Abstract
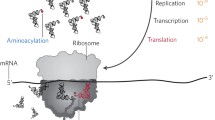
Transfer RNA plays a fundamental role in the protein biosynthesis as an adaptor molecule by functioning as a biological link between the genetic nucleotide sequence in the mRNA and the amino acid sequence in the protein. To perform its role in protein biosynthesis, it has to be accurately recognized by aminoacyl-tRNA synthetases (aaRSs) to generate aminoacyl-tRNAs (aa-tRNAs). The correct pairing between an amino acid with its cognate tRNA is crucial for translational quality control. Production and utilization of mis-charged tRNAs are usually detrimental for all the species, resulting in cellular dysfunctions. Correct aa-tRNAs formation is collectively controlled by aaRSs with distinct mechanisms and/or other trans-factors. However, in very limited instances, mis-charged tRNAs are intermediate for specific pathways or essential components for the translational machinery. Here, from the point of accuracy in tRNA charging, we review our understanding about the mechanism ensuring correct aa-tRNA generation. In addition, some unique mis-charged tRNA species necessary for the organism are also briefly described.
Article PDF
Similar content being viewed by others
References
Ibba M, Söll D. Aminoacyl-tRNA synthesis. Annu Rev Biochem, 2000, 69: 617–650
Giege R. Toward a more complete view of tRNA biology. Nat Struct Mol Biol, 2008, 15: 1007–1014
Francklyn C S, Minajigi A. tRNA as an active chemical scaffold for diverse chemical transformations. FEBS Lett, 2010, 584: 366–375
Mei Y, Stonestrom A, Hou Y M, et al. Apoptotic regulation and tRNA. Protein Cell, 2010, 1: 795–801
Dare K, Ibba M. Roles of tRNA in cell wall biosynthesis. Wiley Interdiscip Rev RNA, 2012, 3: 247–264
Kleiman L, Jones C P, Musier-Forsyth K. Formation of the tRNALys packaging complex in HIV-1. FEBS Lett, 2010, 584: 359–365
Yuan J, Sheppard K, Söll D. Amino acid modifications on tRNA. Acta Biochim Biophys Sin (Shanghai), 2008, 40: 539–553
Sobala A, Hutvagner G. Transfer RNA-derived fragments: origins, processing, and functions. Wiley Interdiscip Rev RNA, 2011, 2: 853–862
Woese C R, Olsen G J, Ibba M, et al. Aminoacyl-tRNA synthetases, the genetic code, and the evolutionary process. Microbiol Mol Biol Rev, 2000, 64: 202–236
Eriani G, Delarue M, Poch O, et al. Partition of tRNA synthetases into two classes based on mutually exclusive sets of sequence motifs. Nature, 1990, 347: 203–206
Cusack S, Berthet-Colominas C, Hartlein M, et al. A second class of synthetase structure revealed by X-ray analysis of Escherichia coli seryl-tRNA synthetase at 2.5 A. Nature, 1990, 347: 249–255
Baldwin A N, Berg P. Transfer ribonucleic acid-induced hydrolysis of valyladenylate bound to isoleucyl ribonucleic acid synthetase. J Biol Chem, 1966, 241: 839–845
Fersht A R. Editing mechanisms in protein synthesis. Rejection of valine by the isoleucyl-tRNA synthetase. Biochemistry, 1977, 16: 1025–1030
Fersht A R, Dingwall C. Evidence for the double-sieve editing mechanism in protein synthesis. Steric exclusion of isoleucine by valyl-tRNA synthetases. Biochemistry, 1979, 18: 2627–2631
Ling J, Reynolds N, Ibba M. Aminoacyl-tRNA synthesis and translational quality control. Annu Rev Microbiol, 2009, 63: 61–78
Ibba M, Söll D. Quality control mechanisms during translation. Science, 1999, 286: 1893–1897
Doring V, Mootz H D, Nangle L A, et al. Enlarging the amino acid set of Escherichia coli by infiltration of the valine coding pathway. Science, 2001, 292: 501–504
