Abstract
Rift Valley fever phlebovirus (RVFV) is a mosquito-transmitted pathogen endemic to sub-Saharan Africa and the Arabian Peninsula. RVFV is a threat to both animal and human health and has costly economic consequences mainly related to livestock production and trade. Competent hosts and vectors for RVFV are widespread, existing outside of endemic countries including the USA. Thus, the possibility of RVFV spreading to the USA or other countries worldwide is of significant concern. RVFV (genus Phlebovirus) is comprised of an enveloped virion containing a three-segmented, negative-stranded RNA genome that is able to undergo genetic reassortment. Reassortment has the potential to produce viruses that are more pathogenic, easily transmissible, and that have wider vector or host range. This is especially concerning because of the wide use of live attenuated vaccine strains throughout endemic countries. This review focuses on the molecular aspects of RVFV, genetic diversity of RVFV strains, and RVFV reassortment.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Rift Valley fever phlebovirus (RVFV) is the etiological agent of a mosquito-transmitted disease endemic in sub-Saharan Africa which has spread to the Arabian Peninsula. Outbreaks of the disease result in high morbidity and mortality among young ruminants including sheep, goats, cattle, and pseudoruminants such as camels, and frequent abortions in pregnant animals [1]. Thus, RVFV is responsible for great economic losses to livestock production and rural farming communities. Moreover, the virus can cause disease in humans ranging from mild flu-like symptoms to severe hepatitis, blindness, encephalitis, and in some cases death [1]. Inactivated and live attenuated vaccines have historically been used for veterinary use during outbreaks in endemic countries [2]. Live attenuated vaccines are controversial because of the concern of reversion back to virulence, either by mutation or by genomic segment reassortment with the circulating wild-type strains. Other RVFV vaccines have been developed using various platforms such as genetically modified live, recombinant, vectored and replicon systems in order to make safer, more effective, and DIVA (differentiating infected from vaccinated animals) compatible vaccines [3]. However, there is still no fully licensed veterinary or human vaccine for use in non-endemic countries.
Transmission of RVFV can occur through contact with infected blood, milk, or tissue, or by RVFV-infected mosquito bites. RVFV-competent mosquito vectors fall under 73 species of mosquitoes in the 8 genera of the family Culicidae [4, 5]. Enzootics/epidemics typically occur following periods of heavy rainfall which creates favorable breeding conditions for mosquito vectors. During inter-epidemic periods, RVFV circulates in nature via horizontal transmission between mosquito vectors and livestock, or wildlife reservoirs such as indigenous wild ruminants. Trans-ovarial or vertical transmission is also widely reported [5]. Field testing for RVFV by immunological assays in endemic areas and in vivo studies has shown that RVFV can infect a wide range of animals [6]. Although no definitive wildlife reservoir has been clearly identified, there is evidence suggesting RVFV may be maintained in buffalo [7, 8]. Competent vectors and host species exist outside of endemic geographic regions including the USA [9,10,11]. Interestingly, cell lines from multiple tissues derived from animals native to North America such as white tailed deer, coyote, and swine were found to support RVFV replication [12]. Thus, RVFV is not only a concern for endemic countries, but presents a risk worldwide.
Molecular properties of RVFV
Classification
RVFV belongs to the genus Phlebovirus, along with Punta Toro, Uukuniemi, sandfly fever-like phleboviruses and other species. The genus Phlebovirus has been historically classified under the family Bunyaviridae along with Orthobunyavirus, Hantavirus, Nairovirus, and Tospovirus. In 2016, the International Committee on Taxonomy of Viruses created a new order, Bunyavirales, which contains nine new families and reorganized the previous genera, thus replacing Bunyaviridae [13]. Phlebovirus is currently classified under the family Phenuiviridae (Fig. 1).
Virion organization
Virion structure
Using cryo-electron tomography, RVF viral particles were determined to be spherical with an average particle diameter of 103 ± 3 nm [14], consistent with previous studies which identified pleomorphic particles with varying diameters ranging from 90 to 110 nm by negative staining [15]. Each of the three viral RNA segments forms ribonucleoprotein (RNP) complexes (viral core; 66 nm diameter) with the viral nucleocapsid protein (N) and the viral RNA-dependent RNA polymerase (L) which are packaged in a lipid bilayer (envelope) obtained during budding into the Golgi complex during virion assembly [16]. The envelope is made up of dimers of the viral glycoproteins, Gn and Gc, organized with icosahedral symmetry and projecting as 12-nm-long cylindrical spikes on the envelope [14, 17, 18]. The 12 pentamers and 110 hexamers are arranged on the viral surface with icosahedral symmetry having a T12 triangulation number [14]. Located 2.2 nm apart, the pentons (12 nm diameter) are positioned around the fivefold symmetry axis and hexons (14 nm diameter) organized around the threefold, quasi threefold, and twofold axes [14, 19]. The virus particle is estimated to contain 720 glycoproteins of both Gn and Gc with each penton made of 10 glycoproteins, and each hexon made of 12 glycoprotein molecules of Gc or Gn monomers, homodimers, or heterodimers [13]. A gap of 4.5 nm exists between the viral envelope and the RNP core [14]. The RNP has a string-like structure and forms a pan-handle structure due to the complementarity of the 5′ and 3′ genomic termini [20].
