Abstract
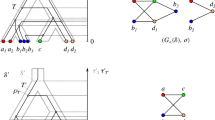
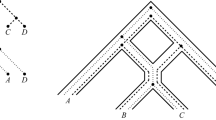
Phylogenetic relationships may be represented by rooted acyclic directed graphs in which each vertex, corresponding to a taxon, possesses a genome. Assume the characters are all binary. A homoplasy occurs if a particular character changes its state more than once in the graph. A vertex is “regular” if it has only one parent and “hybrid” if it has more than one parent. A “regular path” is a directed path such that all vertices after the first are regular. Assume that the network is given and that the genomes are known for all leaves and for the root. Assume that all homoplasies occur only at hybrid vertices and each character has at most one homoplasy. Assume that from each vertex there is a regular path leading to a leaf. In this idealized setting, with other mild assumptions, it is proved that the genome at each vertex is uniquely determined. Hence, for each character the vertex at which a homoplasy occurs in the character is uniquely determined. Without the assumption on regular paths, an example shows that the genomes and homoplasies need not be uniquely determined.
Similar content being viewed by others
References
Bandelt, H.-J., Dress, A., 1986. Reconstructing the shape of a tree from observed dissimilarity data. Adv. Appl. Math. 7, 309–343.
Bandelt, H.-J., Dress, A., 1992. Split decomposition: A new and useful approach to phylogenetic analysis of distance data. Mol. Phylogenet. Evol. 1, 242–252.
Baroni, M., Semple, C., Steel, M., 2004. A framework for representing reticulate evolution. Ann. Comb. 8, 391–408.
Camin, J.H., Sokal, R.R., 1965. A method for deducing branching sequences in phylogeny. Evolution 19, 311–326.
Felsenstein, J., 2006. Inferring Phylogenies. Sinauer Associates, Sunderland, MA.
Dan Gusfield, Gusfield, D., 1991. Efficient algorithms for inferring evolutionary history. Networks 21, 19–28.
Dan Gusfield, Satish Eddhu, and Charles Langley, Gusfield, D., Eddhu, S., Langley, C., 2004a. Optimal, efficient reconstruction of phylogenetic networks with constrained recombination. J. Bioinform. Comput. Biol. 2, 173–213.
Dan Gusfield, Satish Eddhu, and Charles Langley, Gusfield, D., Eddhu, S., Langley, C., 2004b. The fine structure of galls in phylogenetic networks. INFORMS J. Comput. 16, 459–469. Special Issue on Computational Molecular Biology. Bioinformatics.
Hein, J., 1990. Reconstructing evolution of sequences subject to recombination using parsimony. Math. Biosci. 98, 185–200.
Hein, J., 1993. A heuristic method to reconstruct the history of sequences subject to recombination. J. Mol. Evol. 36, 396–405.
Huber, K.T., Moulton, V., 2006. Phylogenetic networks from multi-labelled trees. J. Math. Biol. 52, 613–632.
Huber, K.T., Oxelman, B., Lott, M., Moulton, V., 2006. Reconstructing the evolutionary history of polyploids from multilabeled trees. Mol. Biol. Evol. 23, 1784–1791.
Moret, B.M.E., Nakhleh, L., Warnow, T., Linder, C.R., Tholse, A., Padolina, A., Sun, J., Timme, R., 2004. Phylogenetic networks: Modeling, reconstructibility, and accuracy. IEEE/ACM Trans. Comput. Biol. Bioinform. 1, 13–23.
Nakhleh, L., Warnow, T., Linder, C.R., 2004. Reconstructing reticulate evolution in species—theory and practice. In: Bourne, P.E., Gusfield, D. (Eds.), Proceedings of the Eighth Annual International Conference on Computational Molecular Biology (RECOMB ′04, March 27–31, 2004, San Diego, California), ACM, New York, pp. 337–346.
Semple, C., Steel, M., 2003. Phylogenetics. Oxford University Press, Oxford.
Wang, L., Zhang, K., Zhang, L., 2001. Perfect phylogenetic networks with recombination. J. Comput. Biol. 8, 69–78.
Willson, S.J., 2006a. Unique solvability of certain hybrid networks from their distances. Ann. Combin. 10, 165–178.
Willson, S.J., 2006b. Unique reconstruction of tree-like phylogenetic networks from distances between leaves. Bull. Math. Biol. 68, 919–944.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Willson, S.J. Unique Determination of Some Homoplasies at Hybridization Events. Bull. Math. Biol. 69, 1709–1725 (2007). https://doi.org/10.1007/s11538-006-9187-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11538-006-9187-4