Abstract
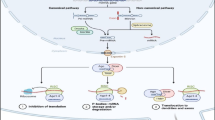
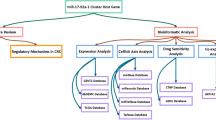
Diagnostic markers are needed for achieving a cure in esophageal cancer, detecting tumor cells earlier. Exosomes are bioactive vesicles secreted by cells into surrounding body fluids. Exosome formation, cargo content, and delivery have major impact in cancer development. This is the first isolation of exosomes from serum of patients with adenocarcinoma of the esophagus and comparison of exosomal miRNA profiles with matching primary tumor and normal tissues. RNA was extracted for miRNA profiling by real-time TaqMan miR arrays. The miR profiles of exosomal cargo, matching tumor, and normal tissue of a subgroup of adenocarcinoma patients have been compared. “Exosomal onco-miRs” such as miR-223-5p, miR-223-3p, miR-483-5p, miR-409-3p, miR-196b-5p, miR-192-5p, miR-146a-5p, and miR-126-5p have been identified as part of exosomal cargo being overexpressed in corresponding tumor compared to normal. Upregulation of miR-223-5p and miR-483-5p in adenocarcinoma (p = 0.034, p = 0.017) has been verified by an independent cohort of 43 patients with T2-3 adeno- and squamous cell carcinoma. In contrast, miR-224-5p, miR-452-5p, miR-23b-5p, miR-203-5p, miR-1201-5p, miR-149-5p, miR-671-3p, miR-944-5p, miR-27b-3p, and miR-22-3p have been identified to be significantly downregulated in adenocarcinoma versus normal and merely or not detectable in exosomes. “Exosomal onco-miRs” are a novel, stable, and noninvasive source for diagnosis and therapy monitoring of esophageal cancer. Oncogenic shuttle miRNAs present in exosomes may contribute to understanding how tumor cells spread their oncogenic potential to the environment. The “exosomal onco-miRs” identified seem to play a major role and may be applied for noninvasive diagnosis and therapy monitoring of adenocarcinoma of the esophagus.






Similar content being viewed by others
Abbreviations
- CD9, 63, 81:
-
Cluster of differentiation 9, 63, 81
- Ct:
-
Cycle threshold
- CT:
-
Computer tomography
- EAC:
-
Esophageal adenocarcinoma
- E2F1:
-
Transcription factor
- ELISA:
-
Enzyme-linked immunosorbent assay
- ESCC:
-
Esophageal squamous cell carcinoma
- FBXW7:
-
F-box and WD repeat domain-containing 7
- FFPE:
-
Formalin-fixed paraffin-embedded
- MIR:
-
MicroRNA
- MMP-9:
-
Metalloproteinase 9
- NTA:
-
Nanoparticle tracking analysis
- PTPN1:
-
Tyrosine-protein phosphatase non-receptor type 1
- RT:
-
Reverse transcription
- RT-PCR:
-
Real-time polymerase chain reaction
- SCLC:
-
Squamous cell lung carcinoma
- TSG101:
-
Tumor susceptibility gene 101
- T:
-
Tumor stage
- USF2:
-
Upstream stimulatory factor 2
- 3′UTR:
-
3′ untranslated region
- VEGFC:
-
Vascular endothelial growth factor C
References
Bollschweiler E, Wolfgarten E, Gutschow C, Hölscher AH. Demographic variations in the rising incidence of esophageal adenocarcinoma in white males. Cancer. 2001;92:549–55.
Thrift AP, Whiteman DC. The incidence of esophageal adenocarcinoma continues to rise: analysis of period and birth cohort effects on recent trends. Ann Oncol. 2012;12:3155–62.
van Hagen P, Hulshof MC, van Lanschot JJ, Steyerberg EW, van Berge Henegouwen MI, Wijnhoven BP, et al. Preoperative chemoradiotherapy for esophageal or junctional cancer. N Engl J Med. 2012;366:2074–84.
Bollschweiler E, Hölscher AH, Metzger R. Histologic tumor type and the rate of complete response after neoadjuvant therapy for esophageal cancer. Future Oncol. 2010;6:25–35.
