Abstract
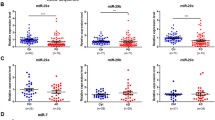
Parkinson’s disease (PD) diagnosis is based on the assessment of motor symptoms, which manifest when more than 50% of dopaminergic neurons are degenerated. To date, no validated biomarkers are available for the diagnosis of PD. The aims of the present study are to evaluate whether plasma and white blood cells (WBCs) are interchangeable biomarker sources and to identify circulating plasma-based microRNA (miRNA) biomarkers for an early detection of PD. We profiled plasma miRNA levels in 99 l-dopa-treated PD patients from two independent data collections, in ten drug-naïve PD patients, and in unaffected controls matched by sex and age. We evaluated expression levels by reverse transcription and quantitative real-time PCR (RT-qPCR) and combined the results from treated PD patients using a fixed effect inverse-variance weighted meta-analysis. We revealed different expression profiles comparing plasma and WBCs and drug-naïve and l-dopa-treated PD patients. We observed an upregulation trend for miR-30a-5p in l-dopa-treated PD patients and investigated candidate target genes by integrated in silico analyses. We could not analyse miR-29b-3p, normally expressed in WBCs, due to the very low expression in plasma. We observed different expression profiles in WBCs and plasma, suggesting that they are both suitable but not interchangeable peripheral sources for biomarkers. We revealed miR-30a-5p as a potential biomarker for PD in plasma. In silico analyses suggest that miR-30a-5p might have a regulatory role in mitochondrial dynamics and autophagy. Further investigations are needed to confirm miR-30a-5p deregulation and targets and to investigate the influence of l-dopa treatment on miRNA expression levels.


Similar content being viewed by others
References
Algarni MA, Stoessl AJ (2016) The role of biomarkers and imaging in Parkinson’s disease. Expert Rev Neurother 16(2):187–203. doi:10.1586/14737175.2016.1135056
Ammerlaan W, Betsou F (2016) Intraindividual temporal miRNA variability in serum, plasma, and white blood cell subpopulations. Biopreserv Biobank 14(5):390–397. doi:10.1089/bio.2015.0125
Andersen CL, Jensen JL, Orntoft TF (2004) Normalization of real-time quantitative reverse transcription-PCR data: a model-based variance estimation approach to identify genes suited for normalization, applied to bladder and colon cancer data sets. Cancer Res 64(15):5245–5250. doi:10.1158/0008-5472.CAN-04-0496
Ashburner M, Ball CA, Blake JA et al (2000) Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nat Genet 25(1):25–29. doi:10.1038/75556
Balzano F, Deiana M, Dei Giudici S et al (2015) miRNA stability in frozen plasma samples. Molecules 20(10):19030–19040. doi:10.3390/molecules201019030
Burgos K, Malenica I, Metpally R et al (2014) Profiles of extracellular miRNA in cerebrospinal fluid and serum from patients with Alzheimer’s and Parkinson’s diseases correlate with disease status and features of pathology. PLoS One 9(5):e94839. doi:10.1371/journal.pone.0094839
Bustin SA, Benes V, Garson JA et al (2009) The MIQE guidelines: minimum information for publication of quantitative real-time PCR experiments. Clin Chem 55(4):611–622. doi:10.1373/clinchem.2008.112797
Cai X, Janku F, Zhan Q et al (2015) Accessing genetic information with liquid biopsies. Trends Genet 31(10):564–575. doi:10.1016/j.tig.2015.06.001
Cardo LF, Coto E, de Mena L et al (2013) Profile of microRNAs in the plasma of Parkinson’s disease patients and healthy controls. J Neurol 260(5):1420–1422. doi:10.1007/s00415-013-6900-8
Chen X, Ba Y, Ma L et al (2008) Characterization of microRNAs in serum: a novel class of biomarkers for diagnosis of cancer and other diseases. Cell Res 18(10):997–1006. doi:10.1038/cr.2008.282
Cheng L, Quek CY, Sun X et al (2013a) The detection of microRNA associated with Alzheimer’s disease in biological fluids using next-generation sequencing technologies. Front Genet 4:150. doi:10.3389/fgene.2013.00150
Cheng HH, Yi HS, Kim Y et al (2013b) Plasma processing conditions substantially influence circulating microRNA biomarker levels. PLoS One 8(6):e64795. doi:10.1371/journal.pone.0064795
