Abstract
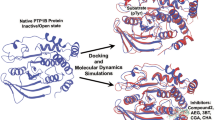
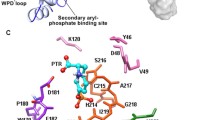
Protein-tyrosine phosphatase 1B (PTP1B) is an attractive drug target for type II diabetes and obesity. The structural motions of its S-loop play crucial roles in WPD-loop closure that is essential for the catalytic mechanism of this protein. In the current studies, totally 20 ns molecular dynamics simulations were employed on both PTP1B and its complex with inhibitors in the explicit solution surroundings with the periodic boundary conditions in order to perform detail exam on the structural flexibility of S-loop. Together with calculating RMSD values and monitoring the distances between active site and the residues in S-loop, it is found that S-loop can move towards to active site and form a tight binding pocket for substrates upon inhibitor binding. And a hydrogen bond network rearrangement was detected in this region, which may cause the transforms of both the tree-dimensional structure and the total accessible surfaces for the residues in S loop. Additionally, the second structures of Ser201 and Gly209 have huge changes for the open system, which is not detected in close system. These findings can reveal the possible mechanism of ligand recognitions and inhibitions, further providing useful information to design novel inhibitors against PTP1B and develop new treatment for type II diabetes and obesity.
Similar content being viewed by others
References
Appiah, E.A., Kennedy B.P. 2003. Protein tyrosine phosphatase: The quest for negative regulators of insulin action. Am J Physiol 84, E663–E670.
Barford, D., Flint, A.J., Tonks, N.K. 1994. Crystal structure of human protein tyrosine phosphatase 1B. Science 263, 1397–1404.
Berendsen, H.J.C., van der Spoel, D., van Drunen R. 1995. GROMACS: a message-passing parallel molecular dynamics implementation. Comp Phys Commun 91, 43–56.
Berman, H.M., Westbrook, J., Feng, Z., Gilliland, G., Bhat T.N., Weissig, H., Shindyalov, I.N., Bourne, P.E. 2000. The protein data bank. Nucleic Acids Res 28, 235–242.
Cheng, A., Tremblay, M.L. 2004. Insulin receptor PTP: PTP1B. Handb Cell Signal 1, 729–732.
Cook, W.S., Unger, R.H. 2002. Protein tyrosine phosphatase 1B: A potential leptin resistance factor of obesity. Dev Cell 2, 385–387.
Darden, T., York, D., Pedersen, L. 1993. Particel mesh Ewald: an N-log(N) method for Ewald sums in large systems. J Chem Phys 98, 10089–10092.
Elchebly, M., Payette, P., Michaliszyn, E., Cromlish, W., Collins, S., Loy, A.L., Normandin, D., Cheng, A., Hagen, J.H., Chan, C.C., Ramachandran, C., Gresser, M.J., Tremblay, M.L., Kennedy, B.P. 1999. Increased insulin sensitivity and obesity resistance in mice lacking the protein tyrosine phosphatase 1B gene. Science 283, 1544–1548.
Gum, R.J., Gaede, L.L., Koterski, S.L., Heindel, M., Clampit, J.E., Zinker, B.A., Trevillyan, J.M., Ulrich, R.G., Jirousek, M.R., Rondinone, C.M. 2003. Reduction of protein tyrosine phosphatase 1B increases insulin-dependent signaling in ob/ob mice. Diabetes 52, 21–28.
Hess, B. 2000. Similarities between principal components of protein dynamics and random diffusion. Phys Rev E 62, 8438–8448.
Hunter, T. 1995. Protein kinases and phosphatase: the Yin and Yang of protein phosphorylation and signalling. Cell 80, 225–236.
Jia, Z., Barford, D., Flint, A.J., Tonks, N.K. 1995. Structural basis for phosphotyrosine peptide recognition by protein tyrosine phosphatase 1B. Science 268, 1754–1758.
Johnson, T.O., Ermolieff, J., Jirousek, M.R. 2002. Protein tyrosine phosphatase 1B inhibitors for diabetes. Nat Rev Drug Discov 1, 696–708.
Klaman, L.D., Boss, O., Peroni, O.D., Kim, J.K., Martino, J.L., Zabolotny, J.M., Moghal, N., Lubkin, M., Kim, Y.B., Sharpe, A.H., Krongrad, A.S., Shulman, G.I., Neel, B.G., Kahn, B.B. 2000. Increased energy expenditure, decreased adiposity, and tissue-specific insulin sensitivity in protein-tyrosine phosphatase 1Bdeficient mice. Mol Cell Biol 20, 5479–5489.
Kolmodin, K., Agvist, J. 2001. The catalytic mechanism of protein tyrosine phosphatases revisited. FEBS Lett 498, 208–213.
