Abstract
Key message
Transgenomics for gene discovery in Populus euphratica.
Abstract
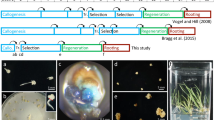
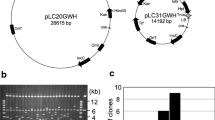
Transgenomics, a member of the omics family of methodologies, is characterized as the introduction of DNA from one organism into another on a genome-wide scale followed by the identification of recipients with altered phenotypes. This strategy allows investigators to identify the gene(s) involved in these phenotypic changes. It is particularly promising for woody plants that have a long life cycle and for which molecular tools are limited. In this study, we constructed a large-insert binary bacterial artificial chromosome library of Populus euphratica, a stress-tolerant poplar species, which included 55,296 clones with average insert sizes of about 127 kb. To date, 1077 of the clones have been transformed into Arabidopsis thaliana via Agrobacterium by the floral dip method. Of these, 69 transgenic lines showed phenotypic changes represented by diverse aspects of plant form and development, 22 of which were reproducibly associated with the same phenotypic change. One of the clones conferring transgenic plants with increased salt tolerance, 002A1F06, was further analyzed and the 127,284 bp insert in this clone harbored eight genes that have been previously reported to be involved in stress resistance. This study demonstrates that transgenomics is useful in the study of functional genomics of woody plants and in the identification of novel gene(s) responsible for economically important traits. Thus, transgenomics can also be used for validation of quantitative trait loci mapped by molecular markers.




Similar content being viewed by others
Change history
26 November 2018
This article (Zhou et al. 2018) has been retracted by the authors because the sequence BIBAC 002A111F06 was incorrectly assigned to the wrong bacterial species. The BIBAC 002A111F06 sequence (GenBank Accession KC129717) reported in the paper was attributed to Populus euphratica Oliv. The BLAST search of this KC129717 sequence against the nr database at NCBI showed that it has very high similarity to a genomic sequence from the gram-negative bacteria Stenotrophomonas maltophilia. The bacterium associates with Populus euphratica Oliv. and DNA isolated from Populus euphratica Oliv. for the construction of the BIBAC clone library inlcuded DNA from Stenotrophomonas maltophilia. Therefore, the phenotype of the transgenic Arabidopsis line carrying the KC129717 sequence cannot be attributed to genes from Populus euphratica Oliv. The authors apologize for the confusion and misinterpretation of our data resulting from the incorrect sequence assignment. All authors agree to this retraction.
References
Ali S, Bakkeren G (2011) Introduction of large DNA inserts into the barley pathogenic fungus, Ustilago hordei, via recombined binary BAC vectors and Agrobacterium-mediated transformation. Curr Genet 57:63–73
Ali S, Bakkeren G (2015) Conversion of BAC clones into binary BAC (BIBAC) vectors and their delivery into basidiomycete fungal cells using Agrobacterium tumefaciens. Methods Mol Biol 1227:199–215
Alonso JM, Stepanova AN, Leisse TJ, Kim CJ, Chen H, Shinn P, Stevenson DK, Zimmerman J, Barajas P, Cheuk R (2003) Genome-wide insertional mutagenesis of Arabidopsis thaliana. Sci Signal 301:653–657
Anggoro DT, Tark-Dame M, Walmsley A, Oka R, de Sain M, Stam M (2017) BIBAC-GW-based vectors for generating reporter lines for site-specific genome editing in planta. Plasmid 89:27–36
Baum DA (2002) Identifying the genetic causes of phenotypic evolution: a review of experimental strategies. Syst Assoc Spec Vol 65:493–507
