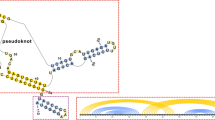
Abstract
This paper studies local connectivity of neutral networks of RNA secondary and pseudoknot structures. A neutral network denotes the set of RNA sequences that fold into a particular structure. It is called locally connected, if in the limit of long sequences, the distance of any two of its sequences scales with their distance in the n-cube. One main result of this paper is that
 is the threshold probability for local connectivity for neutral networks, considered as random subgraphs of n-cubes. Furthermore, we analyze local connectivity for finite sequence length and different alphabets. We show that it is closely related to the existence of specific paths within the neutral network. We put our theoretical results into context with folding algorithms into minimum-free energy RNA secondary and pseudoknot structures. Finally, we relate our structural findings with dynamics by discussing the role of local connectivity in the context of neutral evolution.
is the threshold probability for local connectivity for neutral networks, considered as random subgraphs of n-cubes. Furthermore, we analyze local connectivity for finite sequence length and different alphabets. We show that it is closely related to the existence of specific paths within the neutral network. We put our theoretical results into context with folding algorithms into minimum-free energy RNA secondary and pseudoknot structures. Finally, we relate our structural findings with dynamics by discussing the role of local connectivity in the context of neutral evolution.
Similar content being viewed by others
References
Akutsu, T., 2000. Dynamic programming algorithms for RNA secondary structure prediction with pseudoknots. Discrete Appl. Math. 104, 45–62.
Balister, P.N., Bollobás, B., Frieze, A.M., 2000. The first passage time diameter of the cube.
Batey, R.T., Rambo, R.P., Doudna, J.A., 1999. Tertiary motifs in RNA structure and folding. Angew. Chem. Int. Ed. 38, 2326–2343.
Chen, W.Y.C., Qin, J., Reidys, C.M., 2008. Crossings and nestings of tangled-diagrams. Electron. J. Comb. 15, 86.
Derrida, B., Peliti, L., 1999. Evolution in a flat fitness landscape. Bull. Math. Biol. 53, 355–382.
Flamm, C., Fontana, W., Hofacker, I.L., Schuster, P., 2000. RNA folding at elementary step resolution. RNA 6, 325–338.
Goebel, U., 2000. Neutral networks of minimum free energy structures. PhD Thesis, University of Vienna.
Goebel, U., Forst, C.V., 2000. RNA Pathfinder–global properties of neutral networks. Z. Phys. Chem. 216, 175–192.
Grüner, W., Giegerich, R., Strothmann, D., Reidys, C.M., Weber, J., Hofacker, I.L., Stadler, P.F., Schuster, P., 1996a. Analysis of RNA sequence structure maps by exhaustive enumeration I. Neutral networks. Chem. Mon. 127, 355–374.
Grüner, W., Giegerich, R., Strothmann, D., Reidys, C.M., Weber, J., Hofacker, I.L., Stadler, P.F., Schuster, P., 1996b. Analysis of RNA sequence structure maps by exhaustive enumeration II. Structures of neutral networks and shape space covering. Chem. Mon. 127, 375–389.
Haslinger, C., Stadler, P.F., 1999. RNA structures with pseudo-knots. Bull. Math. Biol. 61, 437–467.
Hofacker, I.L., Fontana, W., Stadler, P.F., Bonhoeffer, L.S., Tacker, M., Schuster, P., 1994. Fast folding and comparison of RNA secondary structures. Monatsh. Chem. 125, 167–188.
Hofacker, I.L., Schuster, P., Stadler, P.F., 1998. Combinatorics of RNA secondary structures. Discrete Appl. Math. 88, 207–237.
Howell, J.A., Smith, T.F., Waterman, M.S., 1980. Computation of generating functions for biological molecules. SIAM J. Appl. Math. 39, 119–133.
Huang, W.D., Reidys, C.M., 2008. Statistics of canonical RNA pseudoknot structures. J. Theor. Biol. doi:10.1016/j.jtbi.2008.04.002.
Huang, W.D., Peng, W.D., Reidys, C.M., 2008. Folding RNA pseudoknot structures, in preparation.
Janson, S., 1990. Poisson approximation for large deviations. Random Struct. Algorithms 1, 221–229.