Bacher J M, de Crecy-Lagard V, Schimmel P R. Inhibited cell growth and protein functional changes from an editing-defective tRNA synthetase. Proc Natl Acad Sci USA, 2005, 102: 1697–1701
Nangle L A, Motta C M, Schimmel P. Global effects of mistranslation from an editing defect in mammalian cells. Chem Biol, 2006, 13: 1091–1100
Lee J W, Beebe K, Nangle L A, et al. Editing-defective tRNA synthetase causes protein misfolding and neurodegeneration. Nature, 2006, 443: 50–55
Schimmel P. Development of tRNA synthetases and connection to genetic code and disease. Protein Sci, 2008, 17: 1643–1652
Giege R, Sissler M, Florentz C. Universal rules and idiosyncratic features in tRNA identity. Nucleic Acids Res, 1998, 26: 5017–5035
Schmidt E, Schimmel P. Mutational isolation of a sieve for editing in a transfer RNA synthetase. Science, 1994, 264: 265–267
Zhu B, Yao P, Tan M, et al. tRNA-independent pretransfer editing by class I leucyl-tRNA synthetase. J Biol Chem, 2009, 284: 3418–3424
Banerjee R, Chen S, Dare K, et al. tRNAs: cellular barcodes for amino acids. FEBS Lett, 2010, 584: 387–395
Gruic-Sovulj I, Uter N, Bullock T, et al. tRNA-dependent aminoacyl-adenylate hydrolysis by a nonediting class I aminoacyl-tRNA synthetase. J Biol Chem, 2005, 280: 23978–23986
Hati S, Ziervogel B, Sternjohn J, et al. Pre-transfer editing by class II prolyl-tRNA synthetase: role of aminoacylation active site in “selective release” of noncognate amino acids. J Biol Chem, 2006, 281: 27862–27872
Splan K E, Ignatov M E, Musier-Forsyth K. Transfer RNA modulates the editing mechanism used by class II prolyl-tRNA synthetase. J Biol Chem, 2008, 283: 7128–7134
Zhou X L, Ruan Z R, Huang Q, et al. Translational fidelity maintenance preventing Ser mis-incorporation at Thr codon in protein from eukaryote. Nucleic Acids Res, 2013, 41: 302–314
Gruic-Sovulj I, Rokov-Plavec J, Weygand-Durasevic I. Hydrolysis of non-cognate aminoacyl-adenylates by a class II aminoacyl-tRNA synthetase lacking an editing domain. FEBS Lett, 2007, 581: 5110–5114
Tan M, Yan W, Liu R J, et al. A naturally occurring nonapeptide functionally compensates for the CP1 domain of leucyl-tRNA synthetase to modulate aminoacylation activity. Biochem J, 2012, 443: 477–484
Ling J, Peterson K M, Simonovic I, et al. The mechanism of pre-transfer editing in yeast mitochondrial threonyl-tRNA synthetase. J Biol Chem, 2012, 287: 28518–28525
Tan M, Zhu B, Zhou X L, et al. tRNA-dependent pre-transfer editing by prokaryotic leucyl-tRNA synthetase. J Biol Chem, 2010, 285: 3235–3244
Nordin B E, Schimmel P. Transiently misacylated tRNA is a primer for editing of misactivated adenylates by class I aminoacyl-tRNA synthetases. Biochemistry, 2003, 42: 12989–12997
Bishop A C, Beebe K, Schimmel P R. Interstice mutations that block site-to-site translocation of a misactivated amino acid bound to a class I tRNA synthetase. Proc Natl Acad Sci USA, 2003, 100: 490–494
Lincecum T L Jr., Tukalo M, Yaremchuk A, et al. Structural and mechanistic basis of pre- and posttransfer editing by leucyl-tRNA synthetase. Mol Cell, 2003, 11: 951–963
Fukunaga R, Yokoyama S. Structural basis for substrate recognition by the editing domain of isoleucyl-tRNA synthetase. J Mol Biol, 2006, 359: 901–912
Nomanbhoy T K, Hendrickson T L, Schimmel P. Transfer RNA-dependent translocation of misactivated amino acids to prevent errors in protein synthesis. Mol Cell, 1999, 4: 519–528
Silvian L F, Wang J, Steitz T A. Insights into editing from an ile-tRNA synthetase structure with tRNAile and mupirocin. Science, 1999, 285: 1074–1077