Schneider PM, Baldus SE, Metzger R, Kocher M, Bongartz, Bollschweiler E, et al. Histomorphologic tumor regression and lymph node metastases determine prognosis following neoadjuvant radiochemotherapy for esophageal cancer: implications for response classification. Ann Surg. 2005;242:684–92.
Hölscher AH, Bollschweiler E, Schröder W, Metzger R, Gutschow C, Drebber U. Prognostic impact of upper, middle, and lower third mucosal or submucosal infiltration in early esophageal cancer. Ann Surg. 2011;254:802–7.
Denzer K, Kleijmeer MJ, Heijnen HF, Stoorvogel W, Geuze HJ, eds: Exosome: from internal vesicle of the multivesicular body to intercellular signalling device. In: J Cell Sci. 2000;113:3365–3374.
Wang G-J, Liu Y, Qin A, Shah SV, Deng ZB, Xiang X, et al. Thymus exosomes-like particles induce regulatory T cells. J Immunol. 2008;181:5242–8.
Lee TH, D’Asti E, Magnus N, Al-Nedawi K, Meehan B, Rak J. Microvesicles as mediators of intercellular communication in cancer—the emerging science of cellular ‘debris’. Semin Immunopathol. 2011;33:455–67.
Kaiser J. MiRNAs rewrite the rules of molecular biology. Res J Biol Technol. 2007;2:3–5.
Gowhar A, Kaiser J, Atya K, Hemant P. RNAi as a novel therapeutic platform technology for oncological solutions—a review. Biotechnol Mol Biol Rev. 2009;4:055–70.
Lu J, Getz G, Miska EA, Alvarez-Saavedra E, Lamb J, Peck D, et al. MicroRNA expression profiles classify human cancers. Nature. 2005;435:834–8.
Marilena V, Croce I, Croce CM. MicroRNA involvement in human cancer. Carcinogenesis. 2012;33:1126–33.
Valadi H, Ekström K, Bossios A, Sjöstrand M, Lee JJ, Lötvall JO. Exosome-mediated transfer of mRNAs and microRNAs is a novel mechanism of genetic exchange between cells. Nat Cell Biol. 2007;9:654–9.
Fidler IJ, Poste G. The “seed and soil” hypothesis revisited. Lancet Oncol. 2008;9:808. PMID:18672217.
Filipazzi P, Bürdek M, Villa A, Rivoltini L, Huber V. Recent advances on the role of tumor exosomes in immunosuppression and disease progression. Semin Cancer Biol. 2012;22:342–9.
Zwicker JI, Liebmann HA, Neuberg D, Lacroix R, Bauer KA, Furie BC, et al. Tumor-derived tissue factor-bearing microparticles are associated with venous thromboembolic events in malignancy. Clin Cancer Res. 2009;15:6830–40.
Lakkaraju A, Rodriguez-Boulan E. Itinerant exosomes: emerging roles in cell and tissue polarity. Trends Cell Biol. 2008;18:199–209.
Jörgensen M, Beek R, Pedersen S, Söndergaard EK, Kreisensen SR, Varming K: Extracellular vesicle (EV) array: microarray capturing of exosomes and other extracellular vesicles for multiplexed phenotyping. J Extracell vesicles 2013;2:doi: 10.3402/jev.v2i0.20920.eCollection.
Takeshita N, Hoshino I, Mori M, Akutsu Y, Hanari N, Yoneyama Y, et al. Serum microRNA expression profile: miR-1246 as a novel diagnostic and prognostic biomarker for oesophageal squamous cell carcinoma. Br J Cancer. 2013;108:644–52.
Tanaka Y, Kamohara H, Kinoshita K, Kurashige J, Ishimoto T, Iwasuki M, et al. Clinical impact of serum exosomal microRNA-21 as a clinical biomarker in human esophageal squamous cell carcinoma. Cancer. 2013;119:1159–67.
Wu C, Wang C, Guan X, Liu Y, Li D, Zhou X, Zhang Y, Chen X, Wang J, Zen K, Zhang CY, Zhang C: Diagnostic and prognostic implications of a serum miRNA panel in oesophageal squamous cell carcinoma. PLoS One 2014, e92292.
Li B, Zhao Y, Guo G, Li W, Zhu ED, Luo X, et al. Plasma microRNAs, miR-223, miR-21 and miR-218, as novel potential biomarkers for gastric cancer detection. PLoS ONE. 2012;7:e41629.
