Abstract
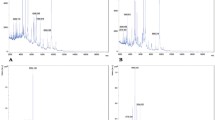
The present study demonstrates isolation and identification of methicillin resistance Staphylococcus aureus (MRSA) strains in the samples collected from burn patients. About 106 swab samples were collected from burn patients of >40% burn injury and were subjected to isolation using nutrient agar followed by screening using Me Re Sa selective medium agar. A total of 10 isolates with identity to S. aureus were obtained and further authenticated using Polymerase Chain Reaction and matrix assisted laser desorption/ionization time of flight mass spectrometry analysis. Presence of mec A gene and the peak pattern observations suggested seven of the 10 isolates are MRSA. Thus, the present study emphasizes the process of identification of MRSA using two different bio-analytical techniques, which authenticate the presence of MRSA.





Similar content being viewed by others
References
Panlilio AL, Culver DH, Gaynes RP, Banerjee S, Henderson TS, Tolson JS, Martone WJ (1992) Methicillin-resistant Staphylococcus aureus in U.S. Hospitals, 1975–1991. Infect Contr Hosp Epidemiol 13:582–586
Kuehnert MJ, Hill HA, Kupronis BA, Tokars JI, Solomon SL, Jernigan DB (2005) Methicillin-resistant-Staphylococcus aureus hospitalizations, United States. Emerg Infect Diseases 11:868–872
Cosgrove SE, Qi YL, Kaye KS, Harbarth S, Karchmer AW, Carmeli Y (2005) The impact of methicillin-resistance in Staphylococcus aureus bacteremia on patient outcomes: mortality, length of stay, and hospital charges. Infect Contr Hosp Epidemiol 26:166–174
Abramson MA, Sexton DJ (1999) Nosocomial methicillin-resistant and methicillin-susceptible, Staphylococcus aureus primary bacteremia: at what costs. Infect Contr Hosp Epidemiol 20:408–411
Witte W, Braulke C, Heuck D, Cuny C (1994) Analysis of nosocomial outbreaks with multiply and methicillin-resistant Staphylococcus aureus (MRSA) in Germany: implications for hospital hygiene. Infection 22:128–134
Krishnamurthy T, Ross PL (1996) Rapid identification of bacteria by direct matrix-assisted laser desorption/ionization mass spectrometric analysis of whole cells. Rapid Commun Mass Spectrom 10:1992–1996
Eady EA, Cove JH (2003) Staphylococcal resistance revisited: community-acquired methicillin resistant Staphylococcus aureus-an emerging problem for the management of skin and soft tissue infections. J Clin Microbiol 16:103–124
Velasco D, del Mar Tomas M, Cartelle M, Beceiro A, Perez A, Molina F, Moure R, Villanueva R, Bou G (2005) Evaluation of different methods for detecting methicillin (oxacillin) resistance in Staphylococcus aureus. J Antimicrob Chemother 55:379–382
Cauwelier B, Gordts B, Descheemaecker P, Van Landuyt H (2004) Evaluation of a disk diffusion method with cefoxitin (30 μg) for detection of methicillin-resistant Staphylococcus aureus. Eur J Clin Microbiol Infect Dis 23:389–392
Bernardo K, Pakulat N, Macht M, Krut O, Seifert H, Fleer S, Hünger F, Krönke M (2002) Identification and discrimination of Staphylococcus aureus strains using matrix-assisted laser desorption/ionization-time of flight mass spectrometry. Proteomics 2:747–753
Gregersen T (1978) Rapid method for distinction of Gram-negative from Gram-positive bacteria. App Microbiol Biotechnol 5:123–127
Harbarth S, Dharan S, Liassine N, Herrault P, Auckenthaler R, Pittet D (1999) Randomized, placebo-controlled, double-blind trial to evaluate the efficacy of mupirocin for eradicating carriage of methicillin-resistant Staphylococcus aureus. Antimicrob Agents Chemother 43:1412–1416
Charyulu EM, Sekaran G, Rajakumar GS, Gnanamani A (2009) Antimicrobial activity of secondary metabolite from marine isolate, Pseudomonas sp. against gram positive and negative bacteria including MRSA. Indian J Exp Biol 47:964–968
Ghebremedhin B, Layer F, Konig W, Konig B (2008) Genetic classification and distinguishing of staphylococcus species based on different partial gap, 16S rRNA, hsp60, rpoB, sodA, and tuf gene sequences. J Clin Microbiol 46:1019–1025
Petinaki E, Arvaniti A, Dimitracopoulos G, Spiliopoulou I (2001) Detection of mecA, mecR1 and mecI genes among clinical isolates of methicillin-resistant staphylococci by combined polymerase chain reactions. J Antimicrob Chemother 47:297–304