Fukai S, Nureki O, Sekine S, et al. Structural basis for double-sieve discrimination of L-valine from L-isoleucine and L-threonine by the complex of tRNA(Val) and valyl-tRNA synthetase. Cell, 2000, 103: 793–803
Dulic M, Cvetesic N, Perona J J, et al. Partitioning of tRNA-dependent editing between pre- and post-transfer pathways in class I aminoacyl-tRNA synthetases. J Biol Chem, 2010, 285: 23799–23809
Dock-Bregeon A C, Rees B, Torres-Larios A, et al. Achieving error-free translation; the mechanism of proofreading of threonyl-tRNA synthetase at atomic resolution. Mol Cell, 2004, 16: 375–386
Zhou X L, Du D H, Tan M, et al. Role of tRNA amino acid-accepting end in aminoacylation and its quality control. Nucleic Acids Res, 2011, 39: 8857–8868
Tan M, Zhu B, Liu R J, et al. Interdomain communication modulates the tRNA-dependent pre-transfer editing of leucyl-tRNA synthetase. Biochem J, 2013, 449: 123–131
Eldred E W, Schimmel P R. Rapid deacylation by isoleucyl transfer ribonucleic acid synthetase of isoleucine-specific transfer ribonucleic acid aminoacylated with valine. J Biol Chem, 1972, 247: 2961–2964
Nordin B E, Schimmel P. Plasticity of recognition of the 3′-end of mischarged tRNA by class I aminoacyl-tRNA synthetases. J Biol Chem, 2002, 277: 20510–20517
Zhao M W, Zhu B, Hao R, et al. Leucyl-tRNA synthetase from the ancestral bacterium Aquifex aeolicus contains relics of synthetase evolution. EMBO J, 2005, 24: 1430–1439
Korencic D, Ahel I, Schelert J, et al. A freestanding proofreading domain is required for protein synthesis quality control in Archaea. Proc Natl Acad Sci USA, 2004, 101: 10260–10265
Wong F C, Beuning P J, Silvers C, et al. An isolated class II aminoacyl-tRNA synthetase insertion domain is functional in amino acid editing. J Biol Chem, 2003, 278: 52857–52864
Beebe K, Ribas De Pouplana L, Schimmel P. Elucidation of tRNA-dependent editing by a class II tRNA synthetase and significance for cell viability. EMBO J, 2003, 22: 668–675
Roy H, Ling J, Irnov M, et al. Post-transfer editing in vitro and in vivo by the beta subunit of phenylalanyl-tRNA synthetase. EMBO J, 2004, 23: 4639–4648
Ling J, So B R, Yadavalli S S, et al. Resampling and editing of mischarged tRNA prior to translation elongation. Mol Cell, 2009, 33: 654–660
Dock-Bregeon A, Sankaranarayanan R, Romby P, et al. Transfer RNA-mediated editing in threonyl-tRNA synthetase. The class II solution to the double discrimination problem. Cell, 2000, 103: 877–884
Nureki O, Vassylyev D G, Tateno M, et al. Enzyme structure with two catalytic sites for double-sieve selection of substrate. Science, 1998, 280: 578–582
Sankaranarayanan R, Dock-Bregeon A C, Romby P, et al. The structure of threonyl-tRNA synthetase-tRNA(Thr) complex enlightens its repressor activity and reveals an essential zinc ion in the active site. Cell, 1999, 97: 371–381
Fukunaga R, Yokoyama S. Aminoacylation complex structures of leucyl-tRNA synthetase and tRNALeu reveal two modes of discriminator-base recognition. Nat Struct Mol Biol, 2005, 12: 915–922
Tukalo M, Yaremchuk A, Fukunaga R, et al. The crystal structure of leucyl-tRNA synthetase complexed with tRNALeu in the post-transfer-editing conformation. Nat Struct Mol Biol, 2005, 12: 923–930
Palencia A, Crepin T, Vu M T, et al. Structural dynamics of the aminoacylation and proofreading functional cycle of bacterial leucyl-tRNA synthetase. Nat Struct Mol Biol, 2012, 19: 677–684