Li J, Guo Y, Liang X, Sun M, Wang G, De W, et al. MicroRNA-223 functions as an oncogene in human gastric cancer cell line by targeting FBXW7/hCdc4. J Cancer Res Clin Oncol. 2012;138:763–74.
Yang M, Chen J, Su F, Yu B, Su F, Lin L, Liu Y, Huang JD, Song E: Microvesicles secreted by macrophages shuttle invasion-potentiating microRNAs into breast cancer cells. Mol Cancer 2011;117.
Li S, Li Z, Guo F, Qin X, Liu B. miR-223 regulates migration and invasion by targeting Artemin in human esophageal carcinoma. J Biomed Sci. 2011;18:24–32.
Gaedcke J, Grade M, Camps J, Sökilde R, Kaczkowski B, Schetter AJ, et al. The rectal cancer microRNAome–microRNA expression in rectal cancer and matched normal mucosa. Clin Cancer Res. 2012;18:4919–30.
Patterson EE, Holloway AK, Weng J, Fojo T, Kebebew E. MicroRNA profiling of adrenocortical tumors reveals miR-483 as a marker of malignancy. Cancer. 2011;117:1630–9.
Song Q, Xu Y, Yag C, Chen Z, Jia C, Chen J. miR-483-5p promotes invasion and metastasis of lung adenocarcinoma by targeting RhoGDI1 and ALCAM. Cancer Res. 2014;74:3031–42.
Chiba M, Kimura M, Asari S. Exosomes secreted from human colorectal cancer cell lines contain mRNAs, microRNAs and natural antisense RNAs, that can transfer into the human hepatoma HepG2 and lung cancer A549 cell lines. Oncol Rep. 2012;28:1551–8.
Saad R, Chen Z, Zhu S, Jia P, Zhao Z, Washington MK, et al. Deciphering the unique microRNA signature in human esophageal adenocarcinoma. Cancer Lett PLoS ONE. 2013;8:e64463.
Liu SG, Qin XG, Zhao BS, Qi B, Yao WJ, Wang TY, et al. Differential expression of miRNAs in esophageal cancer tissue. Oncol Lett. 2013;5:1639–42.
Weng C, Dong H, Chen G, Zhai Y, Baj R, Hu H, et al. miR-409-3p inhibits HT1080 cell proliferation, vascularization and metastasis b targeting angiogenin. Cancer Lett. 2012;323:171–9.
Li C, Nie H, Wang M, Su L, Li J, Yu B. MicroRNA-409-3p regulates cell proliferation and apoptosis by targeting PHF10 in gastric cancer. Cancer Lett. 2012;320:189–99.
Cristóbal I, Aguilera O, García-Foncillas J, Zazo S, Madoz-Gúrpide J, Rojo F. Clinical significance of miR-126 in colorectal cancer. Genes Chromosome Cancer. 2014;53:881.
Yin J, Bai Z, Song J, Yang Y, Wang J, Han W, et al. Differential expression of serum miR-126, miR-141 and miR-21 as novel biomarkers for early detection of liver metastasis in colorectal cancer. Chin J Cancer Res. 2014;26:95–103.
Jia AY, Castillo-Martin M, Bonal DM, Sánchez-Carbayo M, Silva JM, Cordon-Cardo C. MicroRNA-126 inhibits invasion in bladder cancer via regulation of ADAM9. Br J Cancer. 2014;110:2945–54.
Guo B, Li J, Liu L, Hou N, Chang D, Zhao L, et al. Dysregulation of miRNAs and their potential as biomarkers for the diagnosis of gastric cancer. Biomed Rep. 2013;1:907–12.
Ding H, Wu YL, Wang YX, Zhu FF. Characterization of the microRNA expression profile of cervical squamous cell carcinoma metastases. Asian Pac J Cancer Prev. 2014;15:1675–9.
Wang HB, Jiang ZB, Li M: Research on the typical miRNA and target genes in squamous cell carcinoma and adenocarcinoma of esophagus cancer with DNA microarray. Pathol Oncol Res. 2014 (Epub ahead of print).
Gu J, Wang Y, Wu X. MicroRNA in the pathogenesis and prognosis of esophageal cancer. Curr Pharm Des. 2013;19:1292–300.