Connolly BS, Lang AE (2014) Pharmacological treatment of Parkinson disease: a review. JAMA 311(16):1670–1683. doi:10.1001/jama.2014.3654
Croft D, Mundo AF, Haw R et al (2014) The Reactome pathway knowledgebase. Nucleic Acids Res 42(Database issue):D472–D477. doi:10.1093/nar/gkt1102
da Silva FC, Iop RD, Vietta GG et al (2016) microRNAs involved in Parkinson’s disease: a systematic review. Mol Med Rep 14(5):4015–4022. doi:10.3892/mmr.2016.5759
D’haene B, Mestdagh P, Hellemans J et al (2012) miRNA expression profiling: from reference genes to global mean normalization. Methods Mol Biol 822:261–272. doi:10.1007/978-1-61779-427-8_18
Delenclos M, Jones DR, McLean PJ et al (2016) Biomarkers in Parkinson’s disease: advances and strategies. Parkinsonism Relat Disord 22(Suppl 1):S106–S110. doi:10.1016/j.parkreldis.2015.09.048
Devine MJ, Plun-Favreau H, Wood NW (2011) Parkinson’s disease and cancer: two wars, one front. Nat Rev Cancer 11(11):812–823. doi:10.1038/nrc3150
Egidi MG, D’Alessandro A, Mandarello G et al (2010) Troubleshooting in platelet storage temperature and new perspectives through proteomics. Blood Transfus 8(Suppl 3):s73–s81. doi:10.2450/2010.012S
Fahn S (2000) The spectrum of levodopa-induced dyskinesias. Ann Neurol 47(4 Suppl 1):S2–S9 discussion S9-11
Fahn S (2003) Description of Parkinson’s disease as a clinical syndrome. Ann N Y Acad Sci 991:1–14
Geekiyanage H, Jicha GA, Nelson PT et al (2012) Blood serum miRNA: non-invasive biomarkers for Alzheimer’s disease. Exp Neurol 235(2):491–496. doi:10.1016/j.expneurol.2011.11.026
Gelb DJ, Oliver E, Gilman S (1999) Diagnostic criteria for Parkinson disease. Arch Neurol 56(1):33–39
Hashimoto M, Bar-On P, Ho G et al (2004) Beta-synuclein regulates Akt activity in neuronal cells. A possible mechanism for neuroprotection in Parkinson’s disease. J Biol Chem 279(22):23622–23629. doi:10.1074/jbc.M313784200
Hellemans J, Vandesompele J (2011) qPCR data analysis—unlocking the secret to successful results. In: Suzanne Kennedy and Nick Oswald—MO BIO Laboratories (ed) PCR Troubleshooting and optimization: the essential guide. Caister Academic Press, p 1–13.
Higgins JP, Thompson SG, Deeks JJ et al (2003) Measuring inconsistency in meta-analyses. BMJ 327(7414):557–560. doi:10.1136/bmj.327.7414.557
Hoss AG, Labadorf A, Beach TG et al (2016) microRNA profiles in Parkinson’s disease prefrontal cortex. Front Aging Neurosci 8:36. doi:10.3389/fnagi.2016.00036
Hsu SD, Tseng YT, Shrestha S et al (2014) miRTarBase update 2014: an information resource for experimentally validated miRNA-target interactions. Nucleic Acids Res 42(Database issue):D78–D85. doi:10.1093/nar/gkt1266
Kang K, Peng X, Luo J et al (2012) Identification of circulating miRNA biomarkers based on global quantitative real-time PCR profiling. J Anim Sci Biotechnol 3(1):4 1891-3-4. doi:10.1186/2049-1891-3-4
Khoo SK, Petillo D, Kang UJ et al (2012) Plasma-based circulating microRNA biomarkers for Parkinson’s disease. J Park Dis 2:321–331. doi:10.3233/JPD-012144
Kitagishi Y, Wada Y, Matsuda S (2014) Roles of PI3K/AKT/PTEN pathway in the pathogenesis of Parkinson’s disease and the neuropsychiatric symptoms. Int Neuropsychiatr Dis J 2:1–12
Klinkenberg M, Gispert S, Dominguez-Bautista JA et al (2012) Restriction of trophic factors and nutrients induces PARKIN expression. Neurogenetics 13(1):9–21. doi:10.1007/s10048-011-0303-8
Kocer A, Yaman A, Niftaliyev E et al (2013) Assessment of platelet indices in patients with neurodegenerative diseases: mean platelet volume was increased in patients with Parkinson’s disease. Curr Gerontol Geriatr Res 2013:986254. doi:10.1155/2013/986254
Li N, Pan X, Zhang J et al (2017) Plasma levels of miR-137 and miR-124 are associated with Parkinson’s disease but not with Parkinson’s disease with depression. Neurol Sci. doi:10.1007/s10072-017-2841-9
Ma L, Wei L, Wu F et al (2013) Advances with microRNAs in Parkinson’s disease research. Drug Des Devel Ther 7:1103–1113. doi:10.2147/DDDT.S48500
Margis R, Margis R, Rieder CR (2011) Identification of blood microRNAs associated to Parkinsonis disease. J Biotechnol 152(3):96–101. doi:10.1016/j.jbiotec.2011.01.023