Genome organization
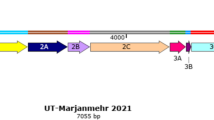
The genome is composed of a tri-segmented, negative-stranded RNA that encodes for seven proteins (Fig. 2). The 6.4 kb large (L) segment encodes the viral RNA-dependent RNA polymerase (RdRp; L protein). The 3.2 kb medium (M) segment encodes a single open reading frame (ORF) which produces the envelope glycoproteins Gn and Gc, and non-structural proteins: NSm (14 kDa) and the 78 kDa Gn/NSm fusion protein; these proteins are produced by alternative usage of multiple AUG start codons at the 5′ end of the mRNA and co-translational cleavage of the respective precursor proteins [21,22,23,24]. Translation initiation from the first AUG results in the synthesis of the 78 kDa Gn/NSm protein, and the NSm protein is produced from the second AUG. Protease cleavage at the N-terminus of Gn releases a mature NSm protein. Translation from the third AUG was recently reported to generate a truncated NSm protein termed NSm′ [25]. This protein lacks 38 amino acids from the N-terminus of NSm and when compared with NSm, is predominantly expressed in infected Vero cells [25]. Translation from the fourth or fifth AUG produces Gn and Gc, both of which have a single ectodomain with 1 or 4 potential N-glycosylation sites, respectively [21,22,23,24]. The 1.7 kb small (S) segment is ambisense, with the nucleoprotein N encoded in the genomic-sense, and NSs in the antigenomic orientation. Transcription termination sites are present at both viral-sense and antiviral-sense RNA located between the N and NSs ORFs. Short untranslated regions (UTRs) flank each genomic segment which hybridize to form circular RNP structures and are important for both transcription and replication [26].
Genomic organization of Rift Valley fever virus. The encoded open reading frames are shown as solid black arrows on the large (L), medium (M), and small (S) segments of the viral genome. Untranslated regions (UTR) flank each segment. The length of each genomic segment is indicated in kilo bases (kb). Transcribed mRNAs are depicted as dashed arrows with the start codons (AUG) indicated as black boxes, and the resulting proteins shown as shaded boxes. Proteins encoded by the M segment are produced by posttranslational cleavage of translated polyproteins. The nucleoprotein is generated by an mRNA transcribed from the negative-sense genomic viral RNA (vRNA). The non-structural S protein (NSs) is represented on the genomic-sense S vRNA as a white box and is produced from an mRNA transcribed from the antigenomic copy vRNA generated during viral replication. The octagon represents an intergenic transcription termination site
Viral protein functions
Gn and Gc are important for viral attachment and entry [14]. The envelope glycoprotein Gn is responsible for recruitment of RdRp to the Golgi membranes [27]. Gn is also independently involved in the packaging of N into virions [27]. The glycoproteins are also involved in efficient viral release from infected cells, for which the primary stimulus is the encapsidated genome [27, 28]. The N protein encapsidates genomic RNA by protein–RNA and protein–protein interactions and forms the RNP [13, 17]. The L protein forms part of the RNP core and plays a vital role in the primary transcription of mRNAs transcribed from the viral genome soon after viral entry and directs the genomic RNA replication in the cytoplasm [27, 29]. Gn, Gc, and N make up the major antigenic proteins and are therefore the common targets for diagnostic and vaccine development.
The non-structural proteins (NSm, 78 kDa Gn/NSm, and NSs) are associated with virulence. Both, NSm, and the 78 kDa protein were shown to be dispensable for viral replication [30]. However, it was observed that infection of Culex quinquefasciatus with RVFV lacking the 78 kDa/NSm displayed reduced transmissibility and infection rate whereas it was completely unable to infect Aedes aegypti mosquitoes [31]. NSm is targeted to the mitochondrial membrane and plays a role in delaying the onset of virus-induced apoptosis [32, 33]. NSs is also dispensable for the viral life cycle but is a major virulence factor for RVFV due to its ability to suppress host antiviral responses [34]. NSs is localized to both the cytoplasm and the nucleus where it forms filamentous structures [35]. One of the earliest known functions of NSs is its ability to inhibit IFN function [36]. It was later found that NSs specifically blocks transcription from the IFN-β promoter by interacting with the sin3A-associated protein (SAP30) through the transcription factor YY1. This interaction allows the repressor complex of SAP30, YY1, and Sin3A-associated corepressor factors to stay on the IFN-β promoter, resulting in the inhibition of transcriptional activation of the IFN-β promoter [37].
Another major virulence function of NSs is its ability to suppress general host gene expression by inhibiting the essential transcription factor TFIIH. NSs independently interacts with two subunits of TFIIH: p44, resulting in its sequestration [38] and p62, resulting in its proteasomal degradation [39,40,41]. Besides its effect on the host transcription machinery, NSs also prevents the induction of antiviral genes in infected cells by promoting the proteasomal degradation of dsRNA-dependent protein kinase (PKR) [28, 42]. PKR is activated in virus-infected cells, which results in increased phosphorylation of the translation initiation factor eIF2α which in turn results in translation suppression and a general inhibition of viral gene expression [43, 44]. Thus, inhibition of PKR by NSs allows RVFV protein synthesis to continue in virus-infected cells. A study by Kalveram et al. (2013) revealed that the host transcription inhibition and PKR degradation functions of NSs are independent of each other [45].
NSs also interacts with the host pericentromeric DNA sequences, thus inducing chromosome cohesion and segregation defects [46]. Another interesting observation is the NSs-induced nuclear relocalization of the host Poly(A) binding protein 1 (PABP1) during RVFV infection. PABP1 binds to the polyA-tail of cellular mRNAs and enhances host mRNA translation by bridging the polyA-tail with the 5′cap-binding eIF4F complex. RVFV-transcribed viral mRNAs, like other phleboviruses, do not have a polyA-tail and hence is independent of PABP1 function. By relocalizing PABP1 during RVFV infection, NSs ensures the inhibition of host mRNAs translation, thereby increasing the chances of viral mRNA translation by freeing up the host cell translation machinery [47]. Besides these functions, the multifunctional protein NSs also induces an increase in reactive oxygen species, activates the p53 signaling pathway, inhibits mRNA export from the nucleus, induces DNA damage response via the ataxia-telangectasia mutated (ATM) signaling pathway, and also inhibits cell cycle arrest either at the G0/G1 or S phase [48,49,50,51,52].
