Abstract
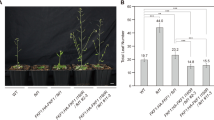
The EARLY FLOWERING 4 (ELF4) gene has been reported as a requirement for photoperiod perception and circadian function. In this study, an ELF4 homolog gene EARLY FLOWERING 4-like4 (EFL4) was isolated from Doritaenopsis “Tinny Tender” (Doritaenopsis Happy Smile × Happy Valentine) and designated as DhEFL4. The acquired DhEFL4 cDNA was 593 bp with a 345 bp open reading frame encoding 114 amino acids. The putative DhEFL4 protein contained a conserved region, DUF1313, which is shared by the ELF4 family. Amino acid sequence alignment showed that DhEFL4 shared high similarity to ELF4 and ELF4-like homologs from other species. The DhEFL4 promoter was also obtained, and 241 cis-elements were found. The diurnal expression of DhEFL4, which may be affected by the EVENINGAT (EE) and circadian motif was analyzed by quantitative PCR under short- and long-day conditions. Reverse-transcription PCR analysis results showed that DhEFL4 is an evening-expressed circadian gene. DhEFL4 was expressed in both vegetative and reproductive organs, and its transcripts reached higher levels in the buds. To further study the function of the DhEFL4 gene, DhEFL4 was overexpressed in Arabidopsis thaliana by Agrobacterium-mediated transformation. The flowering of these transgenic plants was more delayed than that of wild-type Arabidopsis by 12−14 days, and flower stems increased. Results indicated that DhEFL4 may perform an important function in the regulation of flowering.





Similar content being viewed by others
Abbreviations
- ELF4:
-
EARLY FLOWERING 4
- EFL4:
-
EARLY FLOWERING 4-LIKE4
- EE:
-
EVENINGAT
- SD:
-
Short day
- LD:
-
Long day
- CCA1:
-
CIRCADIAN CLOCK ASSOCIATED 1
- LHY:
-
LATE ELONGATED HYPOCOTYL
- TOC1:
-
TIMING OF CAB EXPRESSION 1
- ELF3:
-
EARLY FLOWERING 3
- LUX:
-
LUX ARRHYTHMO
- CaMV:
-
Cauliflower mosaic virus
- EST:
-
Expressed sequence tag
- Lhc:
-
Light-harvesting complex protein genes
- Lhc b:
-
Light-harvesting chlorophyll a/b binding protein genes of PSII
- CO:
-
CONSTANS
- RACE:
-
Rapid amplification of cDNA ends
- ZT:
-
Zeitgeber time
- qRT-PCR:
-
Quantitative real-time PCR
- PCR:
-
Polymerase chain reaction
- MeJA:
-
Methyl jasmonate
- MS:
-
Murashige and Skoog
- ORF:
-
Open reading frame
- RT-PCR:
-
Reverse transcription-polymerase chain reaction
- WT:
-
Wild-type
References
Adeyemo OS, Kolmos E, Tohme J, Chavariaga P, Fregene M, Davis SJ (2011) Identification and characterization of the cassava core-clock gene EARLY FLOWERING 4. Trop Plant Biol 4:117–125. doi:10.1007/s12042-011-9065-6
Alabadi D, Oyama T, Yanovsky MJ, Harmon FG, Mas P, Kay SA (2001) Reciprocal regulation between TOC1 and LHY/CCA1 within the Arabidopsis circadian clock. Science 293:880–883. doi:10.1126/science.1061320
Alabadi D, Yanovsky MJ, Mas P, Harmer SL, Kay SA (2002) Critical role for CCA1 and LHY in maintaining circadian rhythmicity in Arabidopsis. Curr Biol 12:757–761. doi:10.1016/s0960-9822(02)00815-1
Barrett JW, Beech RN, Dancik BP, Strobeck C (1994) A genomic clone of a type-I cab gene encoding a light-harvesting chlorophyll a/b binding-protein of photosystem-II identified from lodgepole pine. Genome 37:166–172. doi:10.1139/g94-021
Blanchard MG, Runkle ES (2006) Temperature during the day, but not during the night, controls flowering of Phalaenopsis orchids. J Exp Bot 57:4043–4049. doi:10.1093/jxb/erl176
Clough SJ, Bent AF (1998) Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J 16:735–743. doi:10.1046/j.1365-313x.1998.00343.x
Cui YY, Pandey DM, Hahn EJ, Paek KY (2004) Effect of drought on physiological aspects of Crassulacean acid metabolism in Doritaenopsis. Plant Sci 167:1219–1226. doi:10.1016/j.plantsci.2004.06.011
Doyle JJ, Doyle JL, Brown A (1990) A chloroplast-DNA phylogeny of the wild perennial relatives of soybean (Glycine subgenus Glycine): congruence with morphological and crossing groups. Evolution 44:371-389
Doyle MR et al (2002) The ELF4 gene controls circadian rhythms and flowering time in Arabidopsis thaliana. Nature 419:74–77. doi:10.1038/nature00954
