurst, G., Yan, C., Sims, M., Bristow, A.W.T., Stokes, P., Parker, D., Mosely, J.A.: Using electron induced dissociation (EID) on an LC time-scale to characterize a mixture of analogous small organic molecules. J. Am. Soc. Mass Spectrom. 23, 850–857 (2012)
Acknowledgements
This work was supported by EPSRC (EP/L023490/1). The Advion Triversa Nanomate used in this research was funded through Birmingham Science City Translational Medicine, Experimental Medicine Network of Excellence Project with support from Advantage West Midlands. The solariX-XR mass spectrometer used in this research was funded by the BBSRC (BB/M012492/1). Supplementary data supporting this research is openly available from the University of Birmingham data archive at DOI https://doi.org/10.25500/eData.bham.00000206.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
ESM 1
(DOCX 852 kb)
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Lopez-Clavijo, A.F., Griffiths, R.L., Goodwin, R.J.A. et al. Liquid Extraction Surface Analysis (LESA) Electron-Induced Dissociation and Collision-Induced Dissociation Mass Spectrometry of Small Molecule Drug Compounds. J. Am. Soc. Mass Spectrom. 29, 2218–2226 (2018). https://doi.org/10.1007/s13361-018-2042-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13361-018-2042-7