Replication and packaging
RVFV replication occurs in the cell cytoplasm and generally takes approximately 10–12 h to complete one cycle (Fig. 3a). Replication has been shown to start locally, near the endosomal fusion site [53]. The RNPs are released into the cytosol and transcribed, translated, and replicated within the first 6 h of infection, with an estimated 40 min doubling time [53]. Recruitment of genome segments to the Golgi occurs at 6–8 hpi, most likely through interactions between N and the cytoplasmic tail of Gn [27, 53, 54]. Gc is targeted to the endoplasmic reticulum (ER) by virtue of an ER-targeting motif, while Gn is retained at the Golgi by means of a Golgi-retention signal, thus allowing co-localization of both glycoproteins at the Golgi apparatus. The cytoplasmic tail of Gn recruits the RNP and the RdRp, and after assembly, the progeny virions bud from the Golgi [55,56,57]. While previous work suggested the packaging mechanism for RVFV is a selective process [58, 59], more recent evidence supports a non-selective process [53]. A recent study by Schreur and Kortekaas (2016) showed that RVF virions have heterogeneous segment composition, with 40% containing no genome segments, up to 25% lacking M, and only around 10% of virions contained all three genome segments [53]. Other studies have also demonstrated the plasticity of RVFV packaging through the efficient production of RVF replicon variants with two and four segmented genomes [60,61,62]. Furthermore, during coinfection with different virus strains, it is possible for replicated genomic segments from the parent infecting viruses to be packaged into virions, generating reassortant viruses (Fig. 3b). It is also known that during assembly, both negative- and positive-sense viral genomic RNAs are incorporated into the virion, thus allowing for the immediate transcription of both the N and NSs mRNAs [29, 63].
Rift Valley fever virus replication cycle. a Coinfection with two different virus strains is shown. Following attachment and entry into a host cell, the viral ribonucleoproteins (RNPs) which consist of viral genomic RNAs complexed with nucleoprotein and viral RNA-dependent RNA polymerase are released into the cytosol via endosomal fusion. Circularization of the linear, single-stranded genome segments occurs via hybridization of the flanking untranslated regions. Transcription, translation, and replication occur in the cytoplasm. Newly replicated viral genomic segments and proteins are recruited to the Golgi for assembly and packaging of virus particles. Mature virions are released from the infected cell. b Possible virus reassortants following coinfection with two virus strains are represented by solid- or dotted-line genomic segments
Genetic diversity of RVFV
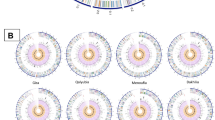
Sequencing and phylogenetic studies conducted over the last 20 years have shown the RVFV genome to be overall highly conserved, with most variances occurring as random single site mutations throughout the genome with no well-defined variable regions. This makes differentiation between strains without genome sequencing difficult, since there are also no well-defined serotypes. The first phylogenetic study based on classification by NSs of 18 different RVFV strains yielded only two genomic lineages: Egyptian and sub-Saharan [64]. Subsequent studies including partial and whole genome sequences have expanded the major lineages to Egypt, West Africa, and East Central Africa [65], then to 7 lineages designated A through G [66, 67], and more recently to 15 lineages classified as A through O [68]. The varying number of lineages identified can be explained by the varying number of analyzed sequences, the use of partial or whole genome sequences, and the methods used for analysis.
The first comprehensive whole genome analysis consisted of 33 RVFV isolates from various African countries and Saudi Arabia over a period spanning 1944–2000, with varying pathogenicity [66]. Included were samples collected during two large outbreaks in 1977–1978 Egypt and 1987 Mauritania, as well as, from 1970 Zimbabwe localized epizootic events. This study illustrated that while the virus strains were ecologically and biologically diverse, the overall genetic diversity was low. Pairwise identity of genome segments between isolates were similar, with nucleotide and amino acid differences of 5% and 2% for M, and 4% and 1% for the L and S segments, respectively. A later study evaluating partial M sequences of 203 isolates also found similar genetic pairwise differences [68].
Interestingly, higher diversity was observed among isolates from Zimbabwe during a low endemic period compared to the large outbreaks in Egypt and Mauritania, suggesting multiple strains concurrently circulating in Zimbabwe, and a single predominant strain in Egypt and Mauritania [66]. In contrast, a genetic analysis of a wide variety of samples collected during the Kenya 2006–2007 RVFV outbreak showed two lineages circulating during that time [69]. This translated to a pairwise nucleotide difference of over 1%, as compared to 0.2% reported for epidemics in Egypt and Mauritania [66]. Thus, these studies demonstrate the conserved nature of the RVFV genome, despite differences in virulence, and that multiple RVFV strains can be found co-circulating during both epidemic and inter-epidemic periods.
RVFV reassortment
Viral reassortment is the exchange of genomic segments between two or more virus strains during coinfection of a host cell (Fig. 3). This phenomenon can occur among segmented RNA viruses, including members of the Phlebovirus and Orthobunyavirus genera. Genomic reassortment among RVFV strains is also known to occur and has been demonstrated experimentally and documented among field isolates. The results from those studies are discussed in the following sections.
