Abstract
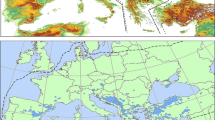
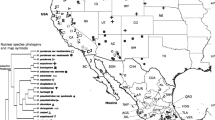
The European Black Pine (Pinus nigra Arn.) has a long and complex history. Genetic distance and frequency analyses identified three differentiated genetic groups, which corresponded to three wide geographical areas: Westerns Mediterranean, Balkan Peninsula and Asia Minor. These groups shared common ancestors (14.75 and 10.72 Ma). The most recent splits occurred after the Messinian Salinity Crisis (4.37 Ma) and the Early–Middle Pleistocene Transitions (0.93 Ma). The posterior ancestral population size (Na) is 260,000–265,000 individuals. Each pool is further fragmented, with evidence of a phylogeographic structure (N st > G st ) typically observed in some natural populations from the Western Mediterranean region and the Balkan Peninsula. The laboratory analysis was performed by fragment analysis—i.e. electrophoretic sizing of polymerase chain reaction fragments, combined with the sequencing analysis of 33 % of all individuals as a control. Intense sampling of chloroplast DNA polymorphisms (3154 individuals and 13 markers: SNPs and SSRs) over the full area of the species’ natural distribution indicated moderate among-population variability (G st(nc) ≤ 0.177) in various parts of its range. These results indicate that the natural populations have long migration histories that differ from one another and that they have been strongly phylogeographically affected by complex patterns of isolation, speciation and fragmentation. Long and varying climatic fluctuations in the region of the principal genetic group have been the probable cause of different forest community associations with different successional patterns resulting in interglacial refugia vs. macro long-term refugia.




Similar content being viewed by others
References
Adams RI, Brown KM, Hamilton MB (2004) The impact of microsatellite electromorph size homoplasy on multilocus population structure estimates in a tropical tree (Corythophora alta) and an anadromous fish (Morone saxatilis). Mol Ecol 13(9):2579–2588
Balaresque P, Bowden GR, Adams SM, et al. (2010) A predominantly Neolithic origin for European paternal lineages. PLoS Biol 8(1):e1000285. doi:10.1371/journal.pbio.1000285
Belkhir K (2000) Genetix ver 4.01: a software for population genetics data analysis. Loboratoire genome et populations. Universite de Montpellier, II, France
Bojović S (1995) Biodiversité du pin noir (Pinus nigra Arn.) en région méditerranéenne. Thèse doctorale. Université d’Aix-Marseille III, France, pp. 1–114
Bonavita S, Vendramin GG, Bernardini V, et al. (2015) The first SSR-based assessment of genetic variation and structure among Pinus laricio Poiret populations within their native area. Plant Biosyst Int J Dealing Asp Plant Biol. doi:10.1080/11263504.2015.1027316
Bucci G, Gonzalez-Martinez SC, Le Provost G, et al. (2007) Range-wide phylogeography and gene zones in Pinus pinaster Ait. revealed by chloroplast microsatellite markers. Mol Ecol 16(10):2137–2153. doi:10.1111/j.1365-294X.2007.03275.x
Chen C, Durand E, Forbes F, et al. (2007) Bayesian clustering algorithms ascertaining spatial population structure: a new computer program and a comparison study. Mol Ecol Notes 7:747–756
Cinget B, Gérardi S, Beaulieu J, et al. (2015) Less pollen-mediated gene flow for more signatures of glacial lineages: congruent evidence from Balsam Fir cpDNA and mtDNA for multiple refugia in eastern and Central North America. PLoS One 10(4):e0122815. doi:10.1371/journal.pone.0122815
Coart E, Van Labeke S, De Loose M, et al. (2006) Chloroplast diversity in the genus Malus: new insights into the relationship between the European wild apple (Malus sylvestris (L.) Mill.) and the domesticated apple (Malus domestica Borkh.). Mol Ecol 15:2171–2182. doi:10.1111/j.1365-294X.2006.02924.x
Critchfield WB, Little EL (1966) Geographic distribution of the pines of the world. U.S.D.A. Forest Service Miscellaneous Publication 991