Brinker M, Brosche M, Vinocur B, Abo Ogiala A, Fayyaz P, Janz D, Ottow EA, Cullmann AD, Altman A, Polle A (2010) Linking the salt transcriptome with physiological responses of a salt-resistant Populus species as a strategy to identify genes important for stress acclimation. Plant Physiol 154:1697–1709
Brosché M, Vinocur B, Alatalo ER, Lamminmäki A, Teichmann T, Ottow EA, Djilianov D, Afif D, Bogeat-Triboulot M-B, Auvinen P, Kangasjärvi J (2005) Gene expression and metabolite profiling of Populus euphratica growing in the Negev desert. Genome Biol 6:R101
Cantu D, Vicente AR, Greve LC, Dewey FM, Bennett AB, Labavitch JM, Powell AL (2008) The intersection between cell wall disassembly, ripening, and fruit susceptibility to Botrytis cinerea. Proc Natl Acad Sci 105:859–864
Chang YL, Henriquez X, Preuss D, Copenhaver GP, Zhang HB (2003) A plant-transformation-competent BIBAC library from the Arabidopsis thaliana Landsberg ecotype for functional and comparative genomics. Theor Appl Genet 106:269–276
Chang YL, Chuang HW, Meksem K, Wu FC, Chang C, Zhang MP, Zhang HB (2011) Characterization of a plant-transformation-ready large-insert BIBAC library of Arabidopsis and bombardment transformation of a large-insert BIBAC of the library into tobacco. Genome 54:437–447
Chauhan R, Farman M, Zhang H, Leong S (2002) Genetic and physical mapping of a rice blast resistance locus, Pi-CO39(t), that corresponds to the avirulence gene AVR1-CO39 of Magnaporthe grisea. Mol Genet Genomics 267:603–612
Chen S, Polle A (2010) Salinity tolerance of Populus. Plant Biol 12:317–333
Cheng H, Endo A, Zhou L, Penney J, HueiChi C, Arroyo A, Leon P, Koshiba T, Sheen J (2002) A unique short-chain dehydrogenase/reductase in Arabidopsis glucose signaling and abscisic acid biosynthesis and functions. Plant Cell Online 14:2723–2743
Clough SJ, Bent AF (1998) Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J 16:735–743
Correa R, Baum DA (2015) Evolutionary transgenomics: prospects and challenges. Front Plant Sci 6:858
Correa R, Stanga J, Larget B, Roznowski A, Shu G, Dilkes B, Baum DA (2012) An assessment of transgenomics as a tool for identifying genes involved in the evolutionary differentiation of closely related plant species. New Phytol 193:494–503
Cui Y, Bi YM, Brugiere N, Arnoldo M, Rothstein SJ (2000) The S locus glycoprotein and the S receptor kinase are sufficient for self-pollen rejection in Brassica. Proc Natl Acad Sci 97:3713–3717
Fang X, Gu S, Xu Z, Chen F, Guo D, Zhang HB, Wu N (2004) Construction of a binary BAC library for an apomictic monosomic addition line of Beta corolliflora in sugar beet and identification of the clones derived from the alien chromosome. Theor Appl Genet 108:1420–1425
Feldmann KA (1991) T-DNA insertion mutagenesis in Arabidopsis: mutational spectrum. Plant J 1:71–82
Feng J, Vick BA, Lee MK, Zhang HB, Jan CC (2006) Construction of BAC and BIBAC libraries from sunflower and identification of linkage group-specific clones by overgo hybridization. Theor Appl Genet 113:23–32
Feng J, Liu Z, Cai X, Jan CC (2013) Toward a molecular cytogenetic map for cultivated sunflower (Helianthus annuus L.) by landed BAC/BIBAC clones. G3 3: 31–40
Frary A, Hamilton CM (2001) Efficiency and stability of high molecular weight DNA transformation an analysis in tomato. Transgenic Res 10:121–132
Ginzberg I, Stein H, Kapulnik Y, Szabados L, Strizhov N, Schell J, Koncz C, Zilberstein A (1998) Isolation and characterization of two different cDNAs of ∆1-pyrroline-5-carboxylate synthase in alfalfa, transcriptionally induced upon salt stress. Plant Mol Biol 38:755–764
Gomase VS, Tagore S (2008) Transgenomics. Gene Ther Mol Biol 12:77–82
Gu R, Fonseca S, Puskás LG, Hackler L, Zvara Á, Dudits D, Pais MS (2004) Transcript identification and profiling during salt stress and recovery of Populus euphratica. Tree Physiol 24:265–276
Hamilton CM (1997) A binary-BAC system for plant transformation with high-molecular-weight DNA. Gene 200:107–116