Jin, E.Y., Reidys, C.M., 2008a. Asymptotic enumeration of RNA structures with pseudoknots. Bull. Math. Biol. 70, 951–970.
Jin, E.Y., Reidys, C.M., 2008a. Central and local limit theorems for RNA structures. J. Theor. Biol. 250, 547–559.
Jin, E.Y., Reidys, C.M., 2008c, Combinatorial design of pseudoknot RNA. Adv. Appl. Math., in press.
Jin, E.Y., Qin, J., Reidys, C.M., 2008. Combinatorics of RNA structures with pseudoknots. Bull. Math. Biol. 70, 45–47
Kimura, M., 1968. Evolutionary rate at the molecular level. Nature 217, 624–626.
Kimura, M., 1983. The Neutral Theory of Molecular Evolution. Cambridge Univ. Press, Cambridge.
Konings, D.A.M., Gutell, R.R., 1995. A comparison of thermodynamic foldings with comparatively derived structures of 16S and 16S-like rRNAs. RNA 1, 559–574.
Loria, A., Pan, T., 1996. Domain structure of the ribozyme from eubacterial ribonuclease P. RNA 2, 551–563.
Lyngso, R., Pedersen, C., 1996. Pseudoknots in RNA secondary structures. In: Physics of Biological Systems: From Molecules to Species. Springer, Berlin.
Science, 2005. Mapping RNA Form and Function. Science 309(2) (5740).
Penner, R.C., Waterman, M.S., 1993. Spaces of RNA secondary structures. Adv. Math. 101, 31–49.
Reidys, C.M., 1997. Random induced subgraphs of generalized n-cubes. Adv. Appl. Math. 19, 360–377.
Reidys, C.M., 2002. Distance in random induced subgraphs of generalized n-cubes. Comb. Probab. Comput. 11, 599–605.
Reidys, C.M., 2008. Large components in random induced subgraphs of n-cubes, Discrete Math., accepted, arXiv:0704.2868.
Reidys, C.M., Stadler, P.F., Schuster, P.K., 1997. Generic properties of combinatory maps and neutral networks of RNA secondary structures. Bull. Math. Biol. 59(2), 339–397.
Rivas, E., Eddy, S., 1999. A dynamic programming algorithm for RNA structure prediction including pseudoknots. J. Mol. Biol. 285, 2053–2068.
Rivas, E., Eddy, S.R., 2000. The language of RNA: a formal grammar that includes pseudoknots. Bioinformatics 16(4), 334–340.
Schultes, E.A., Bartels, P.B., 2000. One sequence, two ribozymes: implications for the emergence of new ribozyme folds. Science 289, 448–452.
Schuster, P., 2002. A testable genotype-phenotype map: Modeling evolution of RNA molecules. In: Laessig, M., Valeriani, A. (Eds.), Biological Evolution and Statistical Physics, pp. 56–83. Springer, New York.
Schuster, P., Fontana, W., Stadler, P.F., Hofacker, I.L., 1994. From sequences to shapes and back: A case study in RNA secondary structures. Proc. Biol. Sci. 255(1344), 279–284.
Searls, D.B., 2002. The language of genes. Nature 420, 211–217.
Uemura, Y., Hasegawa, A., Kobayashi, S., Yokomori, T., 1999. Tree adjoining grammars for RNA structure prediction. Theor. Comput. Sci. 210, 277–303.
Waterman, M.S., 1978. Secondary structure of single - stranded nucleic. acids. Adv. Math. I (suppl.) 1, 167–212.
Waterman, M.S., 1979. Combinatorics of RNA hairpins and cloverleafs. Stud. Appl. Math. 60, 91–96.
Waterman, M.S., Schmitt, W.R., 1994. Linear trees and RNA secondary structure. Discrete Appl. Math. 51, 317–323.
Westhof, E., Jaeger, L., 1992. RNA pseudoknots. Curr. Opin. Struct. Biol. 2, 327–333.
Wong, R., Wyman, M., 1974. The method of Darboux. J. Approx. Theory 10, 159–171.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Reidys, C.M. Local Connectivity of Neutral Networks. Bull. Math. Biol. 71, 265–290 (2009). https://doi.org/10.1007/s11538-008-9356-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11538-008-9356-8