Delplancke M, Alvarez N, Espíndola A, et al. (2012) Gene flow among wild and domesticated almond species: insights from chloroplast and nuclear markers. Evol Appl 5:317–329. doi:10.1111/j.1752-4571.2011.00223.x
Dobrinov I, Doykov G, Gagov V (1982) Forest genetic pool in Bulgaria. Zemizdat, Sofia [in Bulgarian]--
Doyle JJ, Morgante M, Tingey SV, et al. (1998) Size homoplasy in chloroplast microsatellites of wild perennial relatives of soybean (Glycine subgenus glycine). Mol Biol Evol 15(2):215–218
Duminil J, Heuertz M, Doucet J-L, et al. (2010) CpDNA-based species identification and phylogeography: application to African tropical tree species. Mol Ecol 19:5469–5483. doi:10.1111/j.1365-294X.2010.04917.x
Dzialuk A, Muchewicz E, Boratynski A, et al. (2009) Genetic variation of Pinus uncinata (Pinaceae) in the Pyrenees determined with cpSSR markers. Plant Syst Evol 277:197–205. doi:10.1007/s00606-008-0123-y
Eckenwalder JE (2009) Conifers of the world—the complete reference. Timber Press, Oregon, 720p
Emerson BC, Hewitt GM (2005) Phylogeography. Curr Biol 15(10):R367–R371
Estoup A, Jarne P, Cornuet JM (2002) Homoplasy and mutation model at microsatellite loci and their consequences for population genetics analysis. Mol Ecol 11:1591–1604
Evanno G, Regnau S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14:2611–2620. doi:10.1111/j.1365-294X.2005.02553.x
Excoffier L, Schneider S, Roessli D (2002) Arlequin ver 2.001: a software for population genetics data analysis. Department of Anthropology and Ecology, University of Geneva, Switzerland
Falush D, Stephens M, Pritchard JK (2003a) Inference of population structure using multilocus genotype data: linked loci and correlated allele frequencies. Genetics 164:1567–1587
Falush D, Wirth T, Linz B, et al. (2003b) Traces of human migrations in Helicobacter pylori populations. Science 299:1582–1585
Farjon A (2008) A natural history of conifer. Timber Press, Oregon, 304p
Fineschi S (1984) Determination of the origin of an isolated group of trees of Pinus nigra through enzyme gene markers. Silvae Genet 33:169–172
François O, Ancelet S, Guillot G (2006) Bayesian clustering using hidden Markov random fields in spatial population genetics. Genetics 174:805–816
Fukarek P (1958a) Prilog poznavanju crnog bora (Pinus nigra ARN. S. lat.) Beitrag zur Kenntnis der systematischen Stellung, Gliederung und der rezenten Verbreitung der Schwarzkiefer. Rad Poljopr Sumersk Fak Univ Sarajevu 3:3–92 [in German]
Fukarek P (1958b) Standortrassen der Schwarzfhre (Pinus nigra ARN. S. lat.). Centralbl Gesamten Forstwes 75:203–207 [in German]
García-Amorena I, Rubiales JM, Amat EM, et al. (2011) New macrofossil evidence of Pinus nigra Arnold on the northern Iberian Meseta during the Holocene. Rev Palaeobot Palynol 63(3–4):281–288. doi:10.1016/j.revpalbo.2010.10.010
Gealey WK (1989) Plate tectonic evolution of the Mediterranean ’ Middle East region. In: Scotese CR, Sager WW (eds) Mesozoic and Cenozoic plate reconstructions. Elsevier, Amsterdam, pp. 285–306
Godbout J, Fazekas A, Newton CH, et al. (2008) Glacial vicariance in the Pacific northwest: evidence from a lodgepole pine mitochondrial DNA minisatellite for multiple genetically distinct and widely separated refugia. Mol Ecol 17(10):2449–2462. doi:10.1111/j.1365-294X.2008.03761.x
Gomez A, Aguiriano E, Alía R, et al. (2002) Análisis de los recursos genéticos de Pinus pinea L. en España mediante microsatélites del cloroplasto. Invest Agr Sist Recur For 11(1):145–154 [in Spanisch]
Gomez A, Vendramin GG, González-Martínez SC, et al. (2005) Genetic diversity and differentiation of two Mediterranean pines (Pinus halepensis Mill. and Pinus pinaster Ait.) along a latitudinal cline using chloroplast microsatellite markers. Divers Distrib 11:257–263
Grant V (1980) Gene flow and the homogeneity of species populations. Biol Zbl 99:157–169
Grivet D, Petit RJ (2003) Chloroplast DNA phylogeography of the hornbeam in Europe: evidence for a bottleneck at the outset of postglacial colonization. Conserv Genet 4:47–56