Martins M, Rosa A, Guedes LC et al (2011) Convergence of miRNA expression profiling, alpha-synuclein interaction and GWAS in Parkinson’s disease. PLoS One 6(10):e25443. doi:10.1371/journal.pone.0025443
Nagalla S, Shaw C, Kong X et al (2011) Platelet microRNA-mRNA coexpression profiles correlate with platelet reactivity. Blood 117(19):5189–5197. doi:10.1182/blood-2010-09-299719
Paraskevopoulou MD, Georgakilas G, Kostoulas N et al (2013) DIANA-microT web server v5.0: service integration into miRNA functional analysis workflows. Nucleic Acids Res 41(Web Server issue):W169–W173. doi:10.1093/nar/gkt393
Pogue AI, Hill JM, Lukiw WJ (2014) MicroRNA (miRNA): sequence and stability, viroid-like properties, and disease association in the CNS. Brain Res 1584:73–79. doi:10.1016/j.brainres.2014.03.042
Rao P, Benito E, Fischer A (2013) MicroRNAs as biomarkers for CNS disease. Front Mol Neurosci 6:39. doi:10.3389/fnmol.2013.00039
Razick S, Magklaras G, Donaldson IM (2008) iRefIndex: a consolidated protein interaction database with provenance. BMC Bioinformatics 9:405 2105-9-405. doi:10.1186/1471-2105-9-405
Reid TJ, LaRussa VF, Esteban G et al (1999) Cooling and freezing damage platelet membrane integrity. Cryobiology 38(3):209–224
Schlicker A, Domingues FS, Rahnenfuhrer J et al (2006) A new measure for functional similarity of gene products based on Gene Ontology. BMC Bioinformatics 7:302
Serafin A, Foco L, Zanigni S et al (2015) Overexpression of blood microRNAs 103a, 30b, and 29a in L-dopa-treated patients with PD. Neurology 84(7):645–653. doi:10.1212/WNL.0000000000001258
Shah R, Tanriverdi K, Levy D et al (2016) Discordant expression of circulating microRNA from cellular and extracellular sources. PLoS One 11(4):e0153691. doi:10.1371/journal.pone.0153691
Shinde S, Mukhopadhyay S, Mohsen G et al (2015) Biofluid-based microRNA Biomarkers for Parkinson’s disease: an overview and update. AIMS Medical Science 2(1):15–25
Sourvinou IS, Markou A, Lianidou ES (2013) Quantification of circulating miRNAs in plasma: effect of preanalytical and analytical parameters on their isolation and stability. J Mol Diagn 15(6):827–834. doi:10.1016/j.jmoldx.2013.07.005
Tanos B, Pendergast AM (2006) Abl tyrosine kinase regulates endocytosis of the epidermal growth factor receptor. J Biol Chem 281(43):32714–32723
Vandesompele J, De Preter K, Pattyn F et al (2002) Accurate normalization of real-time quantitative RT-PCR data by geometric averaging of multiple internal control genes. Genome Biol 3(7):0034
Vlachos IS, Paraskevopoulou MD, Karagkouni D et al (2015) DIANA-TarBase v7.0: indexing more than half a million experimentally supported miRNA:mRNA interactions. Nucleic Acids Res 43(Database issue):D153–D159. doi:10.1093/nar/gku1215
Wang X, McCullough KD, Franke TF et al (2000) Epidermal growth factor receptor-dependent Akt activation by oxidative stress enhances cell survival. J Biol Chem 275(19):14624–14631
Weichenberger CX, Blankenburg H, Palermo A et al (2015) Dintor: functional annotation of genomic and proteomic data. BMC Genomics 16(1):1081 015-2279-5. doi:10.1186/s12864-015-2279-5
Yasuko K, Yoko W, Satoru M (2014) Roles of PI3K/AKT/PTEN pathway in the pathogenesis of Parkinson’s disease and the neuropsychiatric symptoms. Int Neuropsychiatric Dis J 2(1):1–12
Zeka F, Mestdagh P, Vandesompele J (2015) RT-qPCR-based quantification of small non-coding RNAs. Methods Mol Biol 1296:85–102. doi:10.1007/978-1-4939-2547-6_9
Acknowledgements
The authors are grateful to the study participants for their participation and collaboration in this research project. We thank Clemens Egger and Daniele di Domizio for technical support. We thank Alessandro de Grandi and Ilaria Bozzolan for their support in biobank-related issues. The authors acknowledge the help of Agatha Eisendle, Edith Kompatscher, and Monika Mair in recruiting the study participants. The authors are grateful to Deborah Mascalzoni, Fabiola Del Greco M., and Francisco Domingues for their support in ethical, statistical, and bioinformatics issues. This work was supported by the Department for Innovation and Research and University of the Autonomous Province of Bolzano.