Ye J, Wu X, Wu D, Wu P, Ni C, Zhang Z, et al. P miRNA-27b targets vascular endothelial growth factor C to inhibit tumor progression and angiogenesis in colorectal cancer. PLoS ONE. 2013;8(4):e60687. doi:10.1371/journal.pone.0060687.
Lin ZY, Huang YQ, Zhang YQ, Han ZD, He HC, Ling XH, et al. MicroRNA-224 inhibits progression of human prostate cancer by TRIB1. Int J Cancer. 2014;135:541–50.
Wang Y, Ren J, Gao Y, Ma JZ, Toh HC, Chow P, et al. MicroRNA-224 targets SMAD familiy member 4 to promote cell proliferation and negatively influence patient survival. PLoS ONE. 2013;8:e68744. doi:10.1371/journal.pone.0068744.
Shen SN, Wang LF, Jia YF, Hao YQ, Zhang L, Wang H. Upregulation of microRNA-224 is associated with aggressive progression and poor prognosis in human cervical cancer. Diagn Pathol. 2013;8:69. doi:10.1186/1746-1596-8-69.
Veerla S, Lindgren D, Kvist A, Frigyesi A, Staaf J, Persson H, et al. MiRNA expression in urothelial carcinomas: important roles of miR-10a, miR-222, miR-125b, miR-7 and miR-452 for tumor stage and metastasis, and frequent homozygous losses of miR-31. Int Cancer. 2009;124:2236–42.
Van Schooneveld E, Wouters MC, van der Auwera I, Peeters DJ, Wildiers H, Van Dam PA, et al. Expression profiling of cancerous and normal breast tissues identifies micro RNAs that are differentially expressed in serum from patients with (metastatic) breast cancer and healthy volunteers. Breast Cancer Res. 2012;14:R34.
COLOFOL steering group, Christensen LL, Tobiasen H, Holm A, Schepeler T, Ostenfeld MS, et al. MiRNA-362-3p induces cell cycle arrest through targeting of E2F1, USF2 and PTPN1 and is associated with recurrence of colorectal cancer. Int J Cancer. 2013;133:67–78.
Ma J, Mannoor K, Gao L, Tan A, Guarnera MA, Zhan M, et al. Characterization of microRNA transcriptome in lung cancer by next-generation deep sequencing. Mol Oncol. 2014. doi:10.1016/j.molonc.
Ali S, Ahmad A, Aboukameel A, Ahmed A, Bao B, Banerjee S, et al. Deregulation of miR-146a expression in a mouse model of pancreatic cancer affecting EGFR signaling. Cancer Lett. 2014;351:134–42.
Wu C, Li M, Hu C, Duan H. Prognostic role of microRNA polymorphisms in patients with advanced esophageal squamous cell carcinoma receiving platinum-based chemotherapy. Cancer Chemother Pharmacol. 2013;73:335–41.
Ke Y, Zhao W, Xiong J, Cao R. miR-149 inhibits non-small-cell lung cancer cells EMT by targeting FOXM1. Biochem Res Int. 2013. doi:10.1155/2013/506731.
Mairinger FD, Ting S, Werner R, Walter RF, Hager T, Vollbrecht C, et al. Different micro-RNA expression profiles distinguish subtypes of neuroendocrine tumors of the lung: results of a profiling study. Mod Pathol. 2014. doi:10.1038/modpathol.2014.74.
Duellman T, Warren C, Yang J. Single nucleotide polymorphism-specific regulation of matrix metalloproteinase-9 by multiple miRNAs targeting the coding exon. Nucleic Acids Res. 2014;42:5518–31.
Acknowledgments
We acknowledge Dr. Thomas Benen, Malven Instruments, Herrenberg, for the skillful introduction of NSA instrument for visualization and specification of size and concentration of the exosome samples. We are thankful for excellent technical assistance of Michaela Heitmann, Susanne Neiß, and Anke Wienand-Dorweiler.
Conflicts of interest
None
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Warnecke-Eberz, U., Chon, SH., Hölscher, A.H. et al. Exosomal onco-miRs from serum of patients with adenocarcinoma of the esophagus: comparison of miRNA profiles of exosomes and matching tumor. Tumor Biol. 36, 4643–4653 (2015). https://doi.org/10.1007/s13277-015-3112-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13277-015-3112-0