Hale ML, Borland AM, Gustafsson MHG, et al. (2004) Causes of size homoplasy among chloroplast microsatellites in closely related Clusia species. J Mol Evol 58:182–190
Hardy OJ, Vekemans X (2002) spagedi: a versatile computer program to analyse spatial genetic structure at the individual or population levels. Mol Ecol Notes 2:618–620. doi:10.1046/j.1471-8278.2002.00305.x
He T, Pausas JG, Belcher CM, et al. (2012) Fire-adapted traits of Pinus arose in the fiery Cretaceous. New Phytol 194:751–759
Heuertz M, Carnevale S, Fineschi S, et al. (2006) Chloroplast DNA phylogeography of three European ashes, Fraxinus sp. (Oleaceae): roles of hybridization and contrasting mating systems. Mol Ecol 15:2131–2140
Heuertz M, Teufel J, González-Martínez SC, et al. (2010) Geography determines genetic relationships between species of mountain pine (Pinus mugo complex) in Western Europe. J Biogeogr 37(3):541–556. doi:10.1111/j.1365-2699.2009.02223.x
Hodel RG, Gonzales E (2013) Phylogeography of sea oats (Uniola paniculata), a dune-building coastal grass in Southeastern North America. J Hered, June, doi:10.1093/jhered/est035
Hohn M, Gugerli F, Abran P, et al. (2009) Variation in the chloroplast DNA of Swiss stone pine (Pinus cembra L.) reflects contrasting post-glacial history of populations from the Carpathians and the Alps. J Biogeogr. doi:10.1111/j.1365-2699.2009.02122.x
Hudson RR (1990) Gene genealogies and the coalescent process. Oxf Surv Evol Biol 7:1–44
Iversen J (1973) The development of Denmark’s nature since the last glacial. Dan Geol Undersogelse 7C:1–126
Jaramillo-Correa JP, Aguirre-Planter E, Khasa DP, et al. (2008) Ancestry and divergence of subtropical montane forest isolates: molecular biogeography of the genus Abies (Pinaceae) in southern Mexico and Guatemala. Mol Ecol 17(10):2476–2490. doi:10.1111/j.1365-294X.2008.03762.x
Joos F, Colin PI (2004) A paleo-perspective on changes in atmospheric CO2 and climate. The global carbon cycle: integrating humans, climate, and the natural world. Scope 62:165–186
Kalinowski ST (2005) HP-Rare: a computer program for performing rarefaction on measures of allelic diversity. Mol Ecol Notes 5:187–189
Kalinowski ST (2009) How well do evolutionary trees describe genetic relationships between populations? Heredity 102:506–513
Kayser M, Brauer S, Cordaux R, et al. (2006) Melanesian and Asian origins of Polynesians: mtDNA and Y chromosome gradients across the Pacific. Mol Biol Evol 23(11):2234–2244
Kingman JFC (1982) The coalescent. Stoch Process Appl 13(3):235–248. doi:10.1016/0304-4149(82)90011-4
Lande R, Engen S, Saether BE (1999) Spatial scale of population synchrony: environmental correlation vs. dispersal and density regulation. Am Nat 154:271–281
Lande R, Engen S, Saether BE (2003) Stochastic population dynamics in ecology and conservation. Oxford University Press, Oxford
Lehman N (1998) Conservation biology: genes are not enough. Curr Biol 8:R722–R724
Liu K, Muse SV (2005) PowerMarker: an integrated analysis environment for genetic marker analysis. Bioinformatics 21:2128–2129
Lumaret R, López de Heredia U, Soto A (2009) Origin and genetic variability of cork-oak. In: Aronson J, Pereira JS, Pausas J (eds) Cork oak woodlands on the edge: ecology, adaptive management and restoration. Island Press, Washington, DC, pp. 25–32
Lumaret R, Tryphon-Dionnet M, Michaud H, et al. (2005) Phylogeographical variation of chloroplast DNA in cork oak (Quercus suber). Ann Bot 96:853–861. doi:10.1093/aob/mci237
Manni F, Guerard E, Heyer E (2004) Geographic patterns of (genetic, morphologic, linguistic) variation: how barriers can be detected by using Monmonier’s algorithm. Hum Biol 76:173–190
Maslin MA, Ridgwell AJ (2005) Mid-Pleistocene revolution and the ‘eccentricity myth’. In: Head, M.J., Gibbard, P.L. (Ed), “Early–Middle Pleistocene transitions: the land–ocean evidence”, Geol Soc Lond, Spec Publ, 247: 19–34.
