Abstract
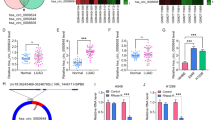
Both intra-tumor macrophage and T-box transcription factor Brachyury (T) have been proved to play important roles in tumor progression and metastasis. However, it is still unknown whether T could regulate the infiltration of macrophages. Here, we report that the Brachyury expression in human lung tumors was inversely correlated with the infiltration of macrophages. Brachyury suppressed the capability of human lung cancer cells to attract macrophages. Using PCR array, we found that Brachyury inhibited expression of several chemokines, including CCL2, CCL4, and CXCL10. Accordingly, knockdown of CCL2 and CCL4 in lung cancer cells suppressed macrophage invasion under coculture conditions. Furthermore, we found that Brachyury expression was inversely correlated with CCL2 and CCL4 expression in human lung tumors. Taken together, our findings shed light on the novel role of Brachyury in regulation of macrophage infiltration.





Similar content being viewed by others
References
Siegel R, Ma J, Zou Z, Jemal A. Cancer statistics, 2014. CA Cancer J Clin. 2014;64:9–29.
Cirkel GA, Gadellaa-van Hooijdonk CG, Koudijs MJ, Willems SM, Voest EE. Tumor heterogeneity and personalized cancer medicine: are we being outnumbered? Future Oncol. 2014;10:417–28.
Longo DL. Tumor heterogeneity and personalized medicine. N Engl J Med. 2012;366:956–7.
Wood SL, Pernemalm M, Crosbie PA, Whetton AD. The role of the tumor-microenvironment in lung cancer-metastasis and its relationship to potential therapeutic targets. Cancer Treat Rev. 2014;40:558–66.
Eming SA, Krieg T, Davidson JM. Inflammation in wound repair: molecular and cellular mechanisms. J Invest Dermatol. 2007;127:514–25.
Ovchinnikov DA. Macrophages in the embryo and beyond: much more than just giant phagocytes. Genesis. 2008;46:447–62.
Mills CD. M1 and m2 macrophages: oracles of health and disease. Crit Rev Immunol. 2012;32:463–88.
Deodhar AK, Rana RE. Surgical physiology of wound healing: a review. J Postgrad Med. 1997;43:52–6.
Greenhalgh DG. The role of apoptosis in wound healing. Int J Biochem Cell Biol. 1998;30:1019–30.
Barrientos S, Stojadinovic O, Golinko MS, Brem H, Tomic-Canic M. Growth factors and cytokines in wound healing. Wound Repair Regen. 2008;16:585–601.
Papaioannou VE. T-box genes in development: from hydra to humans. Int Rev Cytol. 2001;207:1–70.
Tada M, Smith JC. T-targets: clues to understanding the functions of t-box proteins. Dev Growth Differ. 2001;43:1–11.
Dobrovolskaia-Zavadskaia N. Regarding the spontaneous mortification of the tail of a new-born mouse and the existence of a hereditary characteristic (factor). C R Seances Soc Biol Fil. 1927;97:114–6.
Wilkinson DG, Bhatt S, Herrmann BG. Expression pattern of the mouse t gene and its role in mesoderm formation. Nature. 1990;343:657–9.
Klaus A, Birchmeier W. Wnt signalling and its impact on development and cancer. Nat Rev Cancer. 2008;8:387–98.
16Li Y, Luo H, Liu T, Zacksenhaus E, Ben-David Y. The ets transcription factor fli-1 in development, cancer and disease. Oncogene. 2014.
Hsu I, Vitkus S, Da J, Yeh S. Role of oestrogen receptors in bladder cancer development. Nat Rev Urol. 2013;10:317–26.
Takebe N, Nguyen D, Yang SX. Targeting notch signaling pathway in cancer: clinical development advances and challenges. Pharmacol Ther. 2014;141:140–9.
Turner N, Grose R. Fibroblast growth factor signalling: from development to cancer. Nat Rev Cancer. 2010;10:116–29.
Cao L, Yu Y, Bilke S, Walker RL, Mayeenuddin LH, Azorsa DO, et al. Genome-wide identification of pax3-fkhr binding sites in rhabdomyosarcoma reveals candidate target genes important for development and cancer. Cancer Res. 2010;70:6497–508.