Hamilton CM, Frary A, Lewis C, Tankley SD (1996) Stable transfer of intact high molecular weight DNA into plant chromosomes. Proc Natl Acad Sci 93:9975–9979
Hamilton CM, Frary A, Xu Y, Tanksley SD, Zhang HB (1999) Construction of tomato genomic DNA libraries in a binary-BAC (BIBAC) vector. Plant J 18:223–229
He R (2016) Multigene engineering in rice using high-capacity Agrobacterium tumefaciens BIBAC vectors. Methods Mol Biol 1385:29–37
He R, Wang Y, Shi Z, Ren X, Zhu L, Weng Q, He G (2003) Construction of a genomic library of wild rice and Agrobacterium-mediated transformation of large insert DNA linked to BPH resistance locus. Gene 321:113–121
Igarashi Y, Yoshiba Y, Sanada Y, Yamaguchi-Shinozaki K, Wada K, Shinozaki K (1997) Characterization of the gene for ∆1-pyrroline-5-carboxylate synthetase and correlation between the expression of the gene and salt tolerance in Oryza sativa L. Plant Mol Biol 33:857–865
Jansson S, Douglas CJ (2007) Populus: a model system for plant biology. Annu Rev Plant Biol 58:435–458
Koncz C, Schell J (1986) The promoter of TL-DNA gene 5 controls the tissue-specific expression of chimaeric genes carried by a novel type of Agrobacterium binary vector. Mol Genet Genomics 204:383–396
Koornneef M, Meinke D (2010) The development of Arabidopsis as a model plant. Plant J 61:909–921
Krysan PJ, Young JC, Sussman MR (1999) T-DNA as an insertional mutagen in Arabidopsis. Plant Cell Online 11:2283–2290
Lee MK, Zhang Y, Zhang M, Goebel M, Kim HJ, Triplett BA, Stelly DM, Zhang HB (2013) Construction of a plant-transformation-competent BIBAC library and genome sequence analysis of polyploid Upland cotton (Gossypium hirsutum L.). BMC Genomics 14:208
Lichtenzveig J, Scheuring C, Dodge J, Abbo S, Zhang HB (2004) Construction of BAC and BIBAC libraries and their applications for generation of SSR markers for genome analysis of chickpea, Cicer arietinum L. Theor Appl Genet 110:492–510
Ma T, Wang J, Zhou G, Yue Z, Hu Q, Chen Y, Liu B, Qiu Q, Tuskan GA, Jianquan L (2013) Genomic insights into salt adaptation in a desert poplar. Nat Commun 4:2797
Mansour MMF, Salama KHA, Al-Mutawa MM (2003) Transport proteins and salt tolerance in plants. Plant Sci 164:891–900
Martinez-Atienza J, Jiang X, Garciadeblas B, Mendoza I, Zhu JK, Pardo JM, Quintero FJ (2007) Conservation of the salt overly sensitive pathway in rice. Plant Physiol 143:1001–1012
Meksem K, Zobrist K, Ruben E, Hyten D, Quanzhou T, Zhang H, foot DL (2000) Two large-insert soybean genomic libraries constructed in a binary vector: applications in chromosome walking and genome wide physical mapping. Theor Appl Genet 101:747–755
Moullet O, Zhang H, Lagudah E (1999) Construction and characterization of a large DNA insert library from the D genome of wheat. Theor Appl Genet 99:305–313
Neale DB, Kremer A (2011) Forest tree genomics: growing resources and applications. Nat Rev Genet 12:111–122
Ottow EA (2005) Populus euphratica displays apoplastic sodium accumulation, osmotic adjustment by decreases in calcium and soluble carbohydrates, and develops leaf succulence under salt stress. Plant Physiol 139:1762–1772
Ozturk ZN, Talame V, Deyholos M, Michalowski CB, Galbraith DW, Gozukirmizi N, Tuberosa R (2002) Monitoring large-scale changes in transcript abundance in drought- and salt-stressed barley. Plant Mol Biol 48:551–573
Park SC, Pham BP, Van Duyet L, Jia B, Lee S, Yu R, Han SW, Yang JK, Hahm KS, Cheong GW (2008) Structural and functional characterization of osmotically inducible protein C (OsmC) from Thermococcus kodakaraensis KOD1. Biochim Biophys Acta 1784:783–788
Quintero FJ, Martinez-Atienza J, Villalta I, Jiang X, Kim WY, Fujii H, Mendoza I, Yun DJ, Zhu JK, Pardo JM (2011) Activation of the plasma membrane Na+/H+ antiporter Salt-Overly-Sensitive 1 (SOS1) by phosphorylation of an auto-inhibitory C-terminal domain. Proc Natl Acad Sci 108:2611–2616