Miller P (1768) The gardeners dictionary. Ed. 8. London
Mirov NT (1967) The genus Pinus. The Ronald Press, New York, 610 p
Navascués M, Emerson BC (2005) Chloroplast microsatellites: measures of genetic diversity and the effect of homoplasy. Mol Ecol 14:1333–1341. doi:10.1111/j.1365-294X.2005.02504.x
Naydenov KD, Mladenov I, Alexandrov A, et al. (2015) Patterns of genetic diversity resulting from bottlenecks in European Black Pine, with implications on local genetic conservation and management practices in Bulgaria. Eur J For Res 134(4):669–681. doi:10.1007/s10342-015-0881-3
Naydenov KD, Naydenov MK, Tremblay F, et al. (2011) Patterns of genetic diversity that result from bottlenecks in Scots Pine and the implications for local genetic conservation and management practices in Bulgaria. New For 42(2):179
Naydenov KD, Senneville S, Beaulieu J, et al. (2007) Glacial vicariance in Eurasia: mitochondrial DNA evidence from Scots Pine for a complex heritage involving genetically distinct refugia at mid-northern latitudes and in Asia Minor. BMC Evol Biol 7:233. doi:10.1186/1471-2148-7-233
Naydenov KD, Tremblay F, Fenton N (2005) Chloroplast microsatellites analysis revealed a high level of differentiation in Jack pine (Pinus banksiana Lamb.) population in Quebec. Belg J Bot 138(2):181–191
Naydenov KD, Tremblay F, Fenton NJ, et al. (2006) Structure of Pinus nigra Arn. populations in Bulgaria revealed by chloroplast microsatellites and terpenes analysis: provenance tests. Biochem Syst Ecol 34:562–574
Naydenov KD, Tremblay F, Ganchev P (2003) Karyotype diversity in of European Black Pine (Pinus nigra Arn.) from Bulgarian provenances. Phyton 43(1):9–28
Nei M, Chesser RK (1983) Estimation of fixation indices and gene diversities. Ann Hum Genet 47:253–259
Palamarev E (1989) Paleobotanical evidences of the Tertiary history and origin of the Mediterranean sclerophyll dendroflora. Woody plants—evolution and distribution since the Tertiary. Plant Syst Evol 162(1/4):93–107
Perdereau AC, Kelleher CT, Douglas GC, Hodkinson TR (2014) High levels of gene flow and genetic diversity in Irish populations of Salix caprea L. inferred from chloroplast and nuclear SSR markers. BMC Plant Biol 14:202. doi:10.1186/s12870-014-0202-x
Petit RJ, Aguinagalde I, de Beaulieu J-L, et al. (2003) Glacial refugia: hotspots but not melting pots of genetic diversity. Science 300(5625):1563. doi:10.1126/science.1083264
Petit RJ, Brewer S, Bordaes S, et al. (2002a) Identification of refugia and post-glacial colonisation routes of European white oaks based on chloroplast DNA and fossil pollen evidence. For Ecol Manag 156:49–74
Petit RJ, Csaikl UM, Bordaes S, et al. (2002b) Chloroplast DNA variation in European white oaks: phylogeography and patterns of diversity based on data from over 2,600 populations. For Ecol Manag 156:5–26
Petit RJ, Excoffier L (2009) Gene flow and species delimitation. Trends Ecol Evol 24(7):386–393
Petit RJ, Hampe A, Cheddadi R (2005) Climate changes and tree phylogeography in the Mediterranean. Taxon 54(4):877–885
Pianka ER (1978) Evolutionary ecology. Harper and Row, New York
Pico FX, Mendez-Vigo B, Martinez-Zapater JM, et al. (2008) Natural genetic variation of Arabidopsis thaliana is geographically structured in the Iberian Peninsula. Genetics 180(2):1009–1021. doi:10.1534/genetics.108.089581
Pons O, Petit RJ (1996) Measuring and testing genetic differentiation with ordered versus unordered alleles. Genetics 144:1237–1245
Powell W, Morgante M, Andre C, et al. (1995b) Hypervariable microsatellites provide a general source of polymorphic DNA markers for the chloroplast genome. Curr Biol 5:1023–1029
