Abstract
Objective
To explore the differentially expressed proteins in normal cervix, cervical intraepithelial neoplasia (CIN) and cervical squamous cell carcinoma (CSCC) tissues by differential proteomics technique.
Methods
Cervical tissues (including normal cervix, CIN and CSCC) were collected in Department of Gynecologic Oncology of Beijing Obstetrics and Gynecology Hospital. Two-dimensional fluorescence difference in gel electrophoresis (2-D DIGE) and DeCyder software were used to detect the differentially expressed proteins. Matrix-assisted laser desorption/ionization-time-of-flight tandem mass spectrometry (MALDI-TOF/TOF MS) was used to identify the differentially expressed proteins. Western blot (WB) and immunohistochemistry (IHC) were performed to validate the expressions of selected proteins among normal cervix, CIN and CSCC.
Results
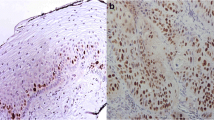
2-D DIGE images with high resolution and good repeatability were obtained. Forty-six differentially expressed proteins (27 up-regulated and 19 down-regulated) were differentially expressed among the normal cervix, CIN and CSCC. 26 proteins were successfully identified by MALDI-TOF/TOF MS. S100A9 (S100 calcium-binding protein A9) was the most significantly up-regulated protein. Eukaryotic elongation factor 1-alpha-1 (eEF1A1) was the most significantly down-regulated protein. Pyruvate kinase isozymes M2 (PKM2) was both up-regulated and down-regulated. The results of WB showed that with the increase in the severity of cervical lesions, the expression of S100A9 protein was significantly increased among the three groups (P = 0.010). The expression of eEF1A1 was reduced but without significant difference (P = 0.861). The expression of PKM2 was significantly reduced (P = 0.000). IHC showed that protein S100A9 was mainly expressed in the cytoplasm, and its positive expression rate was 20.0 % in normal cervix, 70.0 % in CIN and 100.0 % in CSCC, with a significant difference among them (P = 0.006). eEF1A1 was mainly expressed in the cell plasma, and its positive expression rate was 70.0 % in normal cervix, 73.3 % in CIN and 60.0 % in CSCC tissues, without significant difference among them (P = 0.758). PKM2 was mainly expressed in the cell nuclei, and its positive expression rate was 100.0 % in normal cervix, 93.3 % in CIN and 75.0 % in CSCC tissues, showing a difference close to statistical significance (P = 0.059) among them.
Conclusions
There are differentially expressed proteins among normal cervix, CIN and CSCC. S100A9, eEF1A1 and PKM2 may become candidate markers for early diagnosis of cervical cancer and new targets for therapy. It also provides a basis for further studies of the mechanism for CIN developing to CSCC.






Similar content being viewed by others
References
Parkin DM, Bray F, Ferlay J, Pisani P. Global cancer statistics, 2002. CA Cancer J Clin. 2005;55(2):74–108.
Zhao FH, Hu SY, Zhang SW, Chen WQ, Qiao YL. Cervical cancer mortality in 2004–2005 and changes during last 30 years in China. Chin J Prev Med. 2010;44(5):408–12.
Ca´rdenas-Turanzas M, Follen M, Nogueras-Gonzalez GM, Benedet JL, Beck JR, Cantor SB. The accuracy of the Papanicolaou smear in the screening and diagnostic settings. J Low Genit Tract Dis. 2008;12:269–75.
Monsonego J. Cervical cancer prevention—current perspectives. Endocr Dev. 2012;22:222–9.
Woods AG, Sokolowska I, Wetie AG, Wormwood K, Aslebagh R, Patel S, et al. Mass spectrometry for proteomics-based investigation. Adv Exp Med Biol. 2014;806:1–32.
Zhao Q, Duan W, Wu YM, Qian XH, Deng XH. Analysis of serum biomarkers of ovarian epithelial cancers based on 2-DE DIGE and MALDI TOF/TOF. Chin J Oncol. 2008;30(10):754–8.
Boylan KL, Afiuni-Zadeh S, Geller MA, Hickey K, Griffin TJ, Pambuccian SE, et al. A feasibility study to identify proteins in the residual Pap test fluid of women with normal cytology by mass spectrometry-based proteomics. Clin Proteomics. 2014;11(1):30.
Marenholz I. HeizmannCW, Fritz G. S100 proteins in mouse and man: from evolution to function and pathology (including an update of the nomenclature). Biochem Biophys Res Commun. 2004;322:1111–22.
Salama I, Malone PS, Mihaimeed F, Jones JL. A review of the S100 proteins in cancer. Eur J Surg Oncol. 2008;34(4):357–64.