Rothberg JM, Hinz W, Rearick TM, Schultz J, Mileski W, Davey M, Leamon JH, Johnson K, Milgrew MJ, Edwards M (2011) An integrated semiconductor device enabling non-optical genome sequencing. Nature 475:348–352
Santos JM, Freire P, Vicente M, Arraiano CM (1999) The stationary-phase morphogene bola from Escherichia coli is induced by stress during early stages of growth. Mol Microbiol 32:789–798
Savage PA, Vosseller K, Kang C, Larimore K, Riedel E, Wojnoonski K, Jungbluth AA, Allison JP (2008) Recognition of a ubiquitous self antigen by prostate cancer-infiltrating CD8+ T lymphocytes. Science 319:215–220
Shi H, Ishitani M, Kim C, Zhu JK (2000) The Arabidopsis thaliana salt tolerance gene SOS1 encodes a putative Na+/H+ antiporter. Proc Natl Acad Sci 97:6896–6901
Streetz KL, Doyonnas R, Grimm D, Jenkins DD, Fuess S, Perryman S, Lin J, Trautwein C, Shizuru J, Blau H (2008) Hepatic parenchymal replacement in mice by transplanted allogeneic hepatocytes is facilitated by bone marrow transplantation and me diated by CD4 cells. Hepatology 47:706–718
Tao OZ, Zhang HB (1998) Cloning and stable maintenance of DNA fragments over 300 kb in Escherichia coli with conventional plasmid-based vectors. Nucleic Acids Res 26:4901–4904
Tao Q, Wang A, Zhang HB (2002) One large-insert plant-transformation-competent BIBAC library and three BAC libraries of Japonica rice for genome research in rice and other grasses. Theor Appl Genet 105:1058–1066
Tuskan GA, Difazio S, Jansson S, Bohlmann J, Grigoriev I, Hellsten U, Putnam N, Ralph S, Rombauts S, Salamov A (2006) The genome of black cottonwood, Populus trichocarpa (Torr. & Gray). Science 313:1596–1604
Vanderauwera S, Vandenbroucke K, Inze A, van de Cotte B, Muhlenbock P, De Rycke R, Naouar N, Van Gaever T, Van Montagu MC, Van Breusegem F (2012) AtWRKY15 perturbation abolishes the mitochondrial stress response that steers osmotic stress tolerance in Arabidopsis. Proc Natl Acad Sci 109:20113–20118
Waki T, Miyashima S, Nakanishi M, Ikeda Y, Hashimoto T, Nakajima K (2013) A GAL4-based targeted activation tagging system in Arabidopsis thaliana. Plant J 73:357–367
Wang W, Wu Y, Li Y, Xie J, Zhang Z, Deng Z, Zhang Y, Yang C, Lai J, Xie Q (2010) A large insert Thellungiella halophila BIBAC library for genomics and identification of stress tolerance genes. Plant Mol Biol 72:91–99
Wang ZY, Xiong L, Li W, Zhu JK, Zhu J (2011) The plant cuticle is required for osmotic stress regulation of abscisic acid biosynthesis and osmotic stress tolerance in Arabidopsis. Plant Cell Online 23:1971–1984
Wang C, Shi X, Liu L, Li H, Ammiraju JS, Kudrna DA, Xiong W, Wang H, Dai Z, Zheng Y, Lai J, Jin W, Messing J, Bennetzen JL, Wing RA, Luo M (2013) Genomic resources for gene discovery, functional genome annotation, and evolutionary studies of maize and its close relatives. Genetics 195:723–737
Wang Y, Zeng H, Zhou X, Huang F, Peng W, Liu L, Xiong W, Shi X, Luo M (2015a) Transformation of rice with large maize genomic DNA fragments containing high content repetitive sequences. Plant Cell Rep 34:1049–1061
Wang SB, Feng JY, Ren WL, Huang B, Zhou L, Wen YJ, Zhang JS, Dunwell. JM, Shizhong. X, Zhang YM (2015b) Improving power and accuracy of genome-wide association studies via a multi-locus mixed linear model methodology. Sci Rep 6:19444
Weigel D, Ahn JH, Blázquez MA, Borevitz JO, Christensen SK, Fankhauser C, Ferra´ndiz C, Kardailsky I, Malancharuvil EJ, Chory J (2000) Activation tagging in Arabidopsis. Plant Physiol 122:1003–1013
Wu HJ, Zhang Z, Wang JY, Oh DH, Dassanayake M, Liu B, Huang Q, Sun HX, Xia R, Xie Q (2012) Insights into salt tolerance from the genome of Thellungiella salsuginea. Proc Natl Acad Sci 109:12219–12224