Experimental evidence
Viral reassortment was originally utilized to map the mutations which conferred attenuation of the RVFV MP-12 strain [70]. Reassortants isolated from Vero cells coinfected with MP-12 and wild-type ArD38661 were tested in mice, which revealed attenuating mutations on each of the three genome segments. Similarly, Turell et al. [71] used several monoclonal antibodies to investigate the potential for reassortment to occur in the mosquito vector by using a Culex mosquito and hamster model and the RVFV strains ZH501 [72], ArD38661 [73] and MP-12 [74]. Both studies detected only S and M reassortants, as there were no monoclonal antibodies to differentiate the L segment. Despite this limitation, the study by Turell et al. (1990) yielded several important findings regarding RVFV reassortment [71]. First, reassortment was shown to occur in mosquitoes intrathoracically injected with two different virus strains. This was demonstrated between two pathogenic strains (ZH501 and ArD38661), as well as, a pathogenic and attenuated strain (ArD38661 and MP-12). Second, prior infection with one strain (MP-12) was shown to interfere with replication of a second homologous strain in the mosquito (ArD38661). Third, reassortants were isolated from mosquitoes that intermittently fed on a hamster infected with the ArD38661 strain and subsequently fed on a hamster infected with ZH501, as well as, from mosquitoes fed on a hamster coinfected with both strains. Finally, reassortant viruses were transmitted to naïve hamsters by infected mosquito bites. Thus, this study showed that reassortment can occur between two RVFV strains during coinfection, at the vector level. Furthermore, these two studies demonstrated reassortment between a pathogenic and attenuated strain, ArD38661 and MP-12, respectively.
A more recent study used an in vitro model system to investigate the risk of RVFV MP-12 strain to reassort with other related viruses of the family Phenuiviridae of the genus Goukovirus, and Arumowot virus, an unclassified Phlebovirus [75]. The results showed that coinfected cell cultures with either of the two viruses did not produce any viable reassortants with RVFV MP-12. However, reassortment did occur between RVFV MP-12 and a genetic MP-12 variant in coinfected C6/36 mosquito cells. Collectively, these in vitro studies demonstrate that reassortment can readily occur and is a conserved event between RVFV strains.
Examples in nature
Examples of RVFV reassortment that have occurred under natural conditions from 5 separate field and case studies have been reported (Table 1). The first identification of RVFV reassortment from field isolates was by Sall et al. [65]. In that study, 20 isolates were subjected to phylogenetic analysis, revealing 3 major lineages based on geographic regions: West Africa, Egypt and Central-East Africa. Two tests used to analyze the incongruence in topology identified five reassortants. The reassortants identified were isolated from various sources between 1984 and 1993 in Senegal, Guinea, and Mauritania (Table 1) and consisted of reassortment of the S or L genomic segments [65].
Whole genome sequence analysis of 33 RVFV isolates suggested reassortment was not common since there was less incongruence in their topology; however, there was some evidence indicating the occurrence of reassortment events [66]. From the seven phylogenetic lineages (A–G), strain 73HB1230 was found to have S and M segments belonging to lineage B, while the L segment grouped with lineage A. Furthermore, there was phylogenetic evidence of an earlier reassortant event with the S segment of this strain. In addition, there was support for reassortment among M segments of lineages D and E although a later analysis by Grobbelaar et al. (2011) did not find clear evidence to support this as a reassortment event [68].
Grobbelaar et al. [68] performed partial sequencing of the M segments of an additional 170 RVFV field isolates, expanding on the previous 33 sequenced genomes [66]. Altogether, the 203 isolates were collected over a period of 67 years (1944–2010) from Saudi Arabia and 16 countries of Africa. Following phylogenetic analysis, five identified divergent isolates were further sequenced and analyzed for possible genetic reassortment. From that analysis, an M segment reassortant (SA184/10) was identified. The reassortant was isolated from a patient with an accidental needle stick injury while vaccinating sheep, which could have been potentially exposed to both the live attenuated Smithburn vaccine strain and wild-type virus. Phylogenetic investigation revealed that the M segment of the isolated virus grouped with the Smithburn vaccine strain of lineage K, and the S and L segments were closely related to SA54/10 which was isolated from another patient in the same area. In addition, there was evidence of historical reassortment events for three other isolates: H1739 and H1825 from the first recorded human deaths caused by RVF in 1975, and 95EG Cow-2509 isolated from an aborted fetus of a cow vaccinated with 95EG in Egypt [68].
Another M reassortant was identified from an analysis of 31 samples collected during the Kenya 2006–2007 RVFV outbreak [69]. This was later confirmed by Freire et al. (2015) using a set of tools initially used to study reassortment of influenza viruses. Using this analysis framework, pairs of segments were compared revealing five additional M and S reassortants, as well as, a new L/M reassortant, TANTan00107, shown in Table 1 [76].
More recently, an S reassortant was identified from a man working in Angola in 2016 during a non-epidemic period [67]. Similar to the previous studies, phylogenetic analysis revealed seven lineages, A–G. The reassortment occurred between lineages A and E, with the L and M segments most closely related to a South African isolate from sheep in 2009, and S grouping with several strains spanning regions from Egypt to South Africa and Madagascar from the 1970s to 2004 (lineage A). The patient was a fork-lifter in a rural area, frequently exposed to mosquitoes with no history of contact with livestock or febrile humans.
Together, these data (summarized in Table 1) show that reassortment between RVFV strains occurs naturally in the field, and multiple examples of reassortants for each segment type have been identified. Furthermore, reassortants have been isolated from a wide range of sources, and during epidemic and inter-epidemic periods or regions.