Griesbach RJ (2002) Development of Phalaenopsis orchids for the mass-market. Trends in new crops and new uses. Proceedings of the Fifth National Symposium, Atlanta, Georgia, USA, 10–13 November, 2001
Harmer SL, Kay SA (2005) Positive and negative factors confer phase-specific circadian regulation of transcription in Arabidopsis. Plant Cell 17:1926–1940. doi:10.1105/tpc.105.033035
Harmer SL et al (2000) Orchestrated transcription of key pathways in Arabidopsis by the circadian clock. Science 290:2110–2113. doi:10.1126/science.290.5499.2110
Harmer SL, Panda S, Kay SA (2001) Molecular bases of circadian rhythms. Annu Rev Cell Dev Biol 17:215–253. doi:10.1146/annurev.cellbio.17.1.215
Hazen SP, Schultz TF, Pruneda-Paz JL, Borevitz JO, Ecker JR, Kay SA (2005) LUX ARRHYTHMO encodes a Myb domain protein essential for circadian rhythms. Proc Natl Acad Sci U S A 102:10387–10392. doi:10.1073/pnas.0503029102
Higo K, Ugawa Y, Iwamoto M, Korenaga T (1999) Plant cis-acting regulatory DNA elements (PLACE) database: 1999. Nucleic Acids Res 27:297–300
Hsiao Y-Y et al. (2011a) Gene discovery using next-generation pyrosequencing to develop ESTs for Phalaenopsis orchids. BMC Genomics 12 doi:10.1186/1471-2164-12-360
Hsiao YY et al (2011b) Research on orchid biology and biotechnology. Plant Cell Physiol 52:1467–1486. doi:10.1093/pcp/pcr100
Khanna R, Kikis EA, Quail PH (2003) EARLY FLOWERING 4 functions in phytochrome B-regulated seedling de-etiolation. Plant Physiol 133:1530–1538. doi:10.1104/pp. 103.030007
Kiselev KV, Turlenko AV, Zhuravlev YN (2010) Structure and expression profiling of a novel calcium-dependent protein kinase gene PgCDPK1a in roots, leaves, and cell cultures of Panax ginseng plant cell tissue and organ. Culture 103:197–204. doi:10.1007/s11240-010-9767-8
Kolmos E, Davis SJ (2007) ELF4 as a central gene in the circadian clock. Plant Signal Behav 2:370–372. doi:10.4161/psb.2.5.4463
Kolmos E et al (2009) Integrating ELF4 into the circadian system through combined structural and functional studies. HFSP J 3:350–366. doi:10.2976/1.3218766
Liew LC, Hecht V, Laurie RE, Knowles CL, Schoor JKV, Macknight RC, Weller JL (2009) DIE NEUTRALIS and LATE BLOOMER 1 contribute to regulation of the pea circadian clock. Plant Cell 21:3198–3211. doi:10.1105/tpc.109.067223
Lu SX, Knowles SM, Andronis C, Ong MS, Tobin EM (2009) CIRCADIAN CLOCK ASSOCIATED1 and LATE ELONGATED HYPOCOTYL function synergistically in the circadian clock of Arabidopsis. Plant Physiol 150:834–843. doi:10.1104/pp. 108.133272
McWatters HG et al (2007) ELF4 is required for oscillatory properties of the circadian clock. Plant Physiol 144:391–401. doi:10.1104/pp. 107.096206
Michael TP, McClung CR (2002) Phase-specific circadian clock regulatory elements in Arabidopsis. Plant Physiol 130:627–638. doi:10.1104/pp. 004929
Mizoguchi T et al (2002a) LHY and CCA1 are partially redundant genes required to maintain circadian rhythms in Arabidopsis. Dev Cell 2:629–641. doi:10.1016/s1534-5807(02)00170-3
Mizoguchi TWK, Hanzawa Y, Wright L, Mizoguchi M, Song HR, IA C ́, Coupland G (2002b) LHY and CCA1 are partially redundant genes required to maintain circadian rhythms in Arabidopsis. Dev Cell 2:13
Nusinow DA et al (2011) The ELF4-ELF3-LUX complex links the circadian clock to diurnal control of hypocotyl growth. Nature 475:398–U161. doi:10.1038/nature10182
Park DH et al (1999) Control of circadian rhythms and photoperiodic flowering by the Arabidopsis GIGANTEA gene. Science 285:1579–1582. doi:10.1126/science.285.5433.1579
Piechulla B, Merforth N, Rudolph B (1998) Identification of tomato Lhc promoter regions necessary for circadian expression. Plant Mol Biol 38:655–662. doi:10.1023/a:1006094015513
Qin QP et al (2012) The cold awakening of Doritaenopsis ‘Tinny Tender’ orchid flowers: the role of leaves in cold-induced bud dormancy release. J Plant Growth Regul 31:139–155. doi:10.1007/s00344-011-9226-8