Powell W, Morgante M, McDevitt R, et al. (1995a) Polymorphic simple sequence repeat regions in the chloroplast genome: applications to the population genetics of pines. Proc Natl Acad Sci U S A 92:7759–7763
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155(2):945–959
Pritchard JK, Wen W (2003) Documentation for structure software Version 2 [online]. Available from http://pritch.bsd.uchicago.edu/software/readme_2_1/readme.html
Provan J, Bennett KD (2008) Phylogeographic insights into cryptic glacial refugia. Trends Ecol Evol 23(10):564–571. doi:10.1016/j.tree.2008.06.010
Provan J, Soranzo N, Wilson NJ, et al. (1999) A low mutation rate for chloroplast microsatellites. Genetics 153:943–947
Raffi ZA, Dodd RS (2007) Chloroplast DNA supports a hypothesis of glacial refugia over postglacial recolonization in disjunct populations of black pine (Pinus nigra) in Western Europe. Mol Ecol 16(4):723–736
Raufaste N, Bonhomme F (2000) Properties of bias and variance of two multiallelic estimators of Fst. Theor Popul Biol 57:285–296
Robertson A, Hill WG (1984) Deviations from Hardy-Weinberg proportions: sampling variances and use in estimation of inbreeding coefficients. Genetics 107:703–718
Rogers JS (1972) Measures of similarity and genetic distance. In studies in genetics VII. University of Texas Publication 7213, Austin, Texas, p 145−153
Roiron P, Chabal L, Sueiral I, et al. (2013) Palaeobiogeography of Pinus nigra Arn. subsp. salzmannii (Dunal) Franco in the north-western Mediterranean Basin: a review based on macroremains. Rev Palaeobot Palynol 194:1–11. doi:10.1016/j.revpalbo.2013.03.002
Scotti I, Paglia G, Magni F, et al. (2006) Population genetics of Norway spruce (Picea abies Karst.) at regional scale: sensitivity of different microsatellite motif classes in detecting differentiation. Ann For Sci 63:485–491. doi:10.1051/forest:2006029
Semerikov VL, Semerikova SA, Dymshakova OS, et al. (2014) Microsatellite loci polymorphism of chloroplast DNA of Scots Pine (Pinus sylvestris L.) in Asia and Eastern Europe. Russ J Genet 50(6):577–585. doi:10.1134/S1022795414040127
Soto A, Robledo-Arnuncio JJ, Gonzalez-Martinez SC, et al. (2010) Climatic niche and neutral genetic diversity of the six Iberian pine species: a retrospective and prospective view. Mol Ecol 19:1396–1409. doi:10.1111/j.1365-294X.2010.04571.x
Spellman GM, Klicka J (2006) Testing hypotheses of Pleistocene population history using coalescent simulations: phylogeography of the pygmy nuthatch (Sitta pygmaea). Proc R Soc B 273:3057–3063
Stampfli G, Borel G, Cavazza W, et al (2002) The paleotectonic atlas of the periTethyan domain: European Geophysical Society, CD-ROM
Stefanov B (1941/1942) Geographical distribution of coniferous species and their form in nature. Godichnik na Sofiiskia Darjaven Universitet, Sofia, (Bulgaria), XIX and XX, 1–88 [in Bulgarian]
Stefanov B (1943) The phyto-geographical elements of Bulgaria. Thesis of Bulgarian Academy of Sciences, Faculty of Nature and Mathematics, Sofia, (Bulgaria), vol. XXXIX, 19:1–121 [in Bulgarian]
Stewart JR (2003) Comment on ‘Buffered tree population changes in a Quaternary refugium: evolutionary implications’. Science 299:825a. doi:10.1126/science.1079388
Stewart JR, Dalen L (2008) Is the glacial refugium concept relevant for northern species? A comment on Pruett and Winker 2005. Clim Chang 86:1–2. doi:10.1007/s10584-007-9366-9
Stewart JR, Lister AM, Barnes I et al (2010) Refugia revisited: individualistic responses of species in space and time. Proc R Soc B (277): 661–671; DOI:10.1098/rspb.2009.1272.