Frequency of RVFV reassortment
In earlier studies, 7 out of 50 (14%) viruses isolated from Vero cells coinfected with the RVFV MP-12 and ArD38661 strains identified as reassortants [70]. Reassortants were also found from all ten mosquitoes simultaneously coinfected with those same strains, although the percentage of reassortants was not reported [71]. Reassortants made up 27% (32/119) of viruses recovered from mosquitoes coinfected with ZH501 and ArD38661 [71]. However, because those studies lacked the ability to differentiate the L segment, the percentages reflect only S and M reassortants.
More recently, a study with the RVFV MP-12 strain and a genetic variant showed out of 47 plaques isolated from coinfected C6/36 mosquito cells, 83% were reassortants [75]. The reassortants recovered were further differentiated by segment and consisted of 36% L reassortants, 15% M, and 32% were S. It is not known whether reassortment occurs preferentially for a particular segment, but experimental [75] and field data (summarized in Table 1) indicates that reassortment can occur among any of the three genomic segments. Furthermore, there is evidence that RVFV uses a non-selective packaging strategy, suggesting that reassortment between segments may also be non-selective [53].
While experimental infections in mosquitoes suggest mosquitoes are permissive for generation of RVFV reassortants (27% collectively), fewer reassortants (only 2) have been recovered from infected mosquitoes from the field (Fig. 4; Table 1). This most likely reflects the bias of optimized experimental conditions which may not be representative of those encountered in the field, as well as the sources and number sampled. For example, infection of mosquitoes by Turell et al. (1990) was performed by intrathoracic inoculation which bypasses the midgut infection/escape barrier which could correlate to a higher reassortant frequency because the reassortant viruses did not undergo the same selective pressure as orally fed viruses [71]. Concerning is the comparatively high number of reassortant viruses isolated from humans (Fig. 4b), especially given the fact that other reassortant bunyaviruses are associated with severe human illness [77, 78]. Still, it is not clear whether reassortment occurs more frequently in humans, ruminants, other mammals, or mosquitoes; or whether those RVFV reassortants isolated possess altered pathogenicity and/or host range.
Rift Valley fever virus reassortment. a Frequency of experimental and field reported reassortants. The number of reassortants out of total virus isolates evaluated is shown, with percentage of reassortants indicated. b Sources from which reassortants have been isolated in the field are shown; the number of reassortants isolated from each source derived from Table 1 are indicated
Reversion to virulence
Reassortment of an attenuated RVFV with a pathogenic strain resulting in reversion to a virulent phenotype is of significant concern given the wide use of attenuated and modified live vaccine strains in endemic countries. For example, the attenuated live Smithburn vaccine strain has been widely used throughout Africa for more than six decades and, in fact, has been identified as reassorting with a wild-type field strain [68]. The live attenuated MP-12 strain has been pursued as a more desirable alternative to earlier live attenuated strains such as Smithburn. Reassortment between MP-12 and a pathogenic RVFV strain, ArD38661, was demonstrated both in vitro [70] and in vivo [71]. Reassortants isolated from MP-12/ArD3811 coinfected Vero cells and tested for virulence in mice were found to possess an attenuated phenotype for each of the three MP-12 genome segments [70]. Furthermore, genetic analysis showed multiple mutations on each segment of the MP-12 genome conferred attenuation, suggesting the risk of MP-12 reverting to a virulent phenotype via reassortment is low or unlikely [79, 80]. Nonetheless, another study showed the introduction of a few revertant mutations within the MP-12 L and M segments following 25 passages in Vero or MRC-5 cells [81]. Additionally, there is still the risk of other live attenuated vaccine strains that do not possess attenuations in each of the three genome segments to reassort with wild-type viruses.
Reassortment with related viruses
Reassortment of RVFV with other closely related viruses (e.g., other members of Phenuiviridae) is also a concern, especially with the co-circulation of multiple bunyaviruses in the field and the continued discovery of new viruses. For example, Ngari virus was detected during a RVFV outbreak in Mauritania in 2010, with serological evidence of a possible coinfection in a goat [82]. Ngari virus itself is a reassortant of Bunyamwera virus and Batai virus, both classified under the genus Orthobunyavirus [77, 78, 83], and is associated with hemorrhagic fever in humans, although clinical disease in animals is not currently known. Both Bunyamwera and Batai viruses infect humans, animals, and arthropods. The concern is the possibility that reassortment could lead to more virulent strains which are easily transmittable and which could have a wider vector or host range. An experimental approach to reassortment between the MP-12 strain and bunyavirus species closely related to RVFV was performed by Ly et al. (2017) using the Arumowot virus (AMTV) and the Gouleako goukovirus (GOLV), both transmitted by mosquitoes in Africa. The results of this study showed that AMTV and GOLV do not form detectable reassortant strains with the MP-12 strain in coinfected C6/36 cells, most likely due to their incompatibility among the N, L, and Gn/Gc proteins. While reassortment of RVFV with other related viruses may be less likely, as shown by Ly et al. (2017), it should be taken in to consideration and not overlooked [75].
Concluding remarks
While there is strong evidence that RVFV genetic reassortment does occur, there is much that remains unknown. More studies are warranted to address the following critical questions regarding RVFV reassortment.
-
What is the frequency of reassortants recovered from naturally infected vectors (different mosquito species), ruminants (different species), non-ruminants, and humans?
-
What is the potential of emergence of reassortants between wild-type RVFV and attenuated live RVFV vaccine strains (e.g., Smithburn, MP-12), or other members of Phenuiviridae?
-
Is there preference based on a particular segment for reassortment?
-
What are the phenotypic consequences of such reassortants? Are reassortants more fit? More virulent? Have a wider host range?
-
Ultimately, how do all of the above criteria relate to risk to humans and animals?