Rombauts S, Dehais P, Van Montagu M, Rouze P (1999) PlantCARE, a plant cis-acting regulatory element database. Nucleic Acids Res 27:295–296. doi:10.1093/nar/27.1.295
Sakanishi Y, Imanishi H, Ishida G (1980) Effect of temperature on growth and flowering of Phalaenopsis-amabilis. Bull Univ Osaka Prefecture Ser B, Agric Life Sci 32:1–10
Sun X, Qin Q, Zhang J, Zhang C, Zhou M, Paek KY, Cui Y (2012) Isolation and characterization of the FVE gene of a Doritaenopsis hybrid involved in the regulation of flowering. Plant Growth Regul 68:77–86. doi:10.1007/s10725-012-9695-1
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731–2739. doi:10.1093/molbev/msr121
Vaz AP, Figueiredo-Ribeiro Rd Rde C, Kerbauy GB (2004a) Photoperiod and temperature effects on in vitro growth and flowering of P. pusilla, an epiphytic orchid. Plant Physiol Biochem : PPB / Soc Fr Physiol Veg 42:411–415. doi:10.1016/j.plaphy.2004.03.008
Vaz APA, Figueiredo-Ribeiro RDL, Kerbauy GB (2004b) Photoperiod and temperature effects on in vitro growth and flowering of P-pusilla, an epiphytic orchid. Plant Physiol Biochem 42:411–415. doi:10.1016/j.plaphy.2004.03.008
Wang WY, Chen WS, Chen WH, Hung LS, Chang PS (2002) Influence of abscisic acid on flowering in Phalaenopsis hybrida. Plant Physiol Biochem 40:97–100. doi:10.1016/s0981-9428(01)01339-0
Wang WY, Chen WS, Huang KL, Hung LS, Chen WH, Su WR (2003) The effects of day length on protein synthesis and flowering in Doritis pulcherrima. Sci Hortic 97:49–56. doi:10.1016/s0304-4238(02)00128-0
Xuan N, Jin Y, Zhang H, Xie Y, Liu Y, Wang G (2011) A putative maize zinc-finger protein gene, ZmAN13, participates in abiotic stress response. Plant Cell Tissue Organ Cult 107:101–112. doi:10.1007/s11240-011-9962-2
Zakizadeh H, Stummann BM, Luetken H, Mueller R (2010) Isolation and characterization of four somatic embryogenesis receptor-like kinase (RhSERK) genes from miniature potted rose (Rosa hybrida cv. Linda). Plant Cell Tissue Organ Cult 101:331–338. doi:10.1007/s11240-010-9693-9
Acknowledgments
We thank Dr. Zhi John Lu (MOE Key of Bioinformatics, School of Life Science, Tsinghua University) for the useful suggestions on improving the manuscript. This study was supported by the National Natural Science Foundation of China (Grant Nos. 30771762, 31170658, and 31301811) and the Science & Technology Key Project of Zhejiang Province (Grant No. 2012C12909-11).
Authors’ Contribution
W. Chen designed, performed, and managed the research work and wrote the manuscript. Q. Qin improved the manuscript. Y. Zheng and C. Wang supplied the materials. S. Wang analyzed some data. M. Zhou, C. Zhang, and Y. Cui managed the experiment partly.
Conflict of Interest
The authors declare that they have no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Electronic Supplementary Material
Below is the link to the electronic supplementary material.
Online Resource 1
Sequence information on alignment of the DhEFL4 amino acid sequences with ELF4 and ELF4-like from different plant species. (DOCX 16 kb)
Online Resource 2
Sequence information on phylogenetic analysis of the DhEFL4 encoded amino acid sequences from different plant species. (DOCX 83 kb)
Online Resource 3
Nucleotide and deduced amino acid sequences of DhEFL4 cDNA. The DUF1313 superfamily region is indicated with a parenthesis in bold. (GIF 21 kb)
Online Resource 4
Putative promoter region sequence and its transcription factor binding motifs, as predicted from PLACE and PlantCARE. (GIF 138 kb)
Online Resource 5
Cis-acting regulatory elements analysis of promoter sequences by PLACE and PlantCARE. (DOCX 24 kb)
Table S1
Sequences of the primers used in this study. (DOCX 132 kb)
Rights and permissions
About this article
Cite this article
Chen, W., Qin, Q., Zheng, Y. et al. Overexpression of Doritaenopsis Hybrid EARLY FLOWERING 4-like4 Gene, DhEFL4, Postpones Flowering in Transgenic Arabidopsis . Plant Mol Biol Rep 34, 103–117 (2016). https://doi.org/10.1007/s11105-015-0899-1
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11105-015-0899-1