Liu, G.X., Tan, J.Z., Niu, C.Y., Shen, J.H., Luo, X.M., Shen, X., Chen, K.X., Jiang, H.L. 2006. Molecular dynamics simulations of interaction between proteintyrosine phosphatase 1B and a bidentate inhibitor. Acta Pharm Sin 27, 100–110.
Pedersen, A.K., Peters, G.H., Moller, K.B., Iversen, L.F., Kastrup, J.S. 2004. Water-molecule network and active-site flexibility of apo protein tyrosine phosphatase 1B. Acta Crystallogr Sect D 60, 1527–1534.
Peters, G.H., Frimurer, T.M., Andersen, J.N., Olsen, O.H. 2000. Molecular dynamics simulations of proteintyrosine phosphatase 1B. II. Substrate-enzyme interactions and dyanmics. Biophys J 78, 2191–2200.
Stone, R.L., Dixon, J.E. 1994. Protein tyrosine phosphatases. J Biol Chem 269, 31323–31326.
Tonks, N.K., Diltz, C.D., Fischer, E.H. 1988. Purification of the major protein tyrosine phosphatases of human placenta. J Biol Chem 263, 6715–6721.
Van Aalten, D.M.F., Bywater, R., Findlay, J.B., Hendlich, M., Hooft, R.W., Vriend, G. 1996. PRODRG: a program for generating molecular topologies and unique molecular descriptors from coordinates of small molecules. J Comput Aided Mol Des 10, 255–262.
Van Aalten, D.M.F., De Groot, B.L., Findlay, J.B.C., Berendsen, H.J.C., Amadei, A. 1997. A comparison of techniques for calculating protein essential dyanmics. J Comput Chem 18, 169–181.
Wang, J.F., Wei, D.Q., Chen, C., Li, Y.X., Chou, K.C. 2008. Molecular modeling of two CYP2C19 SNPs and its implications for personalized drug design. Protein Pept Lett 15, 27–32.
Wang, J.F., Wei, D.Q., Du, H.L., Li, Y.X., Chou, K.C. 2008. Molecular modeling studies on NADP-dependent of Candida tropicallis strain xylose reductase. Open Bioinformatics J 2, 89–96.
Wang, J.F., Wei, D.Q., Li, L., Zheng, S.Y., Li, Y.X., Chou, K.C. 2007. 3D structure modeling of cytochrome P450 2C19 and its implication for personalized drug design. Biochem Biophys Res Commun 355, 513–519.
Wang, J.F., Wei, D.Q., Lin, Y., Du, H.L., Li, Y.X., Chou, K.C. 2007. Insights from modeling the 3D structure of NAD(P)H-dependent D-xylose reductase of Pichia stipitis and its binding interactions with NAD and NADP. Biochem Biophys Res Commun 359, 323–329.
Wiesmann, C., Barr, K.J., Kung, J., Zhu, J., Erlanson, D.A., Shen, W., Fahr, B.J., Zhong, M., Taylor, L., Randall, M., McDowell, R.S., Hansen, S.K. 2004. Allosteric inhibition of protein tyrosine phosphatase 1B. Nat Struct Mol Biol 11, 730–737.
Wilson, D.P., Wan, Z.K., Xu, W.X., Kirincich, S.J., Follows, B.C., McCarthy, D.J., Foreman, K., Moretto, A., Wu, J.J., Zhu, M., Binnun, E., Zhang, Y.L., Tam, M., Erbe, D.V., Tobin, J., Xu, X., Leung, L., Shilling, A., Tam, S.Y., Mansour, T.S., Lee, J. 2007. Structurebased optimization of protein tyrosine phosphatase 1B inhibitors: From the active site to the second phosphotyrosine binding site. J Med Chem 50, 4681–4698.
Zinker, B.A., Rondinone, C.M., Trevillan, J.M., Gum, R.J., Clampit, J.E., Waring, J.F., Xie, N., Wilcox, D., Jacobson, P., Frost, L., Kroeger, P.E., Reilly, R.M., Koterski, S., Opgenorth, T.J., Ulrich, R.G., Rosby, S., Butler, M., Murray, S.F., McKay, R.A., Bhanot, S., Monia, B.P., Jirousek, M.R. 2002. PTP1B antisense oligo-nucleotide lowers PTP1B protein, normalizesblood glucose, and improves insulin sensitivity in dibetic mice. Proc Natl Acad Sci USA 99, 11357–11362.
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Wang, JF., Gong, K., Wei, DQ. et al. Structural flexibility and interactions of PTP1B’s S-loop. Interdiscip Sci Comput Life Sci 1, 214–219 (2009). https://doi.org/10.1007/s12539-009-0047-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12539-009-0047-4