These questions are not easily answered given the complexity of virus–vector–host interactions, and the constraints of working with a Select Agent. Experimental studies are laborious and ultimately require genetic modification of the viruses used or sequencing to confirm a reassortment event due to the low genetic diversity among RVFV strains. Data collection from larger sampling numbers will be necessary to more accurately assess frequency rates and risk. While certain animal species and virus–vector interaction studies are possible to address the consequence of reassortants, recapitulation of natural conditions in experimental studies with humans is not possible. Thus, study models such as non-human primate and well-documented field cases are necessary for addressing these questions in humans. As molecular virological techniques such as sequencing becomes more affordable and bioinformatic tools become more sophisticated, the elucidation of these questions should become more attainable. The limited number of earlier studies discussed in this review provide insight and pave the way to address these key questions regarding RVFV reassortment.
References
Ikegami T, Makino S (2011) The pathogenesis of Rift Valley fever. Viruses 3:493–519
Indran SV, Ikegami T (2012) Novel approaches to develop Rift Valley fever vaccines. Front Cell Infect Microbiol. https://doi.org/10.3389/fcimb.2012.00131
Faburay B, LaBeaud AD, McVey DS, Wilson WC, Richt JA (2017) Current status of Rift Valley fever vaccine development. Vaccines. https://doi.org/10.3390/vaccines5030029
Linthicum KJ, Britch SC, Anyamba A (2016) Rift Valley fever: an emerging mosquito-borne disease. Annu Rev Entomol 61:395–415
Lumley S et al (2017) Rift Valley fever virus: strategies for maintenance, survival and vertical transmission in mosquitoes. J Gen Virol 98:875–887
Olive MM, Goodman SM, Reynes JM (2012) The role of wild mammals in the maintenance of Rift Valley fever virus. J Wildl Dis 48:241–266
Beechler BR et al (2015) Rift Valley fever in Kruger national park: do buffalo play a role in the inter-epidemic circulation of virus? Transbound Emerg Dis 62:24–32
Mroz C et al (2017) Seroprevalence of Rift Valley fever virus in livestock during inter-epidemic period in Egypt, 2014/15. BMC Vet Res. https://doi.org/10.1186/s12917-017-0993-8
Brustolin M et al (2017) Rift Valley fever virus and European mosquitoes: vector competence of Culex pipiens and Stegomyia albopicta (= Aedes albopictus). Med Vet Entomol 31:365–372
Turell MJ, Wilson WC, Bennett KE (2010) Potential for North American mosquitoes (Diptera: Culicidae) to transmit rift Valley fever virus. J Med Entomol 47:884–889
Turell MJ et al (2015) Potential for Psorophora columbiae and Psorophora ciliata mosquitoes (Diptera: Culicidae) to transmit Rift Valley fever virus. J Med Entomol 52:1111–1116
Gaudreault NN, Indran SV, Bryant PK, Richt JA, Wilson WC (2015) Comparison of Rift Valley fever virus replication in North American livestock and wildlife cell lines. Front Microbiol 6:664
Adams MJ et al (2017) Changes to taxonomy and the International Code of Virus Classification and Nomenclature ratified by the International Committee on Taxonomy of Viruses (2017). Arch Virol 162:2505–2538
Freiberg AN, Sherman MB, Morais MC, Holbrook MR, Watowich SJ (2008) Three-dimensional organization of Rift Valley fever virus revealed by cryoelectron tomography. J Virol 82:10341–10348
Ellis DS, Simpson DI, Stamford S, Abdel Wahab KS (1979) Rift Valley fever virus: some ultrastructural observations on material from the outbreak in Egypt 1977. J Gen Virol 42:329–337
Hornak KE, Lanchy JM, Lodmell JS (2016) RNA encapsidation and packaging in the phleboviruses. Viruses. https://doi.org/10.3390/v8070194
Huiskonen JT, Overby AK, Weber F, Grunewald K (2009) Electron cryo-microscopy and single-particle averaging of Rift Valley fever virus: evidence for GN-GC glycoprotein heterodimers. J Virol 83:3762–3769
Sherman MB, Freiberg AN, Holbrook MR, Watowich SJ (2009) Single-particle cryo-electron microscopy of Rift Valley fever virus. Virology 387:11–15
Rusu M et al (2012) An assembly model of Rift Valley fever virus. Front Microbiol 3:254
Raymond DD, Piper ME, Gerrard SR, Smith JL (2010) Structure of the Rift Valley fever virus nucleocapsid protein reveals another architecture for RNA encapsidation. Proc Natl Acad Sci USA 107:11769–11774
Kakach LT, Suzich JA, Collett MS (1989) Rift Valley fever virus M segment: phlebovirus expression strategy and protein glycosylation. Virology 170:505–510
Wasmoen TL, Kakach LT, Collett MS (1988) Rift Valley fever virus M segment: cellular localization of M segment-encoded proteins. Virology 166:275–280
Suzich JA, Kakach LT, Collett MS (1990) Expression strategy of a phlebovirus: biogenesis of proteins from the Rift Valley fever virus M segment. J Virol 64:1549–1555
Kakach LT, Wasmoen TL, Collett MS (1988) Rift Valley fever virus M segment: use of recombinant vaccinia viruses to study Phlebovirus gene expression. J Virol 62:826–833
Kreher F et al (2014) The Rift Valley fever accessory proteins NSm and P78/NSm-G(N) are distinct determinants of virus propagation in vertebrate and invertebrate hosts. Emerg Microbes Infect 3(10):e71. https://doi.org/10.1038/emi.2014.71