Yang Q, Chen ZZ, Zhou XF, Yin HB, Li X, Xin XF, Hong XH, Zhu JK, Gong Z (2009) Overexpression of SOS (Salt Overly Sensitive) genes increases salt tolerance in transgenic Arabidopsis. Mol Plant 2:22–31
Zhang HB, Zhao X, Ding X, Paterson A, Wing R (1995) Preparation of megabase-size DNA from plant nuclei. Plant J 7:175–184
Zhang HB, Scheuring CF, Zhang M, Zhang Y, Wu C, Dong J, Li Y (2012) Construction of BIBAC and BAC libraries from a variety of organisms for advanced genomics research. Nat Protoc 7:479–499
Zhu JK (2001) Plant salt tolerance. Trends Plant Sci 6:66–71
Zhu B, Su J, Chang M, Verma SDP, Fan YL, Wu R (1998) Overexpression of a ∆1-pyrroline-5-carboxylate synthetase gene and analysis of tolerance to water- and salt-stress in transgenic rice. Plant Sci 139:41–48
Acknowledgements
We would like to thank Dr. Hong-Bin Zhang of A&M University for guidance in BIBAC library construction. We would also like to thank Drs. Ren-Ying Zhuo (The Research Institute of Subtropical Forestry, Chinese Academy of Forestry), Zhou Zhou (Henan University), Hai-Feng Yang (Inner Mongolia Agricultural University), Min-Sheng Yang (Hebei Agricultural University), Xin-Ye Zhang (Hubei Forestry Academy), Xiao-Ping Ao (Central South University), and Xiao-Yu Li (Huangshan University) for their collaboration on the transformation work. This work was supported by the Special Fund on Essential Research for National Non-profit Institutions to the Chinese Academy of Forestry (CAFYBB2011001).
Author information
Authors and Affiliations
Corresponding author
Additional information
This article has been retracted by the authors because the sequence BIBAC 002A111F06 was incorrectly assigned to the wrong bacterial species. The BIBAC 002A111F06 sequence (GenBank Accession KC129717) reported in the paper was attributed to Populus euphratica Oliv. The BLAST search of this KC129717 sequence against the nr database at NCBI showed that it has very high similarity to a genomic sequence from the gram-negative bacteria Stenotrophomonas maltophilia. The bacterium associates with Populus euphratica Oliv. and DNA isolated from Populus euphratica Oliv. for the construction of the BIBAC clone library inlcuded DNA from Stenotrophomonas maltophilia. Therefore, the phenotype of the transgenic Arabidopsis line carrying the KC129717 sequence cannot be attributed to genes from Populus euphratica Oliv. The authors apologize for the confusion and misinterpretation of our data resulting from the incorrect sequence assignment. All authors agree to this retraction.
Electronic supplementary material
Below is the link to the electronic supplementary material.
11103_2018_755_MOESM5_ESM.tiff
Figure S1. The repeatable phenotypic changes of the additional T1 transgenic A. thaliana plants with BIBAC clones listed in Figure 2. The reproducible phenotypic changes of transgenic T1 A. thaliana plants with BIBAC clones in Figure 2. (a) 007A2D05, 135A1H05, and 007A1E12, early flowering. (b) 007A2G09, increased number of rosette leaves. (c) 007A2D06 and 007A1D01, abortion. (d) 002A1D09, 002A1B06, and 017A1H12, dwarfism and late flowering. Seedlings were grown for 5 (a, b) or 7 (c, d) weeks. WT: wild-type plants. (TIFF 4606 KB)
11103_2018_755_MOESM6_ESM.tiff
Figure S2. Additional repeatable phenotypic changes of T1 transgenic A. thaliana plants with BIBAC clones. (a) 007B1C08, early bolting and small rosette leaves. (b) 007A2B03, early bolting. (c) 007A1D10, early bolting. (d) 007A2B05, dwarfism. (e) 007A2F04, dwarfism. Seedlings were grown for 5 (a–d) or 7 (e) weeks. WT: wild-type plants. (TIFF 4706 KB)
About this article
Cite this article
Zhou, J., Liu, X., Zhao, ST. et al. RETRACTED ARTICLE: An assessment of transgenomics as a tool for gene discovery in Populus euphratica Oliv.. Plant Mol Biol 97, 525–535 (2018). https://doi.org/10.1007/s11103-018-0755-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11103-018-0755-4