Thoma C. Prostate cancer: Brachyury—a biomarker for progression and prognosis? Nat Rev Urol. 2014.
Pires MM, Aaronson SA. Brachyury: a new player in promoting breast cancer aggressiveness. J Natl Cancer Inst. 2014;106.
Pinto F, Pertega-Gomes N, Pereira MS, Vizcaino JR, Monteiro P, Henrique RM, et al. T-box transcription factor brachyury is associated with prostate cancer progression and aggressiveness. Clin Cancer Res. 2014.
Palena C, Roselli M, Litzinger MT, Ferroni P, Costarelli L, Spila A, et al. Overexpression of the emt driver brachyury in breast carcinomas: association with poor prognosis. J Natl Cancer Inst. 2014;106.
Kobayashi Y, Sugiura T, Imajyo I, Shimoda M, Ishii K, Akimoto N, et al. Knockdown of the t-box transcription factor brachyury increases sensitivity of adenoid cystic carcinoma cells to chemotherapy and radiation in vitro: Implications for a new therapeutic principle. Int J Oncol. 2014;44:1107–17.
Roselli M, Fernando RI, Guadagni F, Spila A, Alessandroni J, Palmirotta R, et al. Brachyury, a driver of the epithelial-mesenchymal transition, is overexpressed in human lung tumors: an opportunity for novel interventions against lung cancer. Clin Cancer Res. 2012;18:3868–79.
Park JC, Chae YK, Son CH, Kim MS, Lee J, Ostrow K, et al. Epigenetic silencing of human t (brachyury homologue) gene in non-small-cell lung cancer. Biochem Biophys Res Commun. 2008;365:221–6.
Singer JB, Harbecke R, Kusch T, Reuter R, Lengyel JA. Drosophila brachyenteron regulates gene activity and morphogenesis in the gut. Development. 1996;122:3707–18.
Zheng J, Yang M, Shao J, Miao Y, Han J, Du J. Chemokine receptor cx3cr1 contributes to macrophage survival in tumor metastasis. Mol Cancer. 2013;12:141.
Iida N, Nakamoto Y, Baba T, Nakagawa H, Mizukoshi E, Naito M, et al. Antitumor effect after radiofrequency ablation of murine hepatoma is augmented by an active variant of cc chemokine ligand 3/macrophage inflammatory protein-1alpha. Cancer Res. 2010;70:6556–65.
Mizutani K, Sud S, McGregor NA, Martinovski G, Rice BT, Craig MJ, et al. The chemokine ccl2 increases prostate tumor growth and bone metastasis through macrophage and osteoclast recruitment. Neoplasia. 2009;11:1235–42.
Herrmann BG, Labeit S, Poustka A, King TR, Lehrach H. Cloning of the t gene required in mesoderm formation in the mouse. Nature. 1990;343:617–22.
Dobrovolskaia-Zavadskaia N. Brachyura, accompanying the bendings and the genetic structure of the tail of the mouse. C R Seances Soc Biol Fil. 1927;97:1583–5.
Yamada A, Koyanagi KO, Watanabe H. In silico and in vivo identification of the intermediate filament vimentin that is downregulated downstream of brachyury during xenopus embryogenesis. Gene. 2012;491:232–6.
Katikala L, Aihara H, Passamaneck YJ, Gazdoiu S, Jose-Edwards DS, Kugler JE, et al. Functional brachyury binding sites establish a temporal read-out of gene expression in the ciona notochord. PLoS Biol. 2013;11:e1001697.
Kilic N, Feldhaus S, Kilic E, Tennstedt P, Wicklein D, Wasielewski R, et al. Brachyury expression predicts poor prognosis at early stages of colorectal cancer. Eur J Cancer. 2011;47:1080–5.
Sarkar D, Shields B, Davies ML, Muller J, Wakeman JA. Brachyury confers cancer stem cell characteristics on colorectal cancer cells. Int J Cancer. 2012;130:328–37.
Pillay N, Plagnol V, Tarpey PS, Lobo SB, Presneau N, Szuhai K, et al. A common single-nucleotide variant in t is strongly associated with chordoma. Nat Genet. 2012;44:1185–7.