Gauliard N, Billecocq A, Flick R, Bouloy M (2006) Rift Valley fever virus noncoding regions of L, M and S segments regulate RNA synthesis. Virology 351:170–179
Piper ME, Sorenson DR, Gerrard SR (2011) Efficient cellular release of Rift Valley fever virus requires genomic RNA. PLoS ONE 6:e18070
Habjan M et al (2009) NSs protein of Rift Valley fever virus induces the specific degradation of the double-stranded RNA-dependent protein kinase. J Virol 83:4365–4375
Ikegami T, Won S, Peters CJ, Makino S (2005) Rift Valley fever virus NSs mRNA is transcribed from an incoming anti-viral-sense S RNA segment. J Virol 79:12106–12111
Gerrard SR, Bird BH, Albarino CG, Nichol ST (2007) The NSm proteins of Rift Valley fever virus are dispensable for maturation, replication and infection. Virology 359:459–465
Crabtree MB et al (2012) Infection and transmission of Rift Valley fever viruses lacking the NSs and/or NSm genes in mosquitoes: potential role for NSm in mosquito infection. PLoS Negl Trop Dis 6:e1639
Won SY, Ikegami T, Peters CJ, Makino S (2007) NSm protein of Rift Valley fever virus suppresses virus-induced apoptosis. J Virol 81:13335–13345
Terasaki K, Won S, Makino S (2013) The C-terminal region of Rift Valley fever virus NSm protein targets the protein to the mitochondrial outer membrane and exerts antiapoptotic function. J Virol 87:676–682
Lihoradova O, Ikegami T (2014) Countermeasure development for Rift Valley fever: deletion, modification or targeting of major virulence factor NSs. Future Virol 9:27–39
Yadani FZ, Kohl A, Prehaud C, Billecocq A, Bouloy M (1999) The carboxy-terminal acidic domain of Rift Valley Fever virus NSs protein is essential for the formation of filamentous structures but not for the nuclear localization of the protein. J Virol 73:5018–5025
Bouloy M et al (2001) Genetic evidence for an interferon-antagonistic function of Rift Valley fever virus nonstructural protein NSs. J Virol 75:1371–1377
Le May N et al (2008) A SAP30 complex inhibits IFN-beta expression in Rift Valley fever virus infected cells. PLoS Pathog 4:e13
Dasgupta A (2004) Targeting TFIIH to inhibit host cell transcription by Rift Valley fever virus. Mol Cell 13:456–458
Cyr N et al (2015) A OmegaXaV motif in the Rift Valley fever virus NSs protein is essential for degrading p62, forming nuclear filaments and virulence. Proc Natl Acad Sci USA 112:6021–6026
Kainulainen M et al (2014) Virulence factor NSs of Rift Valley fever virus recruits the F-box protein FBXO3 to degrade subunit p62 of general transcription factor TFIIH. J Virol 88:3464–3473
Kalveram B, Lihoradova O, Ikegami T (2011) NSs protein of Rift Valley fever virus promotes posttranslational downregulation of the TFIIH subunit p62. J Virol 85:6234–6243
Ikegami T et al (2009) Rift Valley fever virus NSs protein promotes post-transcriptional downregulation of protein kinase PKR and inhibits eIF2alpha phosphorylation. PLoS Pathog 5:e1000287
Dalet A, Gatti E, Pierre P (2015) Integration of PKR-dependent translation inhibition with innate immunity is required for a coordinated anti-viral response. FEBS Lett 589:1539–1545
Gale M Jr, Tan SL, Katze MG (2000) Translational control of viral gene expression in eukaryotes. Microbiol Mol Biol Rev 64:239–280
Kalveram B, Lihoradova O, Indran SV, Head JA, Ikegami T (2013) Using click chemistry to measure the effect of viral infection on host-cell RNA synthesis. J Vis Exp (78):e50809. https://doi.org/10.3791/50809
Mansuroglu Z et al (2010) Nonstructural NSs protein of Rift Valley fever virus interacts with pericentromeric DNA sequences of the host cell, inducing chromosome cohesion and segregation defects. J Virol 84:928–939
Copeland AM, Altamura LA, Van Deusen NM, Schmaljohn CS (2013) Nuclear relocalization of polyadenylate binding protein during Rift Valley fever virus infection involves expression of the NSs gene. J Virol 87:11659–11669
Copeland AM, Van Deusen NM, Schmaljohn CS (2015) Rift Valley fever virus NSs gene expression correlates with a defect in nuclear mRNA export. Virology 486:88–93
Ly HJ, Ikegami T (2016) Rift Valley fever virus NSs protein functions and the similarity to other bunyavirus NSs proteins. Virol J 13:118
Austin D et al (2012) p53 Activation following Rift Valley fever virus infection contributes to cell death and viral production. PLoS ONE 7:e36327
Baer A et al (2012) Induction of DNA damage signaling upon Rift Valley fever virus infection results in cell cycle arrest and increased viral replication. J Biol Chem 287:7399–7410
Narayanan A et al (2014) Reactive oxygen species activate NFkappaB (p65) and p53 and induce apoptosis in RVFV infected liver cells. Virology 449:270–286
Schreur PJW, Kortekaas J (2016) Single-molecule FISH reveals non-selective packaging of Rift Valley fever virus genome segments. PLoS Pathog 12:e1005800
Carnec X, Ermonval M, Kreher F, Flamand M, Bouloy M (2014) Role of the cytosolic tails of Rift Valley fever virus envelope glycoproteins in viral morphogenesis. Virology 448:1–14
Gerrard SR, Rollin PE, Nichol ST (2002) Bidirectional infection and release of Rift Valley fever virus in polarized epithelial cells. Virology 301:226–235
