Abstract
Background
Long non-coding RNAs (LncRNAs) utilize a wide variety of mechanisms to regulate RNAs or proteins on the transcriptional or post-transcriptional levels. Accumulating studies have identified numerous LncRNAs to exert critical effects on different physiological processes, genetic disorders, and human diseases.
Materials and methods
Both clinical tissues from breast cancer patients and cultured cells were used for the qRT-PCR analysis. Specific siRNAs were included to assess the roles of TUG1 with cell viability assay, transwell assay, and cell apoptosis assay, respectively.
Results
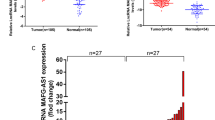
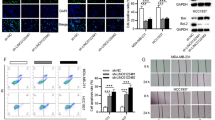
The expression of TUG1 was enhanced in breast cancerous tissues and in highly invasive breast cancer cell lines and was associated with clinical variables, including tumor size, distant metastasis and TNM staging. Knockdown of TUG1 significantly slowed down cell proliferation, cell migration, and invasion in breast cancer cell lines MDA-MB-231 and MDA-MB-436. In addition, cell apoptotic rate was shown to increase upon siTUG1 treatment as evidenced by increases of the activities of caspase-3 and caspase-9.
Conclusion
The identification of TUG1 as a critical mediator of breast cancer progression implied that it might serve as a biomarker for the diagnosis and treatment of breast cancer in clinic.




Similar content being viewed by others
References
Tao Z, Shi A, Lu C, et al. Breast cancer: epidemiology and etiology. Cell Biochem Biophys. 2015;72(2):333–8.
Dittrich A, Gautrey H, Browell D, et al. The HER2 signaling network in breast cancer-like a spider in its web. J Mammary Gland Biol Neoplasia. 2014;19:253–70.
Ferlay J, Soerjomataram I, Dikshit R, et al. Cancer incidence and mortality worldwide: sources, methods and major patterns in GLOBOCAN 2012. Int J Cancer. 2015;136:E359–86.
DeSantis CE, Bray F, Ferlay J, et al. International variation in female breast cancer incidence and mortality rates. Cancer Epidemiol Biomarkers Prev. 2015;24:1495–506.
Cazap E, Buzaid A, Garbino C, et al. Breast cancer in Latin America: experts perceptions compared with medical care standards. Breast. 2010;19:50–4.
Chatenoud L, Bertuccio P, Bosetti C, et al. Trends in mortality from major cancers in the Americas: 1980–2010. Ann Oncol. 2014;25:1843–53.
Meissner HI, Klabunde CN, Han PK, et al. Breast cancer screening beliefs, recommendations and practices: primary care physicians in the United States. Cancer. 2011;117:3101–11.
Lee EY, Muller WJ. Oncogenes and tumor suppressor genes. Cold Spring Harb Perspect Biol. 2010;2:a3236.
Li L, Gao P, Li Y, et al. JMJD2A-dependent silencing of Sp1 in advanced breast cancer promotes metastasis by downregulation of DIRAS3. Breast Cancer Res Treat. 2014;147:487–500.
Djebali S, Davis CA, Merkel A, et al. Landscape of transcription in human cells. Nature. 2012;489:101–8.
Derrien T, Johnson R, Bussotti G, et al. The GENCODE v7 catalog of human long noncoding RNAs: analysis of their gene structure, evolution, and expression. Genome Res. 2012;22:1775–89.
Melissari MT, Grote P. Roles for long non-coding RNAs in physiology and disease. Pflugers Arch. 2016;5:1–4.
Liu H, Li J, Koirala P, et al. Long non-coding RNAs as prognostic markers in human breast cancer. Oncotarget. 2016;7(15):20584–96.
Malih S, Saidijam M, Malih N. A brief review on long noncoding RNAs: a new paradigm in breast cancer pathogenesis, diagnosis and therapy. Tumour Biol. 2016;37(2):1479–85.
Young TL, Matsuda T, Cepko CL. The noncoding RNA taurine upregulated gene 1 is required for differentiation of the murine retina. Curr Biol. 2005;15:501–12.
Sun J, Ding C, Yang Z, et al. The long non-coding RNA TUG1 indicates a poor prognosis for colorectal cancer and promotes metastasis by affecting epithelial-mesenchymal transition. J Transl Med. 2016;14:42.
Li J, Zhang M, An G, et al. LncRNA TUG1 acts as a tumor suppressor in human glioma by promoting cell apoptosis. Exp Biol Med (Maywood). 2016;241(6):644–9.
Fan L, Strasser-Weippl K, Li JJ, et al. Breast cancer in China. Lancet Oncol. 2014;15:e279–89.
Guler G, Himmetoglu C, Jimenez RE, et al. Aberrant expression of DNA damage response proteins is associated with breast cancer subtype and clinical features. Breast Cancer Res Treat. 2011;129:421–32.
Gonzalez-Angulo AM, Timms KM, Liu S, et al. Incidence and outcome of BRCA mutations in unselected patients with triple receptor-negative breast cancer. Clin Cancer Res. 2011;17:1082–9.
Zhang E, He X, Yin D, et al. Increased expression of long noncoding RNA TUG1 predicts a poor prognosis of gastric cancer and regulates cell proliferation by epigenetically silencing of p57. Cell Death Dis. 2016;7:e2109.
Lamouille S, Xu J, Derynck R. Molecular mechanisms of epithelial-mesenchymal transition. Nat Rev Mol Cell Biol. 2014;15:178–96.
Yang J, Mani SA, Donaher JL, et al. Twist, a master regulator of morphogenesis, plays an essential role in tumor metastasis. Cell. 2004;117:927–39.
Pandey MK, Prasad S, Tyagi AK, et al. Targeting cell survival proteins for cancer cell death. Pharmaceuticals (Basel). 2016;9(1):11.
Hale AJ, Smith CA, Sutherland LC, et al. Apoptosis: molecular regulation of cell death. Eur J Biochem. 1996;236:1–26.
Probst-Cousin S, Rickert CH, Schmid KW, et al. Cell death mechanisms in multiple system atrophy. J Neuropathol Exp Neurol. 1998;57:814–21.
Mohan S, Abdul AB, Abdelwahab SI, et al. Typhonium flagelliforme induces apoptosis in CEMss cells via activation of caspase-9, PARP cleavage and cytochrome c release: its activation coupled with G0/G1 phase cell cycle arrest. J Ethnopharmacol. 2010;131:592–600.
Tamura K, Takayama S, Ishii T, et al. Apoptosis and differentiation of Xenopus tail-derived myoblasts by thyroid hormone. J Mol Endocrinol. 2015;54:185–92.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
None.
About this article
Cite this article
Li, T., Liu, Y., Xiao, H. et al. Long non-coding RNA TUG1 promotes cell proliferation and metastasis in human breast cancer. Breast Cancer 24, 535–543 (2017). https://doi.org/10.1007/s12282-016-0736-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12282-016-0736-x