Beebe K, Mock M, Merriman E, et al. Distinct domains of tRNA synthetase recognize the same base pair. Nature, 2008, 451: 90–93
Guo M, Chong Y E, Beebe K, et al. The C-Ala domain brings together editing and aminoacylation functions on one tRNA. Science, 2009, 325: 744–747
Zhou X L, Yao P, Ruan L L, et al. A unique insertion in the CP1 domain of Giardia lamblia leucyl-tRNA synthetase. Biochemistry, 2009, 48: 1340–1347
Fukunaga R, Yokoyama S. Crystal structure of leucyl-tRNA synthetase from the archaeon Pyrococcus horikoshii reveals a novel editing domain orientation. J Mol Biol, 2005, 346: 57–71
Fukunaga R, Fukai S, Ishitani R, et al. Crystal structures of the CP1 domain from Thermus thermophilus isoleucyl-tRNA synthetase and its complex with L-valine. J Biol Chem, 2004, 279: 8396–8402
Fukunaga R, Yokoyama S. Structural basis for non-cognate amino acid discrimination by the valyl-tRNA synthetase editing domain. J Biol Chem, 2005, 280: 29937–29945
Zhu B, Zhao M W, Eriani G, et al. A present-day aminoacyl-tRNA synthetase with ancestral editing properties. RNA, 2007, 13: 15–21
Liu R J, Tan M, Du D H, et al. Peripheral insertion modulates the editing activity of the isolated CP1 domain of leucyl-tRNA synthetase. Biochem J, 2011, 440: 217–227
Lin L, Hale S P, Schimmel P. Aminoacylation error correction. Nature, 1996, 384: 33–34
Chen J F, Guo N N, Li T, et al. CP1 domain in Escherichia coli leucyl-tRNA synthetase is crucial for its editing function. Biochemistry, 2000, 39: 6726–6731
Zhou X L, Wang E D. Two tyrosine residues outside the editing active site in Giardia lamblia leucyl-tRNA synthetase are essential for the post-transfer editing. Biochem Biophys Res Commun, 2009, 386: 510–515
Rock F L, Mao W, Yaremchuk A, et al. An antifungal agent inhibits an aminoacyl-tRNA synthetase by trapping tRNA in the editing site. Science, 2007, 316: 1759–1761
Crepin T, Yaremchuk A, Tukalo M, et al. Structures of two bacterial prolyl-tRNA synthetases with and without a cis-editing domain. Structure, 2006, 14: 1511–1525
Hussain T, Kruparani S P, Pal B, et al. Post-transfer editing mechanism of a D-aminoacyl-tRNA deacylase-like domain in threonyl-tRNA synthetase from Archaea. EMBO J, 2006, 25: 4152–4162
Dwivedi S, Kruparani S P, Sankaranarayanan R. A D-amino acid editing module coupled to the translational apparatus in Archaea. Nat Struct Mol Biol, 2005, 12: 556–557
Hussain T, Kamarthapu V, Kruparani S P, et al. Mechanistic insights into cognate substrate discrimination during proofreading in translation. Proc Natl Acad Sci USA, 2010, 107: 22117–22121
Roy H, Ibba M. Phenylalanyl-tRNA synthetase contains a dispensable RNA-binding domain that contributes to the editing of noncognate aminoacyl-tRNA. Biochemistry, 2006, 45: 9156–9162
Ling J, Yadavalli S S, Ibba M. Phenylalanyl-tRNA synthetase editing defects result in efficient mistranslation of phenylalanine codons as tyrosine. RNA, 2007, 13: 1881–1886
Tardif K D, Horowitz J. Transfer RNA determinants for translational editing by Escherichia coli valyl-tRNA synthetase. Nucleic Acids Res, 2002, 30: 2538–2545
Du X, Wang E D. Tertiary structure base pairs between D- and TpsiC-loops of Escherichia coli tRNA(Leu) play important roles in both aminoacylation and editing. Nucleic Acids Res, 2003, 31: 2865–2872
Yao P, Zhu B, Jaeger S, et al. Recognition of tRNALeu by Aquifex aeolicus leucyl-tRNA synthetase during the aminoacylation and editing steps. Nucleic Acids Res, 2008, 36: 2728–2738
Hale S P, Auld D S, Schmidt E, et al. Discrete determinants in transfer RNA for editing and aminoacylation. Science, 1997, 276: 1250–1252
