Abstract
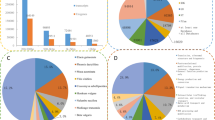
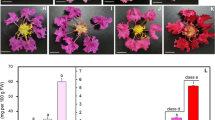
The genus Calanthe includes species of terrestrial orchids that produce attractive flowers with diverse floral traits. Breeding programs have been established to improve the horticultural value of various Calanthe species, but studies to identify the genetic components contributing to the key phenotypic characteristics have not been undertaken. To understand the molecular mechanisms underlying floral development associated with floral morphology, color, and fragrance production, the flower buds of two typical Korean Calanthe species, C. discolor and C. sieboldii, were subjected to gene expression analysis by differential display RT-PCR (DDRT-PCR). A total of 66 non-redundant differentially expressed genes (DEGs) were isolated and sequenced. Of these, 26 and 40 DEGs were found to be highly expressed in C. discolor and C. sieboldii, respectively. Moreover, the expression patterns of a subset of genes presumably implicated in signal transduction, metabolic pathways, and hormonal signaling differed between the two species. The data presented here may improve our understanding of the mechanisms underlying floral development and contribute to advances in orchid biotechnology.



Similar content being viewed by others
Abbreviations
- DEG:
-
Differentially expressed gene
- RT-PCR:
-
Reverse transcription-polymerase chain reaction
- ACP:
-
Annealing control primer
References
Bartley GE, Scolnik PA (1989) Carotenoid biosynthesis in photosynthetic bacteria: genetic characterization of the Rhodobacter capsulatus CrtI protein. J Biol Chem 264:13109–13113
Cho DH, Chung MY, Jee SO, Kim CK, Chung JD, Kim KM (2009) Genetic analysis of flower color traits in Calanthe discolor, C. sieboldii, and variants using molecular linkage map. J Life Sci 19:1239–1244
Gale S, Drinkell C (2007) Calanthe arisanensis Orchidaceae. The Board of Trustees of the Royal Botanic Gardens, Kew. Plate 596:206–210
Gómez-Gómez L, Carrasco P (1998) Differential expression of the S-Adenosyl-l-Methionine synthase genes during pea development. Plant Physiol 117:397–405
Goto T, Kondo T (1991) Structure and molecular stacking of anthocyanins – flower color variation. Angew Chem Int Ed Engl 30:17–33
Hamberger B, Hahlbrock K (2004) The 4-coumarate:CoA ligase gene family in Arabidopsis thaliana comprises one rare, sinapate-activating and three commonly occurring isoenzymes. Proc Natl Acad Sci USA 101:2209–2214
Holton TA, Cornish EC (1995) Genetics and biochemistry of anthocyanin biosynthesis. Plant Cell 7:1071–1083
Hyun MR, Choi JY, Suh JN, So IS, Lee JS (1999a) Isozyme and randomly amplified polymorphic DNA (RAPD) analysis for genetic relationship among Calanthe discolor, C. sieboldii, and C. bicolor native to Cheju Province. Kor J Hort Sci Technol 17:141–143
Hyun MR, Choi JY, Suh JN, So IS, Lee JS (1999b) Studies on distributions and morphological characteristics of Calanthe discolor, C. sieboldii, and C. bicolor native to Cheju Province. Kor J Hort Sci Technol 17:497–499
Iwatsuki K (1995) Dryopteridaceae. In: Iwatsuki K, Yamazaki T, Boufford DE, Ohba H (eds) Pteridophyta and Gymnospermae. Flora of Japan, vol 1. Kodansha, Tokyo, pp 120–173
Kim YS, Kim SH (1989) A taxonomic study on Calanthe in Korea. Kor J Plant Tax 19:273–287
Kim YJ, Kwak CI, Gu YY, Hwang IT, Chun JY (2004) Annealing control primer system for identification of differentially expressed genes on agarose gels. Biotechniques 36:424–434
Kim KS, Kim JS, Park JH (2008) Aseptic germination of F1 hybrid seed by inter-species pollination of Calanthe discolor Lindl. and C. discolor for C. sieboldii (Decne.) Ohwi. Kor J Plant Res 21:341–345
Knudsen JT, Tollsten L, Bergstrom G (1993) Floral scents: a checklist of volatile compounds isolated by head-space techniques. Phytochemistry 33:253–280
Lee JS, Kwack BH (1983) Classification of horticultural cultivars on cultivated Calanthe discolor Lindle native to Korea. J Kor Soc Hort Sci 24:144–148
Ludwig SR, Wessler SR (1990) Maize R gene family: tissue specific helix–loop–helix proteins. Cell 62:849–851
Memon AR (2004) The role of ADP-ribosylation factor and SAR1 in vesicular trafficking in plants. Biochim Biophys Acta 1664:9–30
Mondragón-Palomino M, Theissen G (2009) Why are orchid flowers so diverse? Reduction of evolutionary constraints by paralogues of class B floral homeotic genes. Ann Bot (Lond) 104:583–594
Nemhauser JL, Feldman LJ, Zambryski PC (2000) Auxin and ETTIN in Arabidopsis gynoecium morphogenesis. Development 127:3877–3888
Okada K, Ueda J, Komaki MK, Bell CJ, Shimura Y (1991) Requirement of the auxin polar transport system in early stages of Arabidopsis floral bud formation. Plant Cell 3:677–684
Pfluger J, Zambryski P (2004) The role of SEUSS in auxin response and floral organ patterning. Development 131:4697–4707
Silveira P, Schuiteman A, Vermeulen JJ, Sousa AJ, Silva H, Paiva JP, Ogel ED (2008) The orchids of Timor: checklist and conservation status. Bot J Linn Soc 157:197–215
Stafford HA (1994) Anthocyanins and betalains: evolution of the mutually exclusive pathways. Plant Sci 101:91–98
Tanaka Y, Ohmiya A (2008) Seeing is believing: engineering anthocyanin and carotenoid biosynthetic pathways. Curr Opin Biotechnol 19:190–197
Vernoux T, Kronenberger J, Grandjean O, Laufs P, Traas J (2000) PIN-FORMED 1 regulates cell fate at the periphery of the shoot apical meristem. Development 127:5157–5165
Yang SF, Hoffman NE (1984) Ethylene biosynthesis and its regulation in higher plants. Annu Rev Plant Physiol 35:155–189
Yang T, Poovaiah BW (2003) Calcium/calmodulin-mediated signal network in plants. Trends Plant Sci 8:505–512
Acknowledgements
We thank Dr. Hwang Inhwan (POSTECH, Korea) for providing the pBluescript-T vector. This work was supported by research funds from Chonbuk National University in 2007 (NP-2007-180103), the Korea Research Foundation Grant (KRF2007-412-J05501), and a grant from the Jeonbuk Forest Environment Research Institute.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Park, J.M., Whang, S.S., So, S. et al. Identification of Differentially Expressed Genes in Flower Buds of Calanthe discolor and C. sieboldii . J. Plant Biol. 53, 24–31 (2010). https://doi.org/10.1007/s12374-009-9082-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12374-009-9082-2