Kim HK, Reyzer ML, Choi IJ, Kim CG, Kim HS, Oshima A. Gastric cancer-specific protein profile identified using endoscopic biopsy samples via MALDI mass spectrometry. J Proteome Res. 2010;9(8):4123–30.
Su YJ, Xu F, Yu JP, Yue DS, Ren XB, Wang CL. Up-regulation of the expression of S100A8 and S100A9 in lung adenocarcinoma and its correlation with inflammation and other clinical features. Clin Med J. 2010;123(16):2215–20.
Wu R, Duan L, Ye L, Wang H, Yang X, Zhang Y, et al. S100A9 promotes the proliferation and invasion of HepG2 hepatocellular carcinoma cells via the activation of the MAPK signaling pathway. Int J Oncol. 2013;42(3):1001–10.
Zhu H, Pei HP, Zeng S, Chen J, Shen LF, Zhong MZ, et al. Profiling protein markers associated with the sensitivity to concurrent chemoradiotherapy in human cervical carcinoma. J Proteome Res. 2009;8(8):3969–76.
Zhu XJ, Zheng FY, Zhou SW, Jiang W, Jiang L, Zhu X. Expression and significance of S100A9 in progression of squamous cervical cancer. J Pract Med. 2010;26(8):1350–2.
Qin F, Song Y, Li Z, Zhao L, Zhang Y, Geng L. S100A8/A9 induces apoptosis and inhibits metastasis of CasKi human cervical cancer cells. Pathol Oncol Res. 2010;16(3):353–60.
Rodríguez-Pérez MA, Medina-Aunon A, Encarnación-Guevara SM, Bernal-Silvia S, Barrera-Saldaña H, Albar-Ramírez JP. In silico analysis of protein neoplastic biomarkers for cervix and uterine cancer. Clin Transl Oncol. 2008;10(10):604–17.
Lamberti A, Caraglia M, Longo O, Marra M, Abbruzzese A, Arcari P. The translation elongation factor 1A in tumorigenesis, signal transduction and apoptosis: review article. Amino Acids. 2004;26(4):443–8.
Leclercq TM, Moretti PA, Pitson SM. Guanine nucleotides regulate sphingosine kinase 1 activation by eukaryotic elongation factor 1A and provide a mechanism for eEF1A-associated oncogenesis. Oncogene. 2011;30(3):372–3.
Thornton S, Anand N, Purcell D, Lee J. Not just for housekeeping: protein initiation and elongation factors in cell growth and tumorigenesis. J Mol Med. 2003;81:536–48.
Al-Maghrebi M, Anim JT, Olalu AA. Up-regulation of eukaryotic elongation factor-1 subunits in breast carcinoma. Anticancer Res. 2005;25(3c):2573–7.
Li H, DeSouza LV, Ghanny S, Li W, Romaschin AD, Colgan TJ, et al. Identification of candidate biomarker proteins released by human endometrial and cervical cancer cells using two-dimensional liquid chromatography/tandem mass spectrometry. J Proteome Res. 2007;6(7):2615–22.
Mazurek S. Pyruvate kinase type M2: a key regulator of the metabolic budget system in tumor cells. Int J Biochem Cell Biol. 2011;43(7):969–80.
Wong N, De Melo J, Tang D. PKM2, a central point of regulation in Cancer metabolism. Int J Cell Biol. 2013;2013:242513.
Laudt S, Jeschke S, Koeninger A, Thomas A, Heusner T, Korlach S, et al. Tumor-specific correlation of tumor M2 pyruvate kinase in pre-invasive, invasive and recurrent cervical cancer. Anticancer Res. 2010;30(2):375–81.
Bai XH, Kong L, Zhang AR, Jia JH, Zhang WF, Tang W, et al. Differential expression of M2-PK in normal cervix and cervical cancer tissues. J Shandong Univ (Health Sci). 2006;44(10):981–4.
Kong L, Deng Z, Zhao Y, Wang Y, Sarkar FH, Zhang Y. Down-regulation of phospho-non-receptor Src tyrosine kinases contributes to growth inhibition of cervical cancer cells. Med Oncol. 2011;28(4):1495–506.
Zhao EF, Bao L, Liang L, Li D. Expressions of heat shock protein in cervical cancer and precancerosis. Zhongguo Ying Yong Sheng Li Xue Za Zhi. 2006;22(2):250–3.
Acknowledgments
This work was support by the Beijing Municipal Commission of Education Science and Technology Development Plan (KM200910025025).
Conflict of interest
The authors declare no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Zhao, Q., He, Y., Wang, XL. et al. Differentially expressed proteins among normal cervix, cervical intraepithelial neoplasia and cervical squamous cell carcinoma. Clin Transl Oncol 17, 620–631 (2015). https://doi.org/10.1007/s12094-015-1287-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12094-015-1287-x