Huang Q, Yao P, Eriani G, et al. In vivo identification of essential nucleotides in tRNALeu to its functions by using a constructed yeast tRNALeu knockout strain. Nucleic Acids Res, 2012, 40: 10463–10477
Xu M G, Li J, Du X, et al. Groups on the side chain of T252 in Escherichia coli leucyl-tRNA synthetase are important for discrimination of amino acids and cell viability. Biochem Biophys Res Commun, 2004, 318: 11–16
Yao P, Zhou X L, He R, et al. Unique residues crucial for optimal editing in yeast cytoplasmic Leucyl-tRNA synthetase are revealed by using a novel knockout yeast strain. J Biol Chem, 2008, 283: 22591–22600
Zhou X L, Tan M, Wang M, et al. Post-transfer editing by a eukaryotic leucyl-tRNA synthetase resistant to the broad-spectrum drug AN2690. Biochem J, 2010, 430: 325–333
Lue S W, Kelley S O. An aminoacyl-tRNA synthetase with a defunct editing site. Biochemistry, 2005, 44: 3010–3016
Yao Y N, Wang L, Wu X F, et al. Human mitochondrial leucyl-tRNA synthetase with high activity produced from Escherichia coli. Protein Expr Purif, 2003, 30: 112–116
Roy H, Ling J, Alfonzo J, et al. Loss of editing activity during the evolution of mitochondrial phenylalanyl-tRNA synthetase. J Biol Chem, 2005, 280: 38186–38192
Reynolds N M, Ling J, Roy H, et al. Cell-specific differences in the requirements for translation quality control. Proc Natl Acad Sci USA, 2010, 107: 4063–4068
Li L, Boniecki M T, Jaffe J D, et al. Naturally occurring aminoacyl-tRNA synthetases editing-domain mutations that cause mistranslation in Mycoplasma parasites. Proc Natl Acad Sci USA, 2011, 108: 9378–9383
Yadavalli S S, Ibba M. Selection of tRNA charging quality control mechanisms that increase mistranslation of the genetic code. Nucleic Acids Res, 2013, 41: 1104–1112
Gomes A C, Miranda I, Silva R M, et al. A genetic code alteration generates a proteome of high diversity in the human pathogen Candida albicans. Genome Biol, 2007, 8: R206
Reynolds N M, Lazazzera B A, Ibba M. Cellular mechanisms that control mistranslation. Nat Rev Microbiol, 2010, 8: 849–856
Ahel I, Korencic D, Ibba M, et al. Trans-editing of mischarged tRNAs. Proc Natl Acad Sci USA, 2003, 100: 15422–15427
Ruan L L, Zhou X L, Tan M, et al. Human cytoplasmic ProX edits mischarged tRNAPro with amino acid but not tRNA specificity. Biochem J, 2013, 450: 243–252
Ahel I, Stathopoulos C, Ambrogelly A, et al. Cysteine activation is an inherent in vitro property of prolyl-tRNA synthetases. J Biol Chem, 2002, 277: 34743–34748
An S, Musier-Forsyth K. Trans-editing of Cys-tRNAPro by Haemophilus influenzae YbaK protein. J Biol Chem, 2004, 279: 42359–42362
Ruan B, Söll D. The bacterial YbaK protein is a Cys-tRNAPro and Cys-tRNA Cys deacylase. J Biol Chem, 2005, 280: 25887–25891
An S, Musier-Forsyth K. Cys-tRNA(Pro) editing by Haemophilus influenzae YbaK via a novel synthetase.YbaK.tRNA ternary complex. J Biol Chem, 2005, 280: 34465–34472
Nawaz M H, Merriman E, Yang X L, et al. p23H implicated as cis/trans regulator of AlaXp-directed editing for mammalian cell homeostasis. Proc Natl Acad Sci USA, 2011, 108: 2723–2728
Guo M, Chong Y E, Shapiro R, et al. Paradox of mistranslation of serine for alanine caused by AlaRS recognition dilemma. Nature, 2009, 462: 808–812
Yamane T, Miller D L, Hopfield J J. Discrimination between D- and L-tyrosyl transfer ribonucleic acids in peptide chain elongation. Biochemistry, 1981, 20: 7059–7064