Author information
Authors and Affiliations
Contributions
C. Schwienbacher designed and supervised the study, contributed to the design of the sample collection, and wrote the manuscript. L. Foco designed and performed the statistical analyses and wrote the manuscript. A. Picard contributed to the design of the study, performed the experimental work, and wrote the manuscript. E. Corradi performed the experimental work and wrote the manuscript. A. Serafin contributed to the design of the study and performed the experimental work. J. Panzer performed the experimental work. S. Zanigni designed and contributed to the sample collection and clinical characterization of the patients. H. Blankenburg designed and performed the bioinformatics analyses and wrote the manuscript. M. Facheris designed and contributed to the sample collection and clinical characterization of the patients. G Giannini contributed to the sample collection and clinical characterization of the patients. M. Falla contributed to the clinical characterization of the patients. P. Cortelli critically revised the paper. P. Pramstaller designed and contributed to the sample collection and clinical characterization of the patients and critically revised the manuscript. A. Hicks critically revised the manuscript and contributed to the design of the sample collection and the study.
Corresponding authors
Ethics declarations
The local ethics committee (Comitato etico del comprensorio sanitario di Bolzano; reference numbers: 2008-D2-001090, 62/2012, 58/2013, 59/2013) approved the study, and all participants provided written informed consent.
Conflicts of Interest
C. Schwienbacher, L. Foco, A. Picard, E. Corradi, A. Serafin, J. Panzer, S. Zanigni, H. Blankenburg, M. Facheris, G. Giannini, M. Falla, and A. Hicks declare that they have no competing interests. Prof. Cortelli received honoraria for speaking engagements or consulting activities from Allergan Italia, Lilly Pharma, UCB Pharma S.p.A, Chiesi Farmaceutici, AbbVie srl, and Zambon Italy. Prof. Pramstaller received honoraria for serving on scientific boards and speaking from Novartis, Boehringer, GlaxoSmithKline, Lundbeck, and UCB.
Electronic Supplementary Material
Online Resource 1.
Clinical details of L-dopa-treated PD patients vs controls and of drug-naïve PD patients vs controls. (PDF 121 kb)
Online Resource 2.
TaqMan®miRNA assays and summary of quantitative real-time PCR experimental details. (PDF 96 kb)
Online Resource 3.
Pairplots of the differences between miRNA expression in cases-controls (SET 1, SET 2 and SET 3). The plots display the difference of expression within the matched pair for each of the studied miRNAs and for each data set. (PDF 34 kb)
Online Resource 4.
Results of the meta-analysis (SET 1 and SET 2). (PDF 7 kb)
Online Resource 5.
Forest plot of the meta-analysis for all the analysed miRNAs. The diamond indicates the pooled meta-analysis estimate of the effect (ES) and the 95% confidence interval (95% CI). I2 is provided in brackets. (PDF 4 kb)
Online Resource 6.
MiR-30a-5p target prioritization: whole data set. (XLS 1869 kb)
Online Resource 7.
Correlation between miRNA expression and PD progression in drug-naïve PD patients. (PDF 97 kb)
Online Resource 8.
Analysis on 31 matched pairs from SET 1 with miRNA expression data available in both blood and plasma. (PDF 103 kb)
Online Resource 9.
Simplified scheme of EGFR/PI3K/Akt pathway: a possible biological context for the predicted targets of miR-30a-5p. EGFR, AKT1 and DJ-1 are predicted and prioritized miR-30a-5p targets. Down-regulation of these three genes causes impairment in EGFR/PI3K/Akt pathway, which has important functions in cell survival, particularly in case of oxidative stress. White arrows: activation; red lines: inhibition. (PDF 494 kb)
Rights and permissions
About this article
Cite this article
Schwienbacher, C., Foco, L., Picard, A. et al. Plasma and White Blood Cells Show Different miRNA Expression Profiles in Parkinson’s Disease. J Mol Neurosci 62, 244–254 (2017). https://doi.org/10.1007/s12031-017-0926-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12031-017-0926-9