Svenning JC, Skov F (2004) Limited filling of the potential range in European tree species. Ecol Lett 1:565–573
Taberlet P, Fumagalli L, Wust-Saucy AG, et al. (1998) Comparative phylogeography and postglacial colonization routes in Europe. Mol Ecol 7:453–464
Terrab A, Paun O, Talavera S, et al. (2006) Genetic diversity and population structure in natural populations of Moroccan Atlas cedar (Cedrus atlantica; Pinaceae) determined with cpSSR markers. Am J Bot 93:1274–1280. doi:10.3732/ajb.93.9.1274
Vendramin G, Lelli L, Rossi P, et al. (1996) A set of primer for the amplification of 20 chloroplast microsatellites in Pinaceae. Mol Ecol 5:585–598
Vidakovic M (1991) Conifers: morphology and variation. Graficki Zavod Hrvatske, Croatia
Vinceti B, Loo J, Gaisberger H, et al. (2013) Conservation priorities for Prunus africana defined with the aid of spatial analysis of genetic data and climatic variables. PLoS One 8(3):e59987. doi:10.1371/journal.pone.0059987
Walter R, Epperson BK (2005) Geographic pattern of genetic diversity in Pinus resinosa: contact zone between descendants of glacial refugia. Am J Bot 92(1):92–100
Wang B, Mahani MK, Ng WL, et al. (2014) Extremely low nucleotide polymorphism in Pinus krempfii Lecomte, a unique flat needle pine endemic to Vietnam. Ecol Evol 4(11):2228–2238. doi:10.1002/ece3.1091
Wang Y, Jiang H, Peng S, et al. (2011) Genetic structure in fragmented populations of Hippophae rhamnoides ssp. sinensis in China investigated by ISSR and cpSSR markers. Plant Syst Evol 295(1/4):97–107 http://www.jstor.org/stable/43558222
Weir BS, Cockerham CC (1984) Estimating F-statistics for the analysis of population structure. Evolution 38:1358–1370
Wilson IJ, Weale ME, Balding DJ (2003) Inferences from DNA data: population histories, evolutionary processes and forensic match probabilities. J R Stat Soc A 166:155–201
Wright S (1979) Evolution and the genetics of populations, vol. 4. Variability within and among natural populations. Chicago, 580 pp
Acknowledgments
We would like to thank Irena M. Naydenova and T&T for their technical assistance; the two anonymous organisations for their financial support; and Ph.D. Z. Kaya (Turkey), Ph.D. M. Kostadinovski (Macedonia), M. Topac (Turkey) and Ph.D. C. Varelides (Greece) who all made direct (and indirect) logistical help and supplied some samples. We would also like to thank the Ministers of Forestry, Education and Science of all the countries with participant persons for providing the funding for sample collection and fruitful collaboration. We also wish to thank the Nature Publishing Group Language Editing-NPGLE (www.languageediting.nature.com) and Miss Honor Hedges (CFB-Kingston, ON, Canada) for the careful English revision of this manuscript.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by Z. Kaya
This article is dedicated to the memory of Prof. Dr. Dimitar Velkov from Forest Research Institute, Bulgarian Academy of Science (1921–2001).
Rights and permissions
About this article
Cite this article
Naydenov, K.D., Naydenov, M.K., Alexandrov, A. et al. Ancient split of major genetic lineages of European Black Pine: evidence from chloroplast DNA . Tree Genetics & Genomes 12, 68 (2016). https://doi.org/10.1007/s11295-016-1022-y
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11295-016-1022-y