Soutourina J, Plateau P, Blanquet S. Metabolism of D-aminoacyl-tRNAs in Escherichia coli and Saccharomyces cerevisiae cells. J Biol Chem, 2000, 275: 32535–32542
Wydau S, van der Rest G, Aubard C, et al. Widespread distribution of cell defense against D-aminoacyl-tRNAs. J Biol Chem, 2009, 284: 14096–14104
Ferri-Fioni M L, Schmitt E, Soutourina J, et al. Structure of crystalline D-Tyr-tRNA(Tyr) deacylase. A representative of a new class of tRNA-dependent hydrolases. J Biol Chem, 2001, 276: 47285–47290
Tumbula D L, Becker H D, Chang W Z, et al. Domain-specific recruitment of amide amino acids for protein synthesis. Nature, 2000, 407: 106–110
Nagao A, Suzuki T, Katoh T, et al. Biogenesis of glutaminyl-mt tRNAGln in human mitochondria. Proc Natl Acad Sci USA, 2009, 106: 16209–16214
Frechin M, Senger B, Braye M, et al. Yeast mitochondrial Gln-tRNA(Gln) is generated by a GatFAB-mediated transamidation pathway involving Arc1p-controlled subcellular sorting of cytosolic GluRS. Genes Dev, 2009, 23: 1119–1130
Pujol C, Bailly M, Kern D, et al. Dual-targeted tRNA-dependent amidotransferase ensures both mitochondrial and chloroplastic Gln-tRNAGln synthesis in plants. Proc Natl Acad Sci USA, 2008, 105: 6481–6485
Lapointe J, Duplain L, Proulx M. A single glutamyl-tRNA synthetase aminoacylates tRNAGlu and tRNAGln in Bacillus subtilis and efficiently misacylates Escherichia coli tRNAGln1 in vitro. J Bacteriol, 1986, 165: 88–93
Schulze J O, Masoumi A, Nickel D, et al. Crystal structure of a non-discriminating glutamyl-tRNA synthetase. J Mol Biol, 2006, 361: 888–897
Guo L T, Helgadottir S, Söll D, et al. Rational design and directed evolution of a bacterial-type glutaminyl-tRNA synthetase precursor. Nucleic Acids Res, 2012, 40: 7967–7974
Nakamura A, Yao M, Chimnaronk S, et al. Ammonia channel couples glutaminase with transamidase reactions in GatCAB. Science, 2006, 312: 1954–1958
Oshikane H, Sheppard K, Fukai S, et al. Structural basis of RNA-dependent recruitment of glutamine to the genetic code. Science, 2006, 312: 1950–1954
Ito T, Yokoyama S. Two enzymes bound to one transfer RNA assume alternative conformations for consecutive reactions. Nature, 2010, 467: 612–616
Min B, Pelaschier J T, Graham D E, et al. Transfer RNA-dependent amino acid biosynthesis: an essential route to asparagine formation. Proc Natl Acad Sci USA, 2002, 99: 2678–2683
Charron C, Roy H, Blaise M, et al. Non-discriminating and discriminating aspartyl-tRNA synthetases differ in the anticodon-binding domain. EMBO J, 2003, 22: 1632–1643
Min B, Kitabatake M, Polycarpo C, et al. Protein synthesis in Escherichia coli with mischarged tRNA. J Bacteriol, 2003, 185: 3524–3526
Namgoong S, Sheppard K, Sherrer R L, et al. Co-evolution of the archaeal tRNA-dependent amidotransferase GatCAB with tRNA(Asn). FEBS Lett, 2007, 581: 309–314
Sheppard K, Söll D. On the evolution of the tRNA-dependent amidotransferases, GatCAB and GatDE. J Mol Biol, 2008, 377: 831–844
Bailly M, Blaise M, Lorber B, et al. The transamidosome: a dynamic ribonucleoprotein particle dedicated to prokaryotic tRNA-dependent asparagine biosynthesis. Mol Cell, 2007, 28: 228–239
Blaise M, Bailly M, Frechin M, et al. Crystal structure of a transfer-ribonucleoprotein particle that promotes asparagine formation. EMBO J, 2010, 29: 3118–3129
Commans S, Bock A. Selenocysteine inserting tRNAs: an overview. FEMS Microbiol Rev, 1999, 23: 335–351
Bosl M R, Takaku K, Oshima M, et al. Early embryonic lethality caused by targeted disruption of the mouse selenocysteine tRNA gene (Trsp). Proc Natl Acad Sci USA, 1997, 94: 5531–5534