Gerrard SR, Nichol ST (2002) Characterization of the Golgi retention motif of Rift Valley fever virus G(N) glycoprotein. J Virol 76:12200–12210
Gerrard SR, Nichol ST (2007) Synthesis, proteolytic processing and complex formation of N-terminally nested precursor proteins of the Rift Valley fever virus glycoproteins. Virology 357:124–133
Murakami S, Terasaki K, Narayanan K, Makino S (2012) Roles of the coding and noncoding regions of Rift Valley fever virus RNA genome segments in viral RNA packaging. J Virol 86:4034–4039
Terasaki K, Murakami S, Lokugamage KG, Makino S (2011) Mechanism of tripartite RNA genome packaging in Rift Valley fever virus. Proc Natl Acad Sci USA 108:804–809
Brennan B, Welch SR, McLees A, Elliott RM (2011) Creation of a recombinant Rift Valley fever virus with a two-segmented genome. J Virol 85:10310–10318
Brennan B, Welch SR, Elliott RM (2014) The consequences of reconfiguring the ambisense S genome segment of Rift Valley fever virus on viral replication in mammalian and mosquito cells and for genome packaging. PLoS Pathog 10:e1003922
Wichgers Schreur PJ, Oreshkova N, Moormann RJ, Kortekaas J (2014) Creation of Rift Valley fever viruses with four-segmented genomes reveals flexibility in bunyavirus genome packaging. J Virol 88:10883–10893
Ikegami T, Peters CJ, Makino S (2005) Rift Valley fever virus nonstructural protein NSs promotes viral RNA replication and transcription in a minigenome system. J Virol 79:5606–5615
Sall AA et al (1997) Variability of the NS(S) protein among Rift Valley fever virus isolates. J Gen Virol 78(Pt 11):2853–2858
Sall AA et al (1999) Genetic reassortment of Rift Valley fever virus in nature. J Virol 73:8196–8200
Bird BH, Khristova ML, Rollin PE, Ksiazek TG, Nichol ST (2007) Complete genome analysis of 33 ecologically and biologically diverse Rift Valley fever virus strains reveals widespread virus movement and low genetic diversity due to recent common ancestry. J Virol 81:2805–2816
Liu J et al (2017) The first imported case of Rift Valley fever in China reveals a genetic reassortment of different viral lineages. Emerg Microbes Infect 6:e4
Grobbelaar AA et al (2011) Molecular epidemiology of Rift Valley fever virus. Emerg Infect Dis 17:2270–2276
Bird BH et al (2008) Multiple virus lineages sharing recent common ancestry were associated with a large Rift Valley fever outbreak among livestock in Kenya during 2006–2007. J Virol 82:11152–11166
Saluzzo JF, Smith JF (1990) Use of reassortant viruses to map attenuating and temperature-sensitive mutations of the Rift Valley fever virus MP-12 vaccine. Vaccine 8:369–375
Turell MJ, Saluzzo JF, Tammariello RF, Smith JF (1990) Generation and transmission of Rift Valley fever viral reassortants by the mosquito Culex pipiens. J Gen Virol 71(Pt 10):2307–2312
Meegan JM (1979) The Rift Valley fever epizootic in Egypt 1977-78. 1. Description of the epizootic and virological studies. Trans R Soc Trop Med Hyg 73:618–623
Saluzzo JF, Anderson GW Jr, Hodgson LA, Digoutte JP, Smith JF (1989) Antigenic and biological properties of Rift Valley fever virus isolated during the 1987 Mauritanian epidemic. Res Virol 140:155–164
Caplen H, Peters CJ, Bishop DH (1985) Mutagen-directed attenuation of Rift Valley fever virus as a method for vaccine development. J Gen Virol 66(Pt 10):2271–2277
Ly HJ, Lokugamage N, Nishiyama S, Ikegami T (2017) Risk analysis of inter-species reassortment through a Rift Valley fever phlebovirus MP-12 vaccine strain. PLoS ONE 12:e0185194
Freire CC et al (2015) Reassortment and distinct evolutionary dynamics of Rift Valley fever virus genomic segments. Sci Rep 5:11353
Gerrard SR, Li L, Barrett AD, Nichol ST (2004) Ngari virus is a Bunyamwera virus reassortant that can be associated with large outbreaks of hemorrhagic fever in Africa. J Virol 78:8922–8926
Briese T, Bird B, Kapoor V, Nichol ST, Lipkin WI (2006) Batai and Ngari viruses: M segment reassortment and association with severe febrile disease outbreaks in East Africa. J Virol 80:5627–5630
Vialat P, Muller R, Vu TH, Prehaud C, Bouloy M (1997) Mapping of the mutations present in the genome of the Rift Valley fever virus attenuated MP12 strain and their putative role in attenuation. Virus Res 52:43–50
Ikegami T et al (2015) Rift Valley fever virus MP-12 vaccine is fully attenuated by a combination of partial attenuations in the S, M, and L segments. J Virol 89:7262–7276
Lokugamage N, Ikegami T (2017) Genetic stability of Rift Valley fever virus MP-12 vaccine during serial passages in culture cells. NPJ Vaccines 2. https://doi.org/10.1038/s41541-017-0021-9
Eiden M et al (2014) Ngari virus in goats during Rift Valley fever outbreak, Mauritania, 2010. Emerg Infect Dis 20:2174–2176
Bowen MD et al (2001) A reassortant bunyavirus isolated from acute hemorrhagic fever cases in Kenya and Somalia. Virology 291:185–190
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Additional information
Edited by Joachim Jakob Bugert.
Rights and permissions
About this article
Cite this article
Gaudreault, N.N., Indran, S.V., Balaraman, V. et al. Molecular aspects of Rift Valley fever virus and the emergence of reassortants. Virus Genes 55, 1–11 (2019). https://doi.org/10.1007/s11262-018-1611-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11262-018-1611-y