Abstract
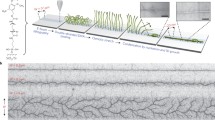
The DNA-array to protein-array technology (DAPA) allows the direct transcription and translation of a coded DNA-array to a protein array in the presence of a cell free expression system. The coupling efficiency of DNA and of the corresponding immobilized proteins is enhanced by using 3-dimensional epoxy surfaces. The production time of protein arrays is considerably reduced and the DNA template array can be reused for producing further protein arrays.
Similar content being viewed by others
Literatur
Stoevesandt O, Taussig MJ, He M (2009) Protein microarrays: high throughput tools for proteomics. Expert Rev Proteomics 6:145–157
Jeong JS, Jiang L, Albino E et al. (2012) Rapid identification of monospecific monoclonal antibodies using a human proteome microarray. Mol Cell Proteomics 11:O111.016253
Hu CJ, Song G, Huang W et al. (2012) Identification of new autoantigens for primary biliary cirrhosis using human proteome microarrays. Mol Cell Proteomics 11:669–680
He M, et al. (2008) Printing protein arrays from DNA arrays. Nat Methods 5:175–177
Stoevesandt O (2012) Protein arraying by cell-free expression and diffusion across a fluid-filled gap. N Biotechnol 29:586–588
Schmidt R, et al. (2013) Optimised ‘on demand’ protein arraying from DNA by cell free expression with the ‘DNA to Protein Array’ (DAPA) technology. J Proteomics 88:141–148
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Stoevesandt, O., Schmidt, R., Heise, C. et al. 3D-Epoxyoberflächen für DNA-codierte Proteinarrays. Biospektrum 20, 453–455 (2014). https://doi.org/10.1007/s12268-014-0461-y
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12268-014-0461-y