Brakstad OG, Aasbakk K, Maeland JA (1992) Detection of Staphylococcus aureus by polymerase chain reaction amplification of the nuc gene. J Clin Microbiol 30:1654–1660
Murakami K, Minamide W, Wada K, Nakamura E, Teraoka H, Watanabe S (1991) Identification of methicillin-resistant strains of staphylococci by polymerase chain reaction. J Clin Microbiol 29:2240–2244
Merlino J, Leroi M, Bradbury R, Veal D, Harbour C (2000) New chromogenic identification and detection of Staphylococcus aureus and methicillin-resistant S. aureus. J Clin Microbiol 38:2378–2380
Wichelhaus TA, Hunfeld KP, Boddinghaus B, Kraiczy P, Schafer V, Brade V (2001) Rapid molecular typing of methicillin resistant Staphylococcus aureus by PCR-RFLP. Infect Control Hosp Epidemiol 22:294–298
Demirev PA, Ho YP, Ryzhov V, Fenselau C (1999) Microorganism identification by mass spectrometry and protein database searches. Anal Chem 71:2732–2738
Edwards-Jones V, Claydon MA, Evason DJ, Walker J, Fox AJ, Gordon DB (2000) Rapid discrimination between methicillin-sensitive and methicillin-resistant Staphylococcus aureus by intact cell mass spectrometry. J Medical Microbiol 49:295–300
Davies S, Zadik PM (1997) Comparison of methods for the isolation of methicillin resistant Staphylococcus aureus. J Clin Pathol 50:257–258
Walker J, Fox AJ, Edwards-Jones V, Gordon DB (2002) Intact cell mass spectrometry (ICMS) used to type methicillin-resistant Staphylococcus aureus: media effects and inter-laboratory reproducibility. J Microbiol Methods 48:117–126
Vandenesch F, Naimi T, Enright MC, Lina G, Nimmo GR, Heffernan H, Liassine N, Bes M, Greenland T, Reverdy ME (2003) Community-acquired methicillin-resistant Staphylococcus aureus carrying Panton-Valentine leukocidin genes: worldwide emergence. Emerg Infect Dis 9:978–984
Enright MC, Robinson DA, Randle G, Feil EJ, Grundmann H, Spratt BG (2002) The evolutionary history of methicillin-resistant Staphylococcus aureus (MRSA). Proc Natl Acad Sci USA 99:7687–7692
Fluit AC, Visser MR, Schmitz FJ (2001) Molecular detection of antimicrobial resistance. Clin Microbiol Rev 14:836–871
Merlino J, Watson J, Rose B, Beard-Pegler M, Gottlieb T, Bradbury R, Harbour C (2002) Detection and expression of methicillin/oxacillin resistance in multidrug-resistant and non-multidrug-resistant Staphylococcus aureus in Central Sydney, Australia. J Antimicrob Chemother 49:793–801
Liang X, Zheng K, Qian MG, Lubman DM (1996) Determination of bacterial protein profiles by matrix-assisted laser desorption/ionization mass spectrometry with high-performance liquid chromatography. Rapid Commun Mass Spectrom 10:1219–1226
Diep BA, Chambers HF, Graber CJ, Szumowski JD, Miller LG, Han LL, Chen JH, Lin F, Lin J, Phan THV (2008) Emergence of multidrug-resistant, community-associated, methicillin-resistant Staphylococcus aureus clone USA300 in men who have sex with men. Ann Intern Med 148:249–257
Watson J, Givney R, Beard-Pegler M, Rose B, Merlino J, Vickery A, Gottlieb T, Bradbury R, Harbour C (2003) Comparative analysis of multidrug-resistant, non-multidrug-resistant, and archaic methicillin-resistant Staphylococcus aureus isolates from Central Sydney, Australia. J Clin Microbiol 41:867–872
Layer F, Ghebremedhin B, Konig W, Konig B (2006) Heterogeneity of methicillin-susceptible Staphylococcus aureus strains at a German university hospital implicates the circulating-strain pool as a potential source of emerging methicillin-resistant S. aureus clones. Am Soc Microbiol 44:2179–2185
Acknowledgments
One of the authors, Mr. E. Madhava Charyulu thanks Director, CLRI for providing the facilities to carry out the work and the author also thanks, Council of Scientific and Industrial Research for Financial Assistance in the form of Senior Research Fellowship and authors thank Dr.V.Jayaraman, Kilapuk Medical College for providing MRSA strains.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Madhava Charyulu, E., Gnanamani, A. & Mandal, A.B. Identification and Discrimination of Methicillin Resistant Staphylococcus aureus Strains Isolated from Burn Wound Sites Using PCR and Authentication with MALDI-TOF–MS. Indian J Microbiol 52, 337–345 (2012). https://doi.org/10.1007/s12088-011-0245-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12088-011-0245-8