Kitamura Y, Sasaki H, Kimura T, Miwa T, Takahashi S, Kawase T, et al. Molecular and clinical risk factors for recurrence of skull base chordomas: gain on chromosome 2p, expression of brachyury, and lack of irradiation negatively correlate with patient prognosis. J Neuropathol Exp Neurol. 2013;72:816–23.
Nibu Y, Jose-Edwards DS, Di Gregorio A. From notochord formation to hereditary chordoma: the many roles of brachyury. Biomed Res Int. 2013;2013:826435.
Barresi V, Ieni A, Branca G, Tuccari G. Brachyury: a diagnostic marker for the differential diagnosis of chordoma and hemangioblastoma versus neoplastic histological mimickers. Dis Markers. 2014;2014:514753.
Scheil-Bertram S, Kappler R, von Baer A, Hartwig E, Sarkar M, Serra M, et al. Molecular profiling of chordoma. Int J Oncol. 2014;44:1041–55.
Haro A, Yano T, Kohno M, Yoshida T, Koga T, Okamoto T, et al. Expression of brachyury gene is a significant prognostic factor for primary lung carcinoma. Ann Surg Oncol. 2013;20 Suppl 3:S509–516.
Quatromoni JG, Eruslanov E. Tumor-associated macrophages: function, phenotype, and link to prognosis in human lung cancer. Am J Transl Res. 2012;4:376–89.
Becker M, Muller CB, De Bastiani MA, Klamt F. The prognostic impact of tumor-associated macrophages and intra-tumoral apoptosis in non-small cell lung cancer. Histol Histopathol. 2014;29:21–31.
El-Nikhely N, Larzabal L, Seeger W, Calvo A, Savai R. Tumor-stromal interactions in lung cancer: novel candidate targets for therapeutic intervention. Expert Opin Investig Drugs. 2012;21:1107–22.
Acknowledgments
This study Project is supported by the National Natural Science Foundation of China (Grant No. 81102036)
Conflicts of interest
None
Author information
Authors and Affiliations
Corresponding author
Additional information
Su Chen and Jian Jiao contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplementary Table 1
Patient information of the specimens enrolled in the study. Case numbers were determined upon hospitalization. M stands for male; F stands for female and the ages were as shown. The diagnosis and the location of the tumors are recorded. TNM stage was determined equivalent to American Joint Committee of Cancer-AJCC staging standard. Scoring of the IHC staining was recorded as the average score from three independent pathologists’ analysis. (PDF 15 kb)
Supplementary Figure 1
Differential expression level of Brachyury in lung cancer cell lines. A. Semi-quantitative RT-PCR was done to test the expression level in the lung cancer cell lines as indicated. Samples were retrieved after 30 cycles of PCR amplification. 18S ribosome RNA was used as loading control to ensure equal loading of each sample. B. Western Blot result of Brachyury expression in the cell lines shown. GAPDH was blotted as loading control. (PDF 30 kb)
Supplementary Figure 2
Generation of H460 cells with stable knockdown of BrachyuryandH1299 cell line with overexpression of Brachyury. A. Semi-quantitative RT-PCR of Brachyury expression in H460 cells transfected with control shRNAs (H460shN) or Brachyury shRNAs (H460shT). 18S was used as loading control. B. Western blot result testing Brachyury expression in H460shN and H460shT cells. GAPDH was used as loading control. C. RT-PCR examination of Brachyury expression in H1299 cells transfected with empty vector (H1299EV) or Brachyury overexpression plasmid (H1299TOE). 18S was used as loading control. D. Western blot result testing Brachyury expression in H1299EV and H1299TOE cells. GAPDH was used as loading control. (PDF 34 kb)
Supplementary Figure 3
The gene list of 84 genes that encode chemokines and their receptors (PDF 13 kb)
Rights and permissions
About this article
Cite this article
Chen, S., Jiao, J., Jiang, D. et al. T-box transcription factor Brachyury in lung cancer cells inhibits macrophage infiltration by suppressing CCL2 and CCL4 chemokines. Tumor Biol. 36, 5881–5890 (2015). https://doi.org/10.1007/s13277-015-3260-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13277-015-3260-2