Kryukov G V, Castellano S, Novoselov S V, et al. Characterization of mammalian selenoproteomes. Science, 2003, 300: 1439–1443
Rayman M P. The importance of selenium to human health. Lancet, 2000, 356: 233–241
Ambrogelly A, Palioura S, Söll D. Natural expansion of the genetic code. Nat Chem Biol, 2007, 3: 29–35
Forchhammer K, Leinfelder W, Boesmiller K, et al. Selenocysteine synthase from Escherichia coli. Nucleotide sequence of the gene (selA) and purification of the protein. J Biol Chem, 1991, 266: 6318–6323
Carlson B A, Xu X M, Kryukov G V, et al. Identification and characterization of phosphoseryl-tRNA[Ser]Sec kinase. Proc Natl Acad Sci USA, 2004, 101: 12848–12853
Palioura S, Sherrer R L, Steitz T A, et al. The human SepSecS-tRNASec complex reveals the mechanism of selenocysteine formation. Science, 2009, 325: 321–325
Zinoni F, Heider J, Bock A. Features of the formate dehydrogenase mRNA necessary for decoding of the UGA codon as selenocysteine. Proc Natl Acad Sci USA, 1990, 87: 4660–4664
Sauerwald A, Zhu W, Major T A, et al. RNA-dependent cysteine biosynthesis in Archaea. Science, 2005, 307: 1969–1972
Kamtekar S, Hohn M J, Park H S, et al. Toward understanding phosphoseryl-tRNACys formation: the crystal structure of Methanococcus maripaludis phosphoseryl-tRNA synthetase. Proc Natl Acad Sci USA, 2007, 104: 2620–2625
Liu Y, Dos Santos P C, Zhu X, et al. Catalytic mechanism of Sep-tRNA:Cys-tRNA synthase: sulfur transfer is mediated by disulfide and persulfide. J Biol Chem, 2012, 287: 5426–5433
O’Donoghue P, Sethi A, Woese C R, et al. The evolutionary history of Cys-tRNACys formation. Proc Natl Acad Sci USA, 2005, 102: 19003–19008
Moura G R, Paredes J A, Santos M A. Development of the genetic code: insights from a fungal codon reassignment. FEBS Lett, 2010, 584: 334–341
Santos M A, Perreau V M, Tuite M F. Transfer RNA structural change is a key element in the reassignment of the CUG codon in Candida albicans. EMBO J, 1996, 15: 5060–5068
Suzuki T, Ueda T, Watanabe K. The ‘polysemous’ codon—a codon with multiple amino acid assignment caused by dual specificity of tRNA identity. EMBO J, 1997, 16: 1122–1134
Zhou X L, Fang Z P, Ruan Z R, et al. Aminoacylation and translational quality control strategy employed by leucyl-tRNA synthetase from a human pathogen with genetic code ambiguity. Nucleic Acids Res, 2013, doi: 10.1093/nar/gkt741
Netzer N, Goodenbour J M, David A, et al. Innate immune and chemically triggered oxidative stress modifies translational fidelity. Nature, 2009, 462: 522–526
Wiltrout E, Goodenbour J M, Frechin M, et al. Misacylation of tRNA with methionine in Saccharomyces cerevisiae. Nucleic Acids Res, 2012, 40: 10494–10506
Jones T E, Alexander R W, Pan T. Misacylation of specific nonmethionyl tRNAs by a bacterial methionyl-tRNA synthetase. Proc Natl Acad Sci USA, 2011, 108: 6933–6938
Author information
Authors and Affiliations
Corresponding author
Additional information
This article is published with open access at Springerlink.com
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 2.0 International License (https://creativecommons.org/licenses/by/2.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
About this article
Cite this article
XiaoLong, Z., EnDuo, W. Transfer RNA: A dancer between charging and mis-charging for protein biosynthesis. Sci. China Life Sci. 56, 921–932 (2013). https://doi.org/10.1007/s11427-013